| mir-19 microRNA precursor family | |
|---|---|
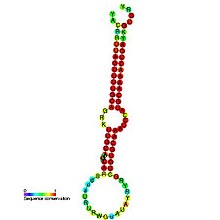 Predicted
secondary structure and
sequence conservation of mir-19 | |
| Identifiers | |
| Symbol | mir-19 |
| Rfam | RF00245 |
| miRBase | MI0000073 |
| miRBase family | MIPF0000011 |
| Other data | |
| RNA type | Gene; miRNA |
| Domain(s) | Eukaryota |
| GO | GO:0035195 GO:0035068 |
| SO | SO:0001244 |
| PDB structures | PDBe |
There maybe 89 known sequences today in the microRNA 19 (miR-19) familly but it will change too fast. They are found into a large number of vertebrate species. The miR-19 microRNA precursor is a small non-coding RNA molecule that regulates gene expression. Within the human and mouse genome there are three copies of this microRNA that are processed from multiple predicted precursor hairpins: [1] [2] [3]
- mouse:
- * miR-19a on chromosome 14 ( MI0000688)
- * miR-19b-1 on chromosome 14 ( MI0000718)
- * miR-19b-2 on chromosome X ( MI0000546)
- human [1]:
- * miR-19a on chromosome 13 ( MI0000073)
- * miR-19b-1 on chromosome 13 ( MI0000074)
- * miR-19b-2 on chromosome X ( MI000075).
MiR-19 has now been predicted or experimentally confirmed ( MIPF0000011). In this case the mature sequence is excised from the 3' arm of the hairpin precursor.
MiRNA seems to generally be found into different cell types, enriched in
neuronal as well as normal and malignant
hematopoietic cells and
tissues
[4].
The presence of miR-19 has been detected in a diverse range of
vertebrate animals including
green anole (Anolis carolinensis)
[5],
primates (gorilla, human,…)
[6]
[7],
cattle (Bos taurus)
[8], dog
[9],
chinese hamster (Cricetulus griseus)
[10],
zebrafish (Danio rerio)
[11],
horse (Equus caballus)
[12],
Takifugu rubripes
[11],
Tetraodon nigroviridis
[11],
chicken (Gallus gallus)
[13]
[14],
gray short-tailed opossum (Monodelphis domestica)
[15],
platypus (Ornithorhynchus anatinus)
[16],
Japanese medaka (Oryzias latipes)
[17],
Xenopus laevis (frog)
[18],
Tasmanian devil (Sarcophilus harrisii)
[19],
pig (Sus scrofa)
[20] and
Zebra Finch (Taeniopygia guttata)
[21].
In some of these species the presence of miR-19 has been shown experimentally, in others the genes encoding miR-19 have been predicted computationally
[1].
MiR-17-92 cluster was identified to encode 6 single mature miRNA ( miR-17, [ [1]], miR-19, miR-20, miR-92, miR-106) containing the first oncogenic miRNA.
MicroRNA from miR-19 family can be expressed from:
- * T-cell acute lymphoblastic leukemia [22]
- * B-cell lymphomas [23]
- * cell lines [22]
- * Cerebellum [24] [25]
- * Purkinje cells [24]
- * HeLa cells [26]
They finaly have tissues-specific miRNA expression. These microRNA are considered as
oncogenes which improve
proliferation,
inhibits
apoptosis and induce
tumor
angiogenesis
[27].
Those miRNA are context-specifics and they have different rôles depending on the place they are.
Ectopic expression of miR-19 represses
CYLD expression, while miR-19 inhibitor treatment induces CYLD protein expression and decreases
NF-kB expression in the
downstream signaling pathway.
Thus, miR-19, CYLD and NF-kB form a regulatory
feedforward loop, which provides new clues for sustained activation of NF-kB in
T-cell acute
lymphoblastic
leukemia.
[22]
MiR-19 is sufficient to induce T-cell lymphoblastic leukemia activating
Notch1 and accelerate the
disease. It's targets are:
- * Bim (Bcl2L11) gene
- * AMP-activated kinase (Prkaa1) gene
- * E2F1 gene
- * the tumour suppressor phosphatases PTEN
- * PP2A (Ppp2r5e) gene
- * Dock5 protein
MiR-19b coordinates a PI3K pathway acting on cell survival in lymphocytes contributing to leukaemogenesis [28] [29] [30].
This patchway is activated through PTEN loss and can contribute to reduce sensitivity to chemotherapy and (in T-ALL) may impact the efficacity of therapeutic gamma-secretase inhibitors.
Baraniskin and al. study show that miR-21, miR-19, and miR-92a levels in cerebrospinal fluid (CSF) seems to be good biomarkers to diagnose a Primary central nervous system lymphoma (PCNSL). They also demonstrate that miRNAs in plasma are in a resistant form to intrinsic RNase activity, and there is a low RNase activity in the CSF [25].
MiR-19 has been identified as a key responsible for the oncogenic activity, reducing the tumor suppressor gene
PTEN expression and activating
AKT/mTOR pathway. This cluster might be important regulator on cancer and aging
[31]
[32].
Mu and al. demonstrated that the expression of endogenous miR-17-92 is required to suppress
apoptosis in
Myc-driven
B-cell
lymphomas. More specificly, miR-19a and miR-19b are required and sufficient to recapitulate the oncogenic properties of the entire cluster
[23]
[33].
Using prediction algorithms, they found miR-19 targets to the prosurvival functions:
In the cell response to stress, the most important is the post-transcriptional control of the important gene expression to cell survival and apoptosis. MiR-19 regulates the Ras homolog B (RhoB) expression in keratinocytes after ultraviolet (UV) radiation exposition. This phenomenon needs the binding of human antigen R (HuR) to the rhoB mRNA 3'-untranslated region. In this case, HuR positively act on miRNA action. The interaction between HuR and miR-19 with rhoB is lost under UV tratment. Here, miR-19, linked to RhoB, act like a protector against keratinocyte apoptosis. A 52- nucleotide-long sequence of the rhoB 3'-UTR spanning bases 818–870, containing the miR-19 and the HuR binding site was sufficient for UV regulation. This event is UV dependant! [34]
One study on multiple myeloma patients permited to identifyed a selective up-regulation of miR-32 and the miR-17-92 cluster. MiR-19a and miR-19b were shown to down regulate SOCS-1 expression (a specific gene that inhibates IL-6 growth signaling). Therefore, miR-17-92 with miR-21, inhibits apoptosis and promotes cell survival [33].
In this case, miR-17-92 cluster promotes
retinoblastoma due to loss of
Rb family members. The mouse retinal development need miR-17-92 overexpresson with Rb and p107 deletion, but it occured frequent emergence of retinoblastoma and
metastasis to the brain.
Here, the cluster oncogenic function is not mediated by a miR-19/PTEN axis toward apoptosis suppression like in
lymphoma or in
leukemia models. MiR-17-92 increase the proliferative capacity of Rb/p107-deficient in
retinal cells.
Moreover, the Rb family members deletion led to compensatory up-regulation of the
cyclin-dependent kinase inhibitor p21Cip1.
Finaly, the cluster overexpression counteracted p21Cip1 up-regulation, promotes proliferation and drove retinoblastoma formation
[35].
Scientists observed that the loss of function of miR-17-92 cluster is induced in smaller
embryos and postnatal death
[36]. The specific role of this cluster in heart and lung
development remain unclear, but the observations described above show that these miRNAs are normally highly expressed in embryonic lung and decrease with maturity. Moreover,
transgenic expression of these miRNAs specifically in lung epithelium results in severe developmental defects with enhanced proliferation and
inhibition of
differentiation of
epithelial cells.
Furthermore, mouse hematopoiesis occurring in the absence of miR-17-92 leads to an isolated defect in B cell development
[36].
The miR-17-92 cluster containing miR-19 miRNA family is also involved into control endothelial cell functions and nerovascularization. MiRNA cluster ( miR-17, miR-18, miR-19 and miR-20) increased during the induction of endothelial cell differenciation in embryonic stem cells (tested on murine) or induce pluripotent stem cells. Eventhough this cluster regulates vascular integrity and angiogenesis, none of each members has a significant impact on the endothelial differenciation of pluripotent stem cells [37].
It has been showing that the 3' UTR of the ATXN1 gene contains 3 target sites for miR-19, and this microRNA shows moderate down regulation of reporter genes containing the ATXN1 3' UTR. Furthermore, it directly bind to the ATXN1 3´UTR to suppress the translation of ATXN1. ATXN1 is also regulated by miR-101, and miR-130. [24]
MiR-19 regulates tissue factor expression at a post-transcriptional level in breast cancer cells, providing a molecular basis for the selective expression of the tissue factor gene. Thanks to bioinformatics anlyses, scientists predicted microRNA- Binding sites for miR-19, miR-20 and miR-106b in the 3'-UTR tissue factor transcript. Experiments confirmed that it negatively regulates gene expression in MCF-7 cells, and overexpression of miR-19 downregulates tissue factor expression in MDA-MB-231 cells (Human breast cancer cell lines). The main action of miR-19 seems to inhibit protein translation of the tissue factor gene in less invasive breast cancer cells [27].
MiR-19 also takes part in
inflammatory responses
enhancing or
repressing pro-inflammatory mediators expression. It positively regulates
Toll-like receptor sigaling with
Dicer1 deletion and miRNA depletion. MiR-19b is an important protagonist in this phenomenon, regulating positively
NF-kB activity.
MiRNA depletion inhibits
cytokines production by NF-kB. This indicates that miRNA control of NF-kB signaling repressors thanks to its relief. Some important regulators of NF-kB signaling (
like A20 (Tnfaip3),
Cyld, and Cezanne (Otud7b)) is targeted by the miR-17-92 cluster.
Moreover, mir-19 targets some members of the Tnfaip3-ubiquitin editing complex (
Tnfaip3/Itch/
Tnip1/
Rnf11). MiR-19 directly involved in the modulation of several NF-kB signaling negative regulators expression, indicating an important role for Rnf11 in the effect of miR-19b on NF-kB signaling.
Finaly, miR-19b exacerbates the
cells crucial inflammatory activation in
rheumatoid arthritis disease
[26]
[29].
- ^
a
b
c Lagos-Quintana, Mariana; Rauhut, Reinhard; Lendeckel, Winfried; Tuschl, Thomas (2001). "Identification of novel genes coding for small expressed RNAs". Science. 294 (5543): 853–858.
doi:
10.1126/science.1064921.
PMID
11679670.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Mourelatos, Z.; Dostie, J.; Paushkin, S.; Sharma, A.; Charroux, B.; Abel, L.; Rappsilber, J.; Mann, M.; Dreyfuss, G. (2002).
"miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs". Genes Dev. 16 (6): 720–728.
doi:
10.1101/gad.974702.
PMC
155365.
PMID
11914277.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Houbaviy, Hristo B.; Murray, Michael F.; Sharp, Phillip A. (2003). "Embryonic stem cell-specific MicroRNAs". Dev Cell. 5 (2): 351–358.
doi:
10.1016/S1534-5807(03)00227-2.
PMID
12919684.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Landgraf, P.; et al. (2007).
"A Mammalian microRNA Expression Atlas Based on Small RNA Library Sequencing". Cell. 129 (7): 1401–1414.
doi:
10.1016 (inactive 2023-08-02).
PMC
2681231.
PMID
17604727.
{{ cite journal}}: Check|doi=value ( help)CS1 maint: DOI inactive as of August 2023 ( link) CS1 maint: date and year ( link) -
^ Lyson TR, Sperling EA, Heimberg AM and al. (2012).
"MicroRNAs support a turtle + lizard clade". Biol Lett. 8 (1): 104–7.
doi:
10.1098/rsbl.2011.0477.
PMC
3259949.
PMID
21775315.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Berezikov E, Guryev V, van de Belt J and al. (2005). "Phylogenetic shadowing and computational identification of human microRNA genes". Cell. 120 (1): 21–4.
doi:
10.1016/j.cell.2004.12.031.
PMID
15652478.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Lui WO, Pourmand N, Patterson BK and al. (2007). "Patterns of known and novel small RNAs in human cervical cancer". Cancer Res. 67 (13): 6031–43.
doi:
10.1158/0008-5472.CAN-06-0561.
PMID
17616659.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Gu Z, Eleswarapu S, Jiang H (2007). "Identification and characterization of microRNAs from the bovine adipose tissue and mammary gland". FEBS Lett. 581 (5): 981–8.
doi:
10.1016/j.febslet.2007.01.081.
PMID
17306260.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Friedländer MR, Chen W, Adamidi C and al. (2008). "Discovering microRNAs from deep sequencing data using miRDeep". Nat Biotechnol. 26 (4): 407–15.
doi:
10.1038/nbt1394.
PMID
18392026.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Hackl M, Jakobi T, Blom J and al. (2011).
"Next-generation sequencing of the Chinese hamster ovary microRNA transcriptome: Identification, annotation and profiling of microRNAs as targets for cellular engineering". J Biotechnol. 153 (1–2): 62–75.
doi:
10.1016/j.jbiotec.2011.02.011.
PMC
3119918.
PMID
21392545.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Chen PY, Manninga H, Slanchev K and al. (2005).
"The developmental miRNA profiles of zebrafish as determined by small RNA cloning". Genes Dev. 19 (11): 1288–93.
doi:
10.1101/gad.1310605.
PMC
1142552.
PMID
15937218.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Zhou M, Wang Q, Sun J and al. (2009). "In silico detection and characteristics of novel microRNA genes in the Equus caballus genome using an integrated ab initio and comparative genomic approach". Genomics. 94 (2): 125–31.
doi:
10.1016/j.ygeno.2009.04.006.
PMID
19406225.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ International Chicken Genome Sequencing Consortium (2004). "Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution". Nature. 432 (7018): 695–716. doi: 10.1038/nature03154. PMID 15592404.
-
^ Yao Y, Zhao Y, Xu H and al. (2008).
"MicroRNA profile of Marek's disease virus-transformed T-cell line MSB-1: predominance of virus-encoded microRNAs". J Virol. 82 (8): 4007–15.
doi:
10.1128/JVI.02659-07.
PMC
2293013.
PMID
18256158.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ Devor EJ, Samollow PB (2008). "In vitro and in silico annotation of conserved and nonconserved microRNAs in the genome of the marsupial Monodelphis domestica". J Hered. 99 (1): 66–72. doi: 10.1093/jhered/esm085. PMID 17965199.
-
^ Murchison EP, Kheradpour P, Sachidanandam R and al. (2008).
"Conservation of small RNA pathways in platypus". Genome Res. 18 (6): 995–1004.
doi:
10.1101/gr.073056.107.
PMC
2413167.
PMID
18463306.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Li SC, Chan WC, Ho MR and al. (2010).
"Discovery and characterization of medaka miRNA genes by next generation sequencing platform". BMC Genomics. 11 (Suppl 4): S8.
doi:
10.1186/1471-2164-11-S4-S8.
PMC
3005926.
PMID
21143817.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: unflagged free DOI ( link) -
^ Watanabe T, Takeda A, Mise K and al. (2005). "Stage-specific expression of microRNAs during Xenopus development". FEBS Lett. 579 (2): 318–24.
doi:
10.1016/j.febslet.2004.11.067.
PMID
15642338.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Murchison EP, Tovar C, Hsu A and al. (2010).
"The Tasmanian devil transcriptome reveals Schwann cell origins of a clonally transmissible cancer". Science. 327 (5961): 84–7.
doi:
10.1126/science.1180616.
PMC
2982769.
PMID
20044575.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Wernersson R, Schierup MH, Jørgensen FG and al. (2005).
"Pigs in sequence space: a 0.66X coverage pig genome survey based on shotgun sequencing". BMC Genomics. 6: 6:70.
doi:
10.1186/1471-2164-6-70.
PMC
1142312.
PMID
15885146.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: unflagged free DOI ( link) -
^ Warren WC, Clayton DF, Ellegren H and al. (2010).
"The genome of a songbird". Nature. 464 (7289): 757–62.
doi:
10.1038/nature08819.
PMC
3187626.
PMID
20360741.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Huashan Ye, Xiaowen Liu, Meng Lv, Yuliang Wu, Shuzhen Kuang, Jing Gong, Ping Yuan, Zhaodong Zhong, Qiubai Li, Haibo Jia, Jun Sun, Zhichao Chen and An-Yuan Guo (2012). "MicroRNA and transcription factor co-regulatory network analysis reveals miR-19 inhibits CYLD in T-cell acute lymphoblastic leukemia". Nucleic Acids Research. 40.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Ping Mu, Yoon-Chi Han, Doron Betel, Evelyn Yao, Massimo Squatrito, Paul Ogrodowski, Elisa de Stanchina, Aleco D’Andrea, Chris Sander, Andrea Ventura (2009).
"Genetic dissection of the miR-17~92 cluster of microRNAs in Myc-induced B-cell lymphomas". Genes Dev. 23 (24): 2806–11.
doi:
10.1101/gad.1872909.
PMC
2800095.
PMID
20008931.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Lee Y, Samaco RC, Gatchel JR, Thaller C, Orr HT, Zoghbi HY (October 2008).
"miR-19, miR-101 and miR-130 co-regulate ATXN1 levels to potentially modulate SCA1 pathogenesis". Nat. Neurosci. 11 (10): 1137–9.
doi:
10.1038/nn.2183.
PMC
2574629.
PMID
18758459.
{{ cite journal}}: CS1 maint: date and year ( link) CS1 maint: multiple names: authors list ( link) - ^
a
b Alexander Baraniskin, Jan Kuhnhenn, Uwe Schlegel, Andrew Chan, Martina Deckert, Ralf Gold, Abdelouahid Maghnouj, Hannah Zöllner, Anke Reinacher-Schick, Wolff Schmiegel, Stephan A. Hahn, Roland Schroers (2011). "Identification of microRNAs in the cerebrospinal fluid as marker for primary diffuse large B-cell lymphoma of the central nervous system". Blood. 117 (11): 3140–3146.
doi:
10.1182/blood-2010-09-308684.
PMID
21200023.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Michael P. Gantier, H. James Stunden, Claire E. McCoy, Mark A. Behlke, Die Wang, Maria Kaparakis-Liaskos, Soroush T. Sarvestani, Yuan H. Yang, Dakang Xu, Sinéad C. Corr, Eric F. Morand, Bryan R. G. Williams (2012).
"A miR-19 regulon that controls NF-iB signaling". Nucleic Acids Research. 40 (16): 8048–8058.
doi:
10.1093/nar/gks521.
PMC
3439911.
PMID
22684508.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Xiaoxi Zhang, Haijun Yu, Jessica R. Lou, Jie Zheng, Hua Zhu, Narcis-Ioan Popescu, Florea Lupu, Stuart E. Lind, and Wei-Qun Ding (2011).
"MicroRNA-19 (miR-19) Regulates Tissue Factor Expression in Breast Cancer Cells". The Journal of Biological Chemistry. 286 (2): 1429–1435.
doi:
10.1074/jbc.M110.146530.
PMC
3020751.
PMID
21059650.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Konstantinos J. Mavrakis1, Andrew L. Wolfe, Elisa Oricchio1, Teresa Palomero and al. (2011).
"Genome-wide RNAi screen identifies miR-19 targets in Notchinduced acute T-cell leukaemia (T-ALL)". Nat Cell Biol. 12 (4): 372–379.
doi:
10.1038/ncb2037.
PMC
2989719.
PMID
20190740.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: numeric names: authors list ( link) - ^ a b Konstantinos J. Mavrakis and Hans-Guido Wendel (2010). "TargetScreen: an unbiased approach to identify functionally important microRNA targets". Cell Cycle. 9 (11): 2080–4. doi: 10.4161/cc.9.11.11807. PMID 20505335.
-
^ Séverine Landais, Sébastien Landry, Philippe Legault and al. (2007). "Oncogenic Potential of the miR-106-363 Cluster and Its Implication in Human T-Cell Leukemia". Cancer Res. 67 (12): 5699–707.
doi:
10.1158/0008-5472.CAN-06-4478.
PMID
17575136.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Johannes Grillari, Matthias Hackl, Regina Grillari-Voglauer (2010).
"miR-17–92 cluster: ups and downs in cancer and aging". Biogerontology. 11 (4): 501–506.
doi:
10.1007/s10522-010-9272-9.
PMC
2899009.
PMID
20437201.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Virginie Olive, Margaux J. Bennett, James C. Walker and al. (2009).
"miR-19 is a key oncogenic component of mir-17-92". Genes Dev. 23 (24): 2839–49.
doi:
10.1101/gad.1861409.
PMC
2800084.
PMID
20008935.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Flavia Pichiorri, Sung-Suk Suh, Marco Ladetto and al. (2008).
"MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis". Proceedings of the National Academy of Sciences. 105 (35): 12885–90.
doi:
10.1073/pnas.0806202105.
PMC
2529070.
PMID
18728182.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ V Glorian, G Maillot, S Polès and al. (2011).
"HuR-dependent loading of miRNA RISC to the mRNA encoding the Ras-related small GTPase RhoB controls its translation during UV-induced apoptosis". Cell Death and Differentiation. 18 (11): 1692–1701.
doi:
10.1038/cdd.2011.35.
PMC
3190107.
PMID
21527938.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Karina Conkrite, Maggie Sundby, Shizuo Mukai and al. (2011).
"miR-17~92 cooperates with RB pathway mutations to promote retinoblastoma". Genes & Development. 25 (16): 1734–45.
doi:
10.1101/gad.17027411.
PMC
3165937.
PMID
21816922.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ a b Joshua T. Mendell (2008). "miRiad roles for the miR-17-92 cluster in development and disease". Cell. 133 (2): 217–22. doi: 10.1016/j.cell.2008.04.001. PMC 2732113. PMID 18423194.
-
^ Karine Tréguer, Eva-Marie Heinrich, Kisho Ohtani and al. (2012). "Role of the MicroRNA-17–92 Cluster in the Endothelial Differentiation of Stem Cells". Journal of Vascular Research. 49 (5): 447–460.
doi:
10.1159/000339429.
PMID
22797777.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link)
- Andrea Ventura, Amanda G. Young, Monte M. Winslow and al. (2008).
"Targeted deletion reveals essential and overlapping functions of the miR-17~92 family of miRNA clustersMechanical stretch up-regulates microRNA-26a and induces human airway smooth muscle hypertrophy by suppressing glycogen synthase kinase-3β". Cell. 132 (5): 875–886.
doi:
10.1016/j.cell.2008.02.019.
PMC
2323338.
PMID
18329372.
{{ cite journal}}: Unknown parameter|DUPLICATE_doi=ignored ( help)CS1 maint: multiple names: authors list ( link) - Lixin Hong, Maoyi Lai, Michelle Chen and al. (2010).
"The miR-17-92 Cluster of microRNAs Confers Tumorigenicity by Inhibiting Oncogene-Induced Senescence". Cancer Res. 70 (21): 8547–8557.
doi:
10.1016/j.cell.2008.02.019.
PMC
2970743.
PMID
20851997.
{{ cite journal}}: Unknown parameter|DUPLICATE_doi=ignored ( help)CS1 maint: multiple names: authors list ( link) - JR-Shiuan Yang, Michael D. Phillips, Doron Betel and al. (2011).
"Widespread regulatory activity of vertebrate microRNA* species". RNA. 17 (2): 312–26.
doi:
10.1016/j.cell.2008.02.019.
PMC
3022280.
PMID
21177881.
{{ cite journal}}: Unknown parameter|DUPLICATE_doi=ignored ( help)CS1 maint: multiple names: authors list ( link) - Joost Kluiver, Johan H. Gibcus, Chris Hettinga and al. (2012).
"Rapid Generation of MicroRNA Sponges for MicroRNA Inhibition". PLOS ONE. 7 (1): e29275.
doi:
10.1371/journal.pone.0029275.
PMC
3253070.
PMID
22238599.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link)
| mir-19 microRNA precursor family | |
|---|---|
 Predicted
secondary structure and
sequence conservation of mir-19 | |
| Identifiers | |
| Symbol | mir-19 |
| Rfam | RF00245 |
| miRBase | MI0000073 |
| miRBase family | MIPF0000011 |
| Other data | |
| RNA type | Gene; miRNA |
| Domain(s) | Eukaryota |
| GO | GO:0035195 GO:0035068 |
| SO | SO:0001244 |
| PDB structures | PDBe |
There maybe 89 known sequences today in the microRNA 19 (miR-19) familly but it will change too fast. They are found into a large number of vertebrate species. The miR-19 microRNA precursor is a small non-coding RNA molecule that regulates gene expression. Within the human and mouse genome there are three copies of this microRNA that are processed from multiple predicted precursor hairpins: [1] [2] [3]
- mouse:
- * miR-19a on chromosome 14 ( MI0000688)
- * miR-19b-1 on chromosome 14 ( MI0000718)
- * miR-19b-2 on chromosome X ( MI0000546)
- human [1]:
- * miR-19a on chromosome 13 ( MI0000073)
- * miR-19b-1 on chromosome 13 ( MI0000074)
- * miR-19b-2 on chromosome X ( MI000075).
MiR-19 has now been predicted or experimentally confirmed ( MIPF0000011). In this case the mature sequence is excised from the 3' arm of the hairpin precursor.
MiRNA seems to generally be found into different cell types, enriched in
neuronal as well as normal and malignant
hematopoietic cells and
tissues
[4].
The presence of miR-19 has been detected in a diverse range of
vertebrate animals including
green anole (Anolis carolinensis)
[5],
primates (gorilla, human,…)
[6]
[7],
cattle (Bos taurus)
[8], dog
[9],
chinese hamster (Cricetulus griseus)
[10],
zebrafish (Danio rerio)
[11],
horse (Equus caballus)
[12],
Takifugu rubripes
[11],
Tetraodon nigroviridis
[11],
chicken (Gallus gallus)
[13]
[14],
gray short-tailed opossum (Monodelphis domestica)
[15],
platypus (Ornithorhynchus anatinus)
[16],
Japanese medaka (Oryzias latipes)
[17],
Xenopus laevis (frog)
[18],
Tasmanian devil (Sarcophilus harrisii)
[19],
pig (Sus scrofa)
[20] and
Zebra Finch (Taeniopygia guttata)
[21].
In some of these species the presence of miR-19 has been shown experimentally, in others the genes encoding miR-19 have been predicted computationally
[1].
MiR-17-92 cluster was identified to encode 6 single mature miRNA ( miR-17, [ [1]], miR-19, miR-20, miR-92, miR-106) containing the first oncogenic miRNA.
MicroRNA from miR-19 family can be expressed from:
- * T-cell acute lymphoblastic leukemia [22]
- * B-cell lymphomas [23]
- * cell lines [22]
- * Cerebellum [24] [25]
- * Purkinje cells [24]
- * HeLa cells [26]
They finaly have tissues-specific miRNA expression. These microRNA are considered as
oncogenes which improve
proliferation,
inhibits
apoptosis and induce
tumor
angiogenesis
[27].
Those miRNA are context-specifics and they have different rôles depending on the place they are.
Ectopic expression of miR-19 represses
CYLD expression, while miR-19 inhibitor treatment induces CYLD protein expression and decreases
NF-kB expression in the
downstream signaling pathway.
Thus, miR-19, CYLD and NF-kB form a regulatory
feedforward loop, which provides new clues for sustained activation of NF-kB in
T-cell acute
lymphoblastic
leukemia.
[22]
MiR-19 is sufficient to induce T-cell lymphoblastic leukemia activating
Notch1 and accelerate the
disease. It's targets are:
- * Bim (Bcl2L11) gene
- * AMP-activated kinase (Prkaa1) gene
- * E2F1 gene
- * the tumour suppressor phosphatases PTEN
- * PP2A (Ppp2r5e) gene
- * Dock5 protein
MiR-19b coordinates a PI3K pathway acting on cell survival in lymphocytes contributing to leukaemogenesis [28] [29] [30].
This patchway is activated through PTEN loss and can contribute to reduce sensitivity to chemotherapy and (in T-ALL) may impact the efficacity of therapeutic gamma-secretase inhibitors.
Baraniskin and al. study show that miR-21, miR-19, and miR-92a levels in cerebrospinal fluid (CSF) seems to be good biomarkers to diagnose a Primary central nervous system lymphoma (PCNSL). They also demonstrate that miRNAs in plasma are in a resistant form to intrinsic RNase activity, and there is a low RNase activity in the CSF [25].
MiR-19 has been identified as a key responsible for the oncogenic activity, reducing the tumor suppressor gene
PTEN expression and activating
AKT/mTOR pathway. This cluster might be important regulator on cancer and aging
[31]
[32].
Mu and al. demonstrated that the expression of endogenous miR-17-92 is required to suppress
apoptosis in
Myc-driven
B-cell
lymphomas. More specificly, miR-19a and miR-19b are required and sufficient to recapitulate the oncogenic properties of the entire cluster
[23]
[33].
Using prediction algorithms, they found miR-19 targets to the prosurvival functions:
In the cell response to stress, the most important is the post-transcriptional control of the important gene expression to cell survival and apoptosis. MiR-19 regulates the Ras homolog B (RhoB) expression in keratinocytes after ultraviolet (UV) radiation exposition. This phenomenon needs the binding of human antigen R (HuR) to the rhoB mRNA 3'-untranslated region. In this case, HuR positively act on miRNA action. The interaction between HuR and miR-19 with rhoB is lost under UV tratment. Here, miR-19, linked to RhoB, act like a protector against keratinocyte apoptosis. A 52- nucleotide-long sequence of the rhoB 3'-UTR spanning bases 818–870, containing the miR-19 and the HuR binding site was sufficient for UV regulation. This event is UV dependant! [34]
One study on multiple myeloma patients permited to identifyed a selective up-regulation of miR-32 and the miR-17-92 cluster. MiR-19a and miR-19b were shown to down regulate SOCS-1 expression (a specific gene that inhibates IL-6 growth signaling). Therefore, miR-17-92 with miR-21, inhibits apoptosis and promotes cell survival [33].
In this case, miR-17-92 cluster promotes
retinoblastoma due to loss of
Rb family members. The mouse retinal development need miR-17-92 overexpresson with Rb and p107 deletion, but it occured frequent emergence of retinoblastoma and
metastasis to the brain.
Here, the cluster oncogenic function is not mediated by a miR-19/PTEN axis toward apoptosis suppression like in
lymphoma or in
leukemia models. MiR-17-92 increase the proliferative capacity of Rb/p107-deficient in
retinal cells.
Moreover, the Rb family members deletion led to compensatory up-regulation of the
cyclin-dependent kinase inhibitor p21Cip1.
Finaly, the cluster overexpression counteracted p21Cip1 up-regulation, promotes proliferation and drove retinoblastoma formation
[35].
Scientists observed that the loss of function of miR-17-92 cluster is induced in smaller
embryos and postnatal death
[36]. The specific role of this cluster in heart and lung
development remain unclear, but the observations described above show that these miRNAs are normally highly expressed in embryonic lung and decrease with maturity. Moreover,
transgenic expression of these miRNAs specifically in lung epithelium results in severe developmental defects with enhanced proliferation and
inhibition of
differentiation of
epithelial cells.
Furthermore, mouse hematopoiesis occurring in the absence of miR-17-92 leads to an isolated defect in B cell development
[36].
The miR-17-92 cluster containing miR-19 miRNA family is also involved into control endothelial cell functions and nerovascularization. MiRNA cluster ( miR-17, miR-18, miR-19 and miR-20) increased during the induction of endothelial cell differenciation in embryonic stem cells (tested on murine) or induce pluripotent stem cells. Eventhough this cluster regulates vascular integrity and angiogenesis, none of each members has a significant impact on the endothelial differenciation of pluripotent stem cells [37].
It has been showing that the 3' UTR of the ATXN1 gene contains 3 target sites for miR-19, and this microRNA shows moderate down regulation of reporter genes containing the ATXN1 3' UTR. Furthermore, it directly bind to the ATXN1 3´UTR to suppress the translation of ATXN1. ATXN1 is also regulated by miR-101, and miR-130. [24]
MiR-19 regulates tissue factor expression at a post-transcriptional level in breast cancer cells, providing a molecular basis for the selective expression of the tissue factor gene. Thanks to bioinformatics anlyses, scientists predicted microRNA- Binding sites for miR-19, miR-20 and miR-106b in the 3'-UTR tissue factor transcript. Experiments confirmed that it negatively regulates gene expression in MCF-7 cells, and overexpression of miR-19 downregulates tissue factor expression in MDA-MB-231 cells (Human breast cancer cell lines). The main action of miR-19 seems to inhibit protein translation of the tissue factor gene in less invasive breast cancer cells [27].
MiR-19 also takes part in
inflammatory responses
enhancing or
repressing pro-inflammatory mediators expression. It positively regulates
Toll-like receptor sigaling with
Dicer1 deletion and miRNA depletion. MiR-19b is an important protagonist in this phenomenon, regulating positively
NF-kB activity.
MiRNA depletion inhibits
cytokines production by NF-kB. This indicates that miRNA control of NF-kB signaling repressors thanks to its relief. Some important regulators of NF-kB signaling (
like A20 (Tnfaip3),
Cyld, and Cezanne (Otud7b)) is targeted by the miR-17-92 cluster.
Moreover, mir-19 targets some members of the Tnfaip3-ubiquitin editing complex (
Tnfaip3/Itch/
Tnip1/
Rnf11). MiR-19 directly involved in the modulation of several NF-kB signaling negative regulators expression, indicating an important role for Rnf11 in the effect of miR-19b on NF-kB signaling.
Finaly, miR-19b exacerbates the
cells crucial inflammatory activation in
rheumatoid arthritis disease
[26]
[29].
- ^
a
b
c Lagos-Quintana, Mariana; Rauhut, Reinhard; Lendeckel, Winfried; Tuschl, Thomas (2001). "Identification of novel genes coding for small expressed RNAs". Science. 294 (5543): 853–858.
doi:
10.1126/science.1064921.
PMID
11679670.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Mourelatos, Z.; Dostie, J.; Paushkin, S.; Sharma, A.; Charroux, B.; Abel, L.; Rappsilber, J.; Mann, M.; Dreyfuss, G. (2002).
"miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs". Genes Dev. 16 (6): 720–728.
doi:
10.1101/gad.974702.
PMC
155365.
PMID
11914277.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Houbaviy, Hristo B.; Murray, Michael F.; Sharp, Phillip A. (2003). "Embryonic stem cell-specific MicroRNAs". Dev Cell. 5 (2): 351–358.
doi:
10.1016/S1534-5807(03)00227-2.
PMID
12919684.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Landgraf, P.; et al. (2007).
"A Mammalian microRNA Expression Atlas Based on Small RNA Library Sequencing". Cell. 129 (7): 1401–1414.
doi:
10.1016 (inactive 2023-08-02).
PMC
2681231.
PMID
17604727.
{{ cite journal}}: Check|doi=value ( help)CS1 maint: DOI inactive as of August 2023 ( link) CS1 maint: date and year ( link) -
^ Lyson TR, Sperling EA, Heimberg AM and al. (2012).
"MicroRNAs support a turtle + lizard clade". Biol Lett. 8 (1): 104–7.
doi:
10.1098/rsbl.2011.0477.
PMC
3259949.
PMID
21775315.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Berezikov E, Guryev V, van de Belt J and al. (2005). "Phylogenetic shadowing and computational identification of human microRNA genes". Cell. 120 (1): 21–4.
doi:
10.1016/j.cell.2004.12.031.
PMID
15652478.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Lui WO, Pourmand N, Patterson BK and al. (2007). "Patterns of known and novel small RNAs in human cervical cancer". Cancer Res. 67 (13): 6031–43.
doi:
10.1158/0008-5472.CAN-06-0561.
PMID
17616659.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Gu Z, Eleswarapu S, Jiang H (2007). "Identification and characterization of microRNAs from the bovine adipose tissue and mammary gland". FEBS Lett. 581 (5): 981–8.
doi:
10.1016/j.febslet.2007.01.081.
PMID
17306260.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Friedländer MR, Chen W, Adamidi C and al. (2008). "Discovering microRNAs from deep sequencing data using miRDeep". Nat Biotechnol. 26 (4): 407–15.
doi:
10.1038/nbt1394.
PMID
18392026.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Hackl M, Jakobi T, Blom J and al. (2011).
"Next-generation sequencing of the Chinese hamster ovary microRNA transcriptome: Identification, annotation and profiling of microRNAs as targets for cellular engineering". J Biotechnol. 153 (1–2): 62–75.
doi:
10.1016/j.jbiotec.2011.02.011.
PMC
3119918.
PMID
21392545.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Chen PY, Manninga H, Slanchev K and al. (2005).
"The developmental miRNA profiles of zebrafish as determined by small RNA cloning". Genes Dev. 19 (11): 1288–93.
doi:
10.1101/gad.1310605.
PMC
1142552.
PMID
15937218.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Zhou M, Wang Q, Sun J and al. (2009). "In silico detection and characteristics of novel microRNA genes in the Equus caballus genome using an integrated ab initio and comparative genomic approach". Genomics. 94 (2): 125–31.
doi:
10.1016/j.ygeno.2009.04.006.
PMID
19406225.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ International Chicken Genome Sequencing Consortium (2004). "Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution". Nature. 432 (7018): 695–716. doi: 10.1038/nature03154. PMID 15592404.
-
^ Yao Y, Zhao Y, Xu H and al. (2008).
"MicroRNA profile of Marek's disease virus-transformed T-cell line MSB-1: predominance of virus-encoded microRNAs". J Virol. 82 (8): 4007–15.
doi:
10.1128/JVI.02659-07.
PMC
2293013.
PMID
18256158.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ Devor EJ, Samollow PB (2008). "In vitro and in silico annotation of conserved and nonconserved microRNAs in the genome of the marsupial Monodelphis domestica". J Hered. 99 (1): 66–72. doi: 10.1093/jhered/esm085. PMID 17965199.
-
^ Murchison EP, Kheradpour P, Sachidanandam R and al. (2008).
"Conservation of small RNA pathways in platypus". Genome Res. 18 (6): 995–1004.
doi:
10.1101/gr.073056.107.
PMC
2413167.
PMID
18463306.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Li SC, Chan WC, Ho MR and al. (2010).
"Discovery and characterization of medaka miRNA genes by next generation sequencing platform". BMC Genomics. 11 (Suppl 4): S8.
doi:
10.1186/1471-2164-11-S4-S8.
PMC
3005926.
PMID
21143817.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: unflagged free DOI ( link) -
^ Watanabe T, Takeda A, Mise K and al. (2005). "Stage-specific expression of microRNAs during Xenopus development". FEBS Lett. 579 (2): 318–24.
doi:
10.1016/j.febslet.2004.11.067.
PMID
15642338.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Murchison EP, Tovar C, Hsu A and al. (2010).
"The Tasmanian devil transcriptome reveals Schwann cell origins of a clonally transmissible cancer". Science. 327 (5961): 84–7.
doi:
10.1126/science.1180616.
PMC
2982769.
PMID
20044575.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Wernersson R, Schierup MH, Jørgensen FG and al. (2005).
"Pigs in sequence space: a 0.66X coverage pig genome survey based on shotgun sequencing". BMC Genomics. 6: 6:70.
doi:
10.1186/1471-2164-6-70.
PMC
1142312.
PMID
15885146.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: unflagged free DOI ( link) -
^ Warren WC, Clayton DF, Ellegren H and al. (2010).
"The genome of a songbird". Nature. 464 (7289): 757–62.
doi:
10.1038/nature08819.
PMC
3187626.
PMID
20360741.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Huashan Ye, Xiaowen Liu, Meng Lv, Yuliang Wu, Shuzhen Kuang, Jing Gong, Ping Yuan, Zhaodong Zhong, Qiubai Li, Haibo Jia, Jun Sun, Zhichao Chen and An-Yuan Guo (2012). "MicroRNA and transcription factor co-regulatory network analysis reveals miR-19 inhibits CYLD in T-cell acute lymphoblastic leukemia". Nucleic Acids Research. 40.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Ping Mu, Yoon-Chi Han, Doron Betel, Evelyn Yao, Massimo Squatrito, Paul Ogrodowski, Elisa de Stanchina, Aleco D’Andrea, Chris Sander, Andrea Ventura (2009).
"Genetic dissection of the miR-17~92 cluster of microRNAs in Myc-induced B-cell lymphomas". Genes Dev. 23 (24): 2806–11.
doi:
10.1101/gad.1872909.
PMC
2800095.
PMID
20008931.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Lee Y, Samaco RC, Gatchel JR, Thaller C, Orr HT, Zoghbi HY (October 2008).
"miR-19, miR-101 and miR-130 co-regulate ATXN1 levels to potentially modulate SCA1 pathogenesis". Nat. Neurosci. 11 (10): 1137–9.
doi:
10.1038/nn.2183.
PMC
2574629.
PMID
18758459.
{{ cite journal}}: CS1 maint: date and year ( link) CS1 maint: multiple names: authors list ( link) - ^
a
b Alexander Baraniskin, Jan Kuhnhenn, Uwe Schlegel, Andrew Chan, Martina Deckert, Ralf Gold, Abdelouahid Maghnouj, Hannah Zöllner, Anke Reinacher-Schick, Wolff Schmiegel, Stephan A. Hahn, Roland Schroers (2011). "Identification of microRNAs in the cerebrospinal fluid as marker for primary diffuse large B-cell lymphoma of the central nervous system". Blood. 117 (11): 3140–3146.
doi:
10.1182/blood-2010-09-308684.
PMID
21200023.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Michael P. Gantier, H. James Stunden, Claire E. McCoy, Mark A. Behlke, Die Wang, Maria Kaparakis-Liaskos, Soroush T. Sarvestani, Yuan H. Yang, Dakang Xu, Sinéad C. Corr, Eric F. Morand, Bryan R. G. Williams (2012).
"A miR-19 regulon that controls NF-iB signaling". Nucleic Acids Research. 40 (16): 8048–8058.
doi:
10.1093/nar/gks521.
PMC
3439911.
PMID
22684508.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Xiaoxi Zhang, Haijun Yu, Jessica R. Lou, Jie Zheng, Hua Zhu, Narcis-Ioan Popescu, Florea Lupu, Stuart E. Lind, and Wei-Qun Ding (2011).
"MicroRNA-19 (miR-19) Regulates Tissue Factor Expression in Breast Cancer Cells". The Journal of Biological Chemistry. 286 (2): 1429–1435.
doi:
10.1074/jbc.M110.146530.
PMC
3020751.
PMID
21059650.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Konstantinos J. Mavrakis1, Andrew L. Wolfe, Elisa Oricchio1, Teresa Palomero and al. (2011).
"Genome-wide RNAi screen identifies miR-19 targets in Notchinduced acute T-cell leukaemia (T-ALL)". Nat Cell Biol. 12 (4): 372–379.
doi:
10.1038/ncb2037.
PMC
2989719.
PMID
20190740.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: numeric names: authors list ( link) - ^ a b Konstantinos J. Mavrakis and Hans-Guido Wendel (2010). "TargetScreen: an unbiased approach to identify functionally important microRNA targets". Cell Cycle. 9 (11): 2080–4. doi: 10.4161/cc.9.11.11807. PMID 20505335.
-
^ Séverine Landais, Sébastien Landry, Philippe Legault and al. (2007). "Oncogenic Potential of the miR-106-363 Cluster and Its Implication in Human T-Cell Leukemia". Cancer Res. 67 (12): 5699–707.
doi:
10.1158/0008-5472.CAN-06-4478.
PMID
17575136.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Johannes Grillari, Matthias Hackl, Regina Grillari-Voglauer (2010).
"miR-17–92 cluster: ups and downs in cancer and aging". Biogerontology. 11 (4): 501–506.
doi:
10.1007/s10522-010-9272-9.
PMC
2899009.
PMID
20437201.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Virginie Olive, Margaux J. Bennett, James C. Walker and al. (2009).
"miR-19 is a key oncogenic component of mir-17-92". Genes Dev. 23 (24): 2839–49.
doi:
10.1101/gad.1861409.
PMC
2800084.
PMID
20008935.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b Flavia Pichiorri, Sung-Suk Suh, Marco Ladetto and al. (2008).
"MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis". Proceedings of the National Academy of Sciences. 105 (35): 12885–90.
doi:
10.1073/pnas.0806202105.
PMC
2529070.
PMID
18728182.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ V Glorian, G Maillot, S Polès and al. (2011).
"HuR-dependent loading of miRNA RISC to the mRNA encoding the Ras-related small GTPase RhoB controls its translation during UV-induced apoptosis". Cell Death and Differentiation. 18 (11): 1692–1701.
doi:
10.1038/cdd.2011.35.
PMC
3190107.
PMID
21527938.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) -
^ Karina Conkrite, Maggie Sundby, Shizuo Mukai and al. (2011).
"miR-17~92 cooperates with RB pathway mutations to promote retinoblastoma". Genes & Development. 25 (16): 1734–45.
doi:
10.1101/gad.17027411.
PMC
3165937.
PMID
21816922.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ a b Joshua T. Mendell (2008). "miRiad roles for the miR-17-92 cluster in development and disease". Cell. 133 (2): 217–22. doi: 10.1016/j.cell.2008.04.001. PMC 2732113. PMID 18423194.
-
^ Karine Tréguer, Eva-Marie Heinrich, Kisho Ohtani and al. (2012). "Role of the MicroRNA-17–92 Cluster in the Endothelial Differentiation of Stem Cells". Journal of Vascular Research. 49 (5): 447–460.
doi:
10.1159/000339429.
PMID
22797777.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link)
- Andrea Ventura, Amanda G. Young, Monte M. Winslow and al. (2008).
"Targeted deletion reveals essential and overlapping functions of the miR-17~92 family of miRNA clustersMechanical stretch up-regulates microRNA-26a and induces human airway smooth muscle hypertrophy by suppressing glycogen synthase kinase-3β". Cell. 132 (5): 875–886.
doi:
10.1016/j.cell.2008.02.019.
PMC
2323338.
PMID
18329372.
{{ cite journal}}: Unknown parameter|DUPLICATE_doi=ignored ( help)CS1 maint: multiple names: authors list ( link) - Lixin Hong, Maoyi Lai, Michelle Chen and al. (2010).
"The miR-17-92 Cluster of microRNAs Confers Tumorigenicity by Inhibiting Oncogene-Induced Senescence". Cancer Res. 70 (21): 8547–8557.
doi:
10.1016/j.cell.2008.02.019.
PMC
2970743.
PMID
20851997.
{{ cite journal}}: Unknown parameter|DUPLICATE_doi=ignored ( help)CS1 maint: multiple names: authors list ( link) - JR-Shiuan Yang, Michael D. Phillips, Doron Betel and al. (2011).
"Widespread regulatory activity of vertebrate microRNA* species". RNA. 17 (2): 312–26.
doi:
10.1016/j.cell.2008.02.019.
PMC
3022280.
PMID
21177881.
{{ cite journal}}: Unknown parameter|DUPLICATE_doi=ignored ( help)CS1 maint: multiple names: authors list ( link) - Joost Kluiver, Johan H. Gibcus, Chris Hettinga and al. (2012).
"Rapid Generation of MicroRNA Sponges for MicroRNA Inhibition". PLOS ONE. 7 (1): e29275.
doi:
10.1371/journal.pone.0029275.
PMC
3253070.
PMID
22238599.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link)