| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 5 | Archive 6 | Archive 7 | Archive 8 | Archive 9 | Archive 10 | → | Archive 15 |
The Chemistry Bond Star
- Thanks and thank you for the fantastic work you are doing on amphetamine and related drug articles. Boghog ( talk) 15:56, 3 January 2014 (UTC)
Disambiguation link notification for January 4
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited C-Met, you added a link pointing to the disambiguation page Clear cell carcinoma ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:59, 4 January 2014 (UTC)
Drug box help: Created a new page
Hi - Yesterday I created a new page on Obeticholic acid which was in the news. I wanted to add a Drugbox (and further chemical details) but couldn't find Help on doing this. Are there template /bots to help with this? The CAS number is 459789-99-2. Thanks for any help. Jrfw51 ( talk) 10:03, 10 January 2014 (UTC)
- Hi. Nice work with the
Obeticholic acid article. Unfortunately there are no bots that I am aware of that can create a filled out {{
infobox drug}} template. I went ahead and manually created it. Cheers.
Boghog (
talk)
14:41, 10 January 2014 (UTC)
- Thanks for creating the Drug infobox. I see you had to reformat many of the references -- and these look better now. Why does the Cite | journal template produce such garbled input? Usually I cut and paste the PMID # but the formating then always needs changing. I see this was discussed above and I would support any extra help on this common feature for us less-than-daily contributors.
Jrfw51 (
talk)
15:31, 10 January 2014 (UTC)
- There are several ways of producing {{ cite journal}} templates. I presume that you were using the RefToolbar. I prefer the User:Diberri's Wikipedia template filling tool ( instructions). While the later requires one extra step (a copy and paste), it produces cleaner and sometimes more complete output. The RefToolbar really needs to be updated, but unfortunately there are no active maintainers of this code. Boghog ( talk) 15:56, 10 January 2014 (UTC)
- Thanks for creating the Drug infobox. I see you had to reformat many of the references -- and these look better now. Why does the Cite | journal template produce such garbled input? Usually I cut and paste the PMID # but the formating then always needs changing. I see this was discussed above and I would support any extra help on this common feature for us less-than-daily contributors.
Jrfw51 (
talk)
15:31, 10 January 2014 (UTC)
Help with citation tool
Hi, Boghog, I work for NCBI, and as discussed here on WP:NIH, I would like to get started working on a way to make it easier for editors to cite PubMed sources. I've read some of your discussions above, and I think you could probably help me scope out the task. I see that you are partial to Diberri's tool, and I'm thinking that maybe that among the things we might do is to host that at NCBI (no promises -- it's just brainstorming at this point). Maybe we could also hook it in with the ref toolbar, although I'm not familiar at all with how that works. I have a bad tendency to try to fix everything at once, and then never get anything done, so if we could scope out something reasonable, I think we could make some real progress. Let me know if you have ideas about this. Klortho ( talk) 07:08, 16 January 2014 (UTC)
Phytoestrogens
Hi Boghog, I don't usually edit pages and don't really know what's customary, but aren't readers of an article like this interested in knowing funders' agendas? I was thinking that people who are concerned about their health would want to know that a study was funded by an organization whose mission is to make sure people keep eating soy. — Preceding unsigned comment added by 75.73.43.96 ( talk) 18:28, 11 January 2014 (UTC)
- Hi. Thanks for your note. I agree who funded the study may have an impact on the reliability of the source. However if there are serious concerns about the source, I think a better course of action is to delete the source and associated text entirely from the article. The particular source in question ( PMID 19524224) appears to be WP:MEDRS compliant (secondary source published in a peer reviewed journal). Editors should not be questioning the reliability of a reliable sources directly in a Wikipedia article unless there is another reliable source to support that doubt. If there is a second reliable source that question the first source, then by all means, include that source in the article. Also:
The text of the article should not needlessly duplicate the names, dates, titles, and other information about the source that you list in the citation.
— WP:MEDMOS
- Details of who funded the study would seem to be excessive. Boghog ( talk) 20:12, 11 January 2014 (UTC)
- Thanks for your response. Now I have a much better idea of how to proceed with this article and how to deal with similar concerns in the future. — Preceding unsigned comment added by 2607:EA00:104:3C00:DC22:EE16:38E0:4E71 ( talk) 18:07, 17 January 2014 (UTC)
Favor to ask
If you're not too busy, could I ask you to look through my reply to Hamiltonstone (the 3 indented bullets) in
Wikipedia:Featured article candidates/Amphetamine/archive1#Boghog) really quickly to make sure I didn't say something stupid in my explanation on the chemistry? I'd appreciate it if you can!
![]() Seppi333 (
Insert 2¢)
17:06, 19 January 2014 (UTC)
Seppi333 (
Insert 2¢)
17:06, 19 January 2014 (UTC)
Hey again Boghog, I'm making a trace amine biosynthesis image that I plan to annotate like template:amphetamine pharmacokinetics, but I have no idea how to construct/label adrenaline consistently with the format I used for the structures - it wasn't included in the papers I used as a reference. Would you be able to give me some guidance on that? Best, Seppi333 ( Insert 2¢) 09:38, 26 January 2014 (UTC)
- Hi Seppi333. Nice work with the biosynthesis graphic. I am not sure exactly what you are asking here. One potential problem is that the R stereochemistry of the benzyl alcohol in norepinephrine is not currently specified in the figure. The absolute stereochemistry of both norepinephrine and epinephrine should be included in the figure. An additional trivial issue is the orientation of the phenyl group. The hydroxyl groups are pointed to the left in the File:Catecholamine and trace amine biosynthesis.png where they are pointed down in File:Norepinephrine structure with descriptor.svg and File:Epinephrine structure.svg. Ideally they should all be drawn in the same orientation. Or are you asking whether to label the structure as adrenaline or epinephrine? (My suggestion is the later since that what the epinephrine article is titled.) Or did you have some other concern? Boghog ( talk) 10:25, 26 January 2014 (UTC)
Ahh alright, I'll update it accordingly. I was confused by the structure difference between the paper I was using as a reference (Fig 2) and wikipedia's articles on the compounds. Seppi333 ( Insert 2¢) 11:03, 26 January 2014 (UTC)
Not done annotating the figure or tweaking the image yet, but did I address the all the chem issues with my changes?
- Yes, looks good. No major issues. There is a minor issue of the bonds not lining up exactly with the labeled atoms to which they are attached. What software are you using to produce these figures? Most chemical drawing software aligns the labeled atoms automatically. Cheers. Boghog ( talk) 17:49, 26 January 2014 (UTC)
- Not that I consider it a chem editor, but Mspaint gets the job done for me, hehe. I actually skewed the side labels slightly to crop it more, but I'll just expand it a little and center them before I use the image in the mainspace (although I just noticed I didn't center NH2 at the top). Seppi333 ( Insert 2¢) 20:54, 26 January 2014 (UTC)
Preferred format for Gene Wiki articles
Last year (13May2003) I heard from a student who was editing the c10orf76 gene page as a project in my bioinformatics analysis course that you would like me to contact you so that I could give my students formatting instructions for the Wikipedia pages on human genes of unknown function that they work on. He suggested that my class (of 44 students) was giving you and others a lot of extra work due to formatting errors in student pages. He forwarded to me some common issues you'd observed, which I'll put into my assignment for this year. If there were other formatting resources you had in mind, please let me know.
Each student wants to make a meaningful contribution to Wikipedia even if it is not a notable one. Our goal is to find sources of information where there are few obvious ones (that is their challenge) so contributing to the page on beta-globin, for instance, wouldn't work.
I look forward to hearing from you. — Preceding unsigned comment added by Wikid25 ( talk • contribs) 20:35, 29 January 2014 (UTC)
- Hi Wikid25. Thank you for contacting me and for adding my formatting requests to your class assignment. To expand on what I previously wrote, there are two problems with creating articles about obscure genes. As I previously mentioned, the first is notability. In order for an article to meet the notability requirement, the subject should have "significant coverage" (i.e., "more than a passing mention") in a reliable source. The second is that there is a high risk that these articles will contain original research, which is not permitted in Wikipedia articles. Straight forward calculations are allowed (see WP:CALC) but synthesis is not. For example, speculation as to function (protein X contains domain Y and domain Y in other proteins has function Z therefore protein X may have function Z) is synthesis. Unless this speculation (protein X may have function Z) is directly support by a reliable source, it should not be include in a Wikipedia article. Unfortunately some of the articles created by your former students are border line in terms of notability and original research. Both of these problems could be avoid if your students were to choose less obscure genes. Boghog ( talk) 04:34, 31 January 2014 (UTC)
few questions again
Hey Boghog, I noticed recently while looking up information on alpha-methylated biogenic amines that there's been some new developments in research on TAARs and their ligands - specifically TAAR5 and TAAR2. I'll probably update the stubs for those articles within the next week or so, but I'm not entirely sure what can be concluded about TAAR2 based upon that paper (its paywalled, but downloadable here). I get that the paper is asserting that TAAR1/TAAR2 are co-expressed in white blood cells and they're "involved" in chemotaxic trafficking of those cells to amines, but does that just suggest or did it actually show that PEA/TYR are hTAAR2 agonists?
- Interesting question. What the paper demonstrates is that TAAR1 and TAAR2 are coexpressed in PMN cells and likely physically interact with each other (e.g., form heterodimers). The paper also demonstrates that trace amines (TAs) are functional in PMN cells (trigger chemotactic migration and cytokine secretion) and through siRNA knockdown experiments, these activities are dependent on the presence of both TAAR1 and TAAR2. TA activity is greatly reduced if either TAAR1 or TAAR2 is knocked down. This suggests that TAAR1/TAAR2 heterodimers are required for full activity. What the paper did not demonstrate is direct binding of TAs to either TAAR1 or TAAR2. It is possible that TAs bind only to TAAR1 or TAAR2 or both partners in the TAAR1/TAAR2 complex. Alternatively TAs could bind to a third protein that signals through the TAAR1/TAAR2 complex. The bottom line is that TAs are probably agonists of TAAR2, but this has not been conclusively proven. What can be said is that TAAR2 is required for full activity of TAs in PMN cells. Boghog ( talk) 07:05, 15 February 2014 (UTC)
My other question is on N-methylhistamine and alpha-methylhistamine. I noticed some databases, like Pubchem,
list the two terms together in the synonyms, but
IUPHAR's HRH4 ligand list contains a mention of N, alpha, and 2 methylhistamine (see agonists section) around 10 times and none are indicated as being endogenouus. Aren't these different enantiomers/racemic forms of the same endogenous compound (the one made by
histamine N-methyltransferase)? Sorry for all the questions; I'm
working on something related that I'm considering on moving to article space at some point.
No rush in responding to this - I know you're busy, so take your time. Regards,
![]() Seppi333 (
Insert 2¢ |
Maintained)
03:56, 13 February 2014 (UTC)
Seppi333 (
Insert 2¢ |
Maintained)
03:56, 13 February 2014 (UTC)
- alpha-Methylhistamine (methyl group attached to the carbon adjacent to the amine group) is definitely not the same molecule as N-Methylhistamine (methyl group attached to the amino group or one of the histidine nitrogen atoms). These two molecules (or actually four molecules since N-methylhistamine is ambiguous) could be described as regio- but definitely not stereoisomers of each other. The Pubchem list should not list N-Methylhistamine as a synonym of alpha-Methylhistamine and therefore is in error. Histamine N-methyltransferase methylates the epsilon histidine nitrogen atom to produce N-Methylhistamine ( Kegg reaction R02155) which is distinct from alpha-methylhistamine. Boghog ( talk) 07:42, 15 February 2014 (UTC)
- Ah, ok that makes sense. Thanks for the explanations - they were really helpful! Best, Seppi333 ( Insert 2¢ | Maintained) 04:03, 16 February 2014 (UTC)
- Sorry to bother you with this again; I need your input in the amph FA again whenever you get a chance - WP:Featured article candidates/Amphetamine/archive2#Boghog2 - just in the 2nd bullet in the synthesis section comments. Seppi333 ( Insert 2¢ | Maintained) 02:21, 19 February 2014 (UTC)
- 2 quick questions - is this correct so far? User:Seppi333/sandbox2
- And I'm showing my chem ignorance by asking this, but what are the 2 remaining reactions for phenylacetone -> benzoic -> hippuric? Thanks for your help with the FAC btw!
Seppi333 (
Insert 2¢ |
Maintained)
10:45, 22 February 2014 (UTC)
- No problem. The graphic so far looks good. Actually what to call the phenylacetone → benzoic transformation is not trivial. The sources that you supply say it is an oxidization, but say nothing about the route. I can find no specifics about the oxidation of phenylacetone in humans. Certain bacteria have a phenylacetone monooxygenase which catalyzes the Baeyer–Villiger oxidation of phenylacetone to benzylacetate. (Warning: what follows is original research) Perhaps humans have a similar enzyme. Once formed benzylacetate can be hydrolyzed to benzyl alchohol which in turn can be oxidized into benzoic acid. Alternatively the benzylic position of phenylacetone can be oxidized to the ketone and the resulting 1,2-diketone could undergo oxidative cleavage to yield benzoic acid (End of OR warning). All you can say is that phenylacetone is (somehow) oxidized into benzoic acid. The second reaction is a conjugation of benzoic acid with glycine to yield hippuric acid (see for example PMID 1811921). I am still working on the expanding the synthesis section for the FAC. Boghog ( talk) 14:19, 22 February 2014 (UTC)
February 2014
Warning, do not remove useful information from articles, like the months of date parameters. See my revert. Debresser ( talk) 16:58, 22 February 2014 (UTC)
- Warning, do not undo constructive edits by other editors. Think before you revert. Boghog ( talk) 18:37, 22 February 2014 (UTC)
- If you make a whole set of edits, it will be your own problem if some of them are not in accordance with Wikipedia guidelines. I reverted to the last version before your removal of citation parameters, and all subsequent edits you will simply have to restore manually. I would like to point out that I seem to remember we have had this discussion before, so you were warned! Debresser ( talk) 19:05, 22 February 2014 (UTC)
- Precisely which of my edits are not in accordance with Wikipedia guidelines? What guideline states that the date parameter must specify the month? In the previous
discussion in which you "warned" me, you simply stated that I was removing useful information which I disputed and you never gave an adequate reply. This
revert with the edit summary
Perhaps this will teach him a lesson
is incredibly arrogant. Editors are supposed to resolve conflicts through discussion, not scolding. Boghog ( talk) 19:35, 22 February 2014 (UTC)
- Precisely which of my edits are not in accordance with Wikipedia guidelines? What guideline states that the date parameter must specify the month? In the previous
discussion in which you "warned" me, you simply stated that I was removing useful information which I disputed and you never gave an adequate reply. This
revert with the edit summary
- The phrase you quote was only part of my edit summary, and the whole edit summary makes better sense than you credit me for.
- Please also post the same "edit warring" warning you were so kind to paste on my talk page on your own talk page, since it is you who is the second party to this "edit war".
- The claim "you never answered my question", or as you say here "you never gave an adequate reply" is the stereotype reply of a person who does not listen to the objective arguments that have been brought forth. I have explained myself adequately, both this time and in the past. Do not remove month parameters from citation templates! Debresser ( talk) 20:55, 22 February 2014 (UTC)
- No, you have not explained yourself adequately and you have still not answered my question. What guideline states that the date parameter must specify the month? I am still waiting for an answer. Boghog ( talk) 21:10, 22 February 2014 (UTC)
- You are asking the wrong question. The question you should ask is "why was I reverted"? And the answer is, because you removed helpful information from the article, namely the months from the citation parameters. There is no rule that these must be here, as there is no rule that any information whatsoever must be in an article. But once it is there, nobody should remove it, as in Wikipedia, we do not remove helpful information without a good reason. That is called "vandalism", among other terms. Even though in this case I understand it was not vandalism motivating you, still, the act of removing information from an article is detrimental to the project and will not be tolerated. Debresser ( talk) 21:47, 22 February 2014 (UTC)
- Thank you for your detailed reply. To reiterate, month is redundant with issue in scientific citations and all the citations in the article in question are scientific. Removing redundant information is not necessarily detrimental to the project. Boghog ( talk) 22:11, 22 February 2014 (UTC)
- I see you point, even though I disagree with it. What do you say we ask at Help talk:Citation Style 1 (which is where the talk page of Cite:Journal redirects to)? Debresser ( talk) 21:12, 23 February 2014 (UTC)
Amphetamine Problems
Hey Boghog. Seppi is being pretty aggressive again, reverting edits solely because I made them. He chopped the entire toxicity section down to two short sentences (which is far too short for the nuanced topic) just to piss me off. He's also strangely, subtly vandalizing a talk page section I added. I correctly guessed the reason for his (admittedly mild) POV, which can be seen in a section of my talk page. He seems to have gone off the deep end, though, and we may need mediation. Exercisephys ( talk) 00:51, 27 February 2014 (UTC)
- I have not been following this too closely, but I have noticed some friction. This is a bit outside my area of expertise, but I will take a look and try to offer some suggestions that might help resolve the dispute. Boghog ( talk) 08:57, 27 February 2014 (UTC)
Hi, can you check the above article - a complete section has been blanked - is it ok? Regards Denisarona ( talk) 16:10, 5 March 2014 (UTC)
- If you have time, maybe also look at Amphiregulin - I don't know much about these subjects. Denisarona ( talk) 16:12, 5 March 2014 (UTC)
- Hi. Thanks for the heads up. I have mixed feelings about these edits. The intention of the further reading sections in bot generated Gene Wiki articles was to provide notability and encourage human editors to expand the articles by providing background references some of which would hopefully be moved in-line. The closest relevant guideline I could find is this:
Some editors list sources that they hope to use in the future to build the article in Further reading. This is neither encouraged nor prohibited.
— Wikipedia:Further_reading#Relation_to_reference_sections
- I myself occasionally remove these further reading sections, especially if there are large numbers of in-line citations. These particular deletions are borderline. These articles have undergone minor expansion and only contain a few in-line citations. In addition, the edit summary Cite in footnote if important for the article ( diff) is a bit condescending. If this editor only deletes a few of these sections, I would tend to ignore it. However if this editor continues these further reading section deletions on large numbs of articles, particularly if the articles have few in-line citations, I would strongly oppose it. 16:52, 5 March 2014 (UTC) Boghog ( talk)
Some stroopwafels for you!

|
For generally continuous high-level editing of articles in the medical and pharmaceutical science area. Keep it up! JFW | T@lk 17:50, 9 March 2014 (UTC) |
- Thanks for the stroopwafels! Have had tasted the delicious cheese from the same region but never before the wafels. Yum, yum. I have always thought MEDRS was a bit over the top, but I am starting to appreciate why MEDRS is necessary. Boghog ( talk) 18:48, 9 March 2014 (UTC)
- Oi, I like these so much too! Debresser ( talk) 19:05, 9 March 2014 (UTC)
Autofixing CS1 cites
Hello, User:Wikid77 here. I noticed you have been discussing ways to correct cite parameters. I am currently writing extra Lua script to autofix (auto-correct) typos when using the wp:CS1 cite templates. When I helped to develop those Lua-based cite templates, during October 2012 to April 2013, the intent was to auto-correct for typos, not issue numerous error messages, and I never imagined Lua would be used to issue thousands of red-error messages when simple auto-correction would have been quite easy. Well, after waiting all year, I have returned to re-focus on autofixing typos in cite templates, and suppress most of the red-error messages. Across Wikipedia editing, many editors are just too busy to nitpick the details and so, autofixing of cite parameters provides a rapid way to solve the problems and make many cite templates almost trivial to use. Last year, I estimated the hand-correction of cites to require over 3 years of manual, hand-crafted edits, and now after another whole year, the backlog is still about 3-5 years if hand-fixed. Although several users are diligently hand-editing the pages to fix cites, many other users are actively inserting invalid cite parameters into almost as many dozens of pages each week. The past year (of tedious cite work) has proven how autofixing is the only hope to rapidly correct the 10,000 pages in the backlog categories. For example:
During early 2014, the unsupported parameters have been fixed at only 100-200 pages per month, as meaning more years of backlog work. I wrote new essay, " wp:Autofixing cites" to explain some simple ways to autofix the major cite parameters and hope people might discuss issues about the autofixing in the talk-page there, " WT:Autofixing cites" where all the complex tactics of fixing URLs and dates could be discussed, in more detail. - Wikid77 ( talk) 21:48, 12 March 2014 (UTC)
Bot
Hey Boghog. Would you be interested in making this bot [1]? Would be happy to send you the reward :-) Doc James ( talk · contribs · email) (if I write on your page reply on mine) 06:33, 15 March 2014 (UTC)
- By the way also agree that it would be nice to move refs over one line rather than many lines. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 11:17, 15 March 2014 (UTC)
Okay it appears we may have consensus. Do you want to start the bot request or should I? What we proposing to do, if I understand correctly is:
- Replacing "cite PMID" and "cite DOI" with "cite journal"
- Replacing "cite ISBN" with "cite book"
- Placing all refs that occur over many lines to over one line
- Probably not a good idea to address the author issue yet
Doc James ( talk · contribs · email) (if I write on your page reply on mine) 00:17, 16 March 2014 (UTC)
- I recommend that you advertise more widely before deciding that you have consensus to make a substantive change that would affect tens of thousands of pages. –
Jonesey95 (
talk)
01:30, 16 March 2014 (UTC)
- The current proposal is to substitute {{
cite pmid}} with equivalent {{
cite journal}} templates, not in tens of thousands of pages, but rather in the "1500 most viewed articles" that are within the scope of
WikiProject Medicine. The proper venue for this discussion is therefore the project's talk page and the
consensus established there is overwhelmingly in favor of this change.
Boghog (
talk)
05:58, 16 March 2014 (UTC)
- Yes there is consensus among the editors who edit this content most. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 08:40, 16 March 2014 (UTC)
- The current proposal is to substitute {{
cite pmid}} with equivalent {{
cite journal}} templates, not in tens of thousands of pages, but rather in the "1500 most viewed articles" that are within the scope of
WikiProject Medicine. The proper venue for this discussion is therefore the project's talk page and the
consensus established there is overwhelmingly in favor of this change.
Boghog (
talk)
05:58, 16 March 2014 (UTC)
Looks like there is agreement for both. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 06:07, 20 March 2014 (UTC)
Is the the citation-template-filling tool on wmflabs working for you?
I get it trying to save index.cgi rather than running the script. Does it need to be authorized again or something? RDBrown ( talk) 12:26, 24 March 2014 (UTC)
I am not quite sure what you mean by "trying to save index.cgi". I just got the citation template filling tool working today by following these instructions, and in particular this solution. Boghog ( talk) 18:53, 24 March 2014 (UTC)
Template data
Is the data stored in a db for citePMID and citeISBN? Or is it a bunch of templates that somehow get polled by the individual articles when they use the cite-call? Ian Furst ( talk) 18:06, 17 March 2014 (UTC)
- I am not sure that I completely understand your question. Which data are you referring to? Currently Wikidata does not store citation data. It is planned for the future, but it has not yet been implemented. There are of course other sources of citation data. For example,
PubMed is a database that stores citations that can be retrieved using a
PMID (e.g.,
PMID
12345). There are several book databases that can be searched by
ISBN and return citation data (e.g.,
0-321-73975-2). Finally there are web tools that will return citation data given a
doi (e.g.,
CrossRef). The long term plan is to capture citation data stored in the individual Wikipedias in Wikidata and check that data against each other and against external databases. Again, I am not exactly sure how this will be implemented since it still is in the planning stage.
Boghog (
talk)
18:34, 17 March 2014 (UTC)
- I think I'm asking the question the wrong way. I've been using citeISBN in some articles. If I place citeISBN into articleA, the data is stored on another page with a title that includes that ISBN number (same with PMID, I think). Is that data just being stored on the template page or does it get entered into a relational database?
Ian Furst (
talk)
18:40, 17 March 2014 (UTC)
- Oh sorry, now I understand. The {{
cite pmid}} and {{
cite isbn}} templates stores the citation data in another template that is
transcluded back into the Wikipedia article. For example, {{cite pmid|10367338}} produces the following citation:
- Collins, A. S.; Sumner, S. C.; Borghoff, S. J.; Medinsky, M. A. (1999). "A physiological model for tert-amyl methyl ether and tert-amyl alcohol: Hypothesis testing of model structures". Toxicological sciences : an official journal of the Society of Toxicology. 49 (1): 15–28. doi: 10.1093/toxsci/49.1.15. PMID 10367338.
- The data is not stored on my talk page. If you click on the small "edit" button at the end of the citation above, it will take you to the template where the data is actually stored.
Boghog (
talk)
18:54, 17 March 2014 (UTC)
- That part I understand. Where I'm unsure is how the article {{cite pmid|}} function calls the data. Are the individual fields (name, title, etc...) that were entered into the template part of a larger backend db (e.g. is the template just a form that I'm seeing and the data is actually stored as individual fields behind the scenes and the cite function is a query. Or does the {{cite pmid|}} trigger a bot that searches the necessary fields. I'm guess I'm trying to figure out how the backend of the cite function works.
Ian Furst (
talk)
19:01, 17 March 2014 (UTC)
- Interesting question. I believe there is no database as such. Rather the data is pulled in from the template on-the-fly as needed. After the data is read from the template, it is inserted directly into the article as if it were there from the beginning. Of course there is a performance penalty for doing this. To minimize the delay in rendering the page, Wikipedia caches the fully composed Web page using Squid (software) so that it can quickly be displayed. Boghog ( talk) 19:15, 17 March 2014 (UTC)
- That part I understand. Where I'm unsure is how the article {{cite pmid|}} function calls the data. Are the individual fields (name, title, etc...) that were entered into the template part of a larger backend db (e.g. is the template just a form that I'm seeing and the data is actually stored as individual fields behind the scenes and the cite function is a query. Or does the {{cite pmid|}} trigger a bot that searches the necessary fields. I'm guess I'm trying to figure out how the backend of the cite function works.
Ian Furst (
talk)
19:01, 17 March 2014 (UTC)
- Oh sorry, now I understand. The {{
cite pmid}} and {{
cite isbn}} templates stores the citation data in another template that is
transcluded back into the Wikipedia article. For example, {{cite pmid|10367338}} produces the following citation:
- I think I'm asking the question the wrong way. I've been using citeISBN in some articles. If I place citeISBN into articleA, the data is stored on another page with a title that includes that ISBN number (same with PMID, I think). Is that data just being stored on the template page or does it get entered into a relational database?
Ian Furst (
talk)
18:40, 17 March 2014 (UTC)
Found my way to some documentation on it and you're right (sorry - should have looked for the solution myself, thought it would be a simple question). It says that it's transcluded from the template. I've done lots of db work but never used a page to act as a record source like this. My suspicion is that the parameter fields in the template are easier to screw up than a source field in a db (not to mention the performance hit). I wonder what causes problems across languages? Also, if a wiki solution is appropriate for data like this with discrete fields. It might be better managed in a db. 19:29, 17 March 2014 (UTC)
- Boghog; I was thinking more about the problems with citations and their lack of portability. My assumption is if a citations template is on a particular languages wiki it's not easily transportable or accessible from another languages wiki. What do you think about beginning a discussion on the hosting of the citation data in a database (that would be accessible to all language wiki's). A particular citation then would have a single uniqueID (PMID/ISBN/DOI or whatever) which would allow enough fields to satisfy all languages. E.g. a PMID entry might be made once in english with the field "article_name_english" but could then be translated to all 248 languages (and the same with other fields). Because it's indexable the call from the article to populate the data would be quick. Hopefully, entry is done once and the translation automated.
Ian Furst (
talk)
23:51, 17 March 2014 (UTC)
- I am not certain how Wikidata works under the hood, but I believe Wikidata is essentially a database. Furthermore Wikidata already contains language specific fields (see for example Q5633421). So in principle, Wikidata could store translated citation titles and journal names for each language. In practice, I am not sure how this would work. Perhaps Google translate could be used to provide first pass translations that could be tweaked by human editors if necessary. In any case, before support for citations in Wikidata is rolled out, these issues will need to be taken into account. See Wikidata road map for current status and future plans. Since citations are not yet on the road map, I think it will be some time before citations are added to Wikidata. Boghog ( talk) 07:56, 26 March 2014 (UTC)
Metabolism of amph (again)
Hey Boghog, I found this information of a toxicity data site: http://www.acutetox.eu/ (PDF: http://www.acutetox.eu/pdf_human_short/35-Amphetamine%20revised.pdf)
Metabolism and excretion
Metabolism of amphetamine generally includes aromatic hydroxylation, aliphatic hydroxylation, and n-dealkylation, resulting in inactive and active metabolites.
Main metabolites of amphetamine are: hippuric acid (16 to 28%), phenylacetone (0.9%), benzoylglucuronide (4%), norephedrine (2%), conjugated p-hydroxyamphetamine (2-4%) and conjugated p-hydroxynorephedrine (0.3%). One of metabolites, p-hydroxyamphetamine is a potent hallucinogen implicated in the development of amphetamine-induced psychosis [1].
The excretion of unchanged amphetamine is dependent on the urine pH. In an acidic urine (pH less than 6.6) 67 to 73% of an ingested dose is excreted as amphetamine. When pH is higher than 6.7, 17 to 43% is excreted unchanged in the urine [1].
Do you think its worth updating the metabolic pathway diagram to include "conjugated p-hydroxyamphetamine"?
Also, what program do you use to generate SVG structure diagrams? I think I may end up redoing the ones I made in that format.
Best, Seppi333 ( Insert 2¢ | Maintained) 02:35, 26 March 2014 (UTC)
- PS: Your synthesis diagrams look really nice.
- Thanks. I am not sure that it really necessary to add conjugation since it is a relatively minor metabolic pathway for amphetamine and the source you supplied does not specify what kind of conjugation is taking place (I assume it is glucuronidation). I use ChemDraw to produce the graphics, save as pdf and use pdf2svg to convert the pdf to svg format. Boghog ( talk) 12:23, 26 March 2014 (UTC)
Please tell why you reverted my paragraph
On the myostatin article. The article was from Nasdaq.com. If youre going to revert someone's edit, please tell why.-- Wyn.junior ( talk) 15:04, 28 March 2014 (UTC)
- Sorry, I should have left an edit summary. The source you have supplied just states that MYO-T12 is just entering clinical trials. Furthermore it is claimed that this dietary supplement somehow reduces myostatin levels. There are no reliable sources that support this claim. Just press releases by the company that contain the disclaimer "These statements have not been evaluated by the Food and Drug Administration. This product is not intended to diagnose, treat, cure or prevent any disease". Per WP:MEDRS, what is need to support medical claims are reliable secondary sources that review the results of control clinical trials.
- Elsewhere in the myostatin article, it states:
- "Some athletes, eager to get their hands on such drugs, turn to the internet, where fake "myostatin blockers" are being sold.". MYO-T12 is being sold a dietary supplement, but without a shred of evidence that is actually increases muscle mass. Until it is proven that it works, MYO-T12 would appear to be just another fake "myostatin blocker". Boghog ( talk) 15:30, 28 March 2014 (UTC)
- The products on the market are all pill based. Is it possible that MYO-T12 is an injection?-- Wyn.junior ( talk) 00:10, 29 March 2014 (UTC)
- I don't think it is likely that MYO-T12 could be injected. One of the main ingredients is egg yoke powder which would case a severe immune reaction if injected directly into the blood stream without first being digested. Follistatin from chicken egg yoke is claimed to be the active ingredient of MYO-T12. One might be able to inject purified chicken follistatin, but I would be very worried about an immune response. Recombinant human follistatin would be safer, but would be expensive to produce.
- PMID 24627466 has demonstrated that native follistatin injected systematically has little effect on muscle regeneration because of short half-life. A re-engineered follistatin with longer half-life had a much greater effect. In summary, MYO-T12 taken orally would not work because the follistatin it contains would be digested like any protein before it reaches the blood stream. Injected re-engineered follistatin may work, but this is not what MYO-T12 is. Bottom line, don't waste your money. This stuff cannot possibly work. Boghog ( talk) 07:48, 29 March 2014 (UTC)
- Also the text that you introduced is internally inconsistent. How could the statements "MYOS Corp is conducting its first clinical trial" and "the world's first clinically proven natural myostatin inhibitor" both be true? Boghog ( talk) 15:39, 28 March 2014 (UTC)
citation-template-filling tool
Editors using
this tool are creating references containing the
deprecated |month= parameter (e.g. in PMID, maybe others).
Are there any plans to amend the output to instead use the |date= parameter? This parameter can take a variety of complete and partial dates. --
79.67.241.76 (
talk)
18:28, 28 March 2014 (UTC)
- Thanks for your message. No one has yet to provide me with a clear explanation for why it was necessary to deprecate this parameter. But yes, I was planning to take care of this. My preference is to skip the month and return only the year in the year parameter since (1) journals themselves rarely include month in reference sections of articles, (2) month is redundant with issue, and (3) it is cleaner and avoids date formatting issues. To date, the only thing have have done with the code is to port it to the new tool server. So it may take a bit of digging to figure out how to do this, but I am working on it. Boghog ( talk) 21:52, 28 March 2014 (UTC)
- Untested, but I think this should change month, year to date. I was doubtful about month, until I checked the IEEE journals I have. Month is given at the top of pages, not issue so month may help in finding articles in bound journals ... if I'm not showing my age.
--- WWW/Wikipedia/TemplateFiller/Source/PubmedId.pm.orig 2009-05-15 11:48:47.000000000 +1000
+++ WWW/Wikipedia/TemplateFiller/Source/PubmedId.pm 2014-04-07 22:10:10.203585151 +1000
@@ -79,6 +79,10 @@ sub template_basic_fields {
my $month = Decode_Month( $self->{month} ) if $self->{month};
$month = Month_to_Text( $month ) if $month;
+ if (defined($month) && defined($self->{year})) {
+ $month = $month . ' ' . $self->{year};
+ $self->{year} = undef;
+ }
tie( my %fields, 'Tie::IxHash' );
%fields = (
@@ -89,8 +93,8 @@ sub template_basic_fields {
volume => { value => $self->{volume} },
issue => { value => $self->{issue} },
pages => { value => $pages },
- year => { value => $self->{year} },
- month => { value => $month, show => 'if-filled' },
+ year => { value => $self->{year}, show => 'if-filled' },
+ date => { value => $month, show => 'if-filled' },
pmid => { value => $self->{pmid} },
pmc => { value => $self->{pmc_id}, show => 'if-filled' },
doi => { value => $self->{doi} },
- Hope this helps/works. RDBrown ( talk) 13:12, 7 April 2014 (UTC)
- For existing references, Monkbot currently checks
|day=|month=|year=(or|month=|year=) are valid and then merges them into|date=. If month data is available in your tool it may be better to return both month and year in|date=. I don't think there's any format other than|date=March 2014available for this. When day is also present, that's when the fun begins! My preference is 28 March 2014 for publication date (e.g. newspapers), and 2014-03-28 for archivedate. Likewise the latter for accessdate, but only when the publication date is not available. -- 79.67.241.76 ( talk) 22:48, 28 March 2014 (UTC)
- For existing references, Monkbot currently checks
- Very specific reasons as to why it is better to return the year in the year parameter and skip the month were presented above. In contrast you merely state that it "may be better to return both month and year in
|date=". Also IMHO, including the day on which an article was published is excessive and completely unnecessary. Finally it makes no sense to include an accessdate in a journal citation. Articles that are published in journals must by definition have been published on a specific date. If an article does not have a publication date (e.g., year + volume + issue), it wasn't published in a journal. Boghog ( talk) 23:28, 28 March 2014 (UTC)
- Very specific reasons as to why it is better to return the year in the year parameter and skip the month were presented above. In contrast you merely state that it "may be better to return both month and year in
- Totally agree about
|accessdate=. Many other editor tools blindly shove accessdate into every reference. Whenever|date=is present I remove accessdate. As for full dates, I agree they are not needed in journal references. I should have clarified my latter comments about Monkbot and dates were meant to be more general, i.e. used for newspaper references and other such things, and not specifically to do with journals. Glad to hear you're considering updating the tools at some point. -- 79.67.241.76 ( talk) 23:53, 28 March 2014 (UTC)
- Totally agree about
Mistake
Sorry, I made a mistake deleting this paragraph. Thank you for correcting it. Humpath ( talk) 09:15, 11 April 2014 (UTC)
- No worries. Cheers. Boghog ( talk) 10:07, 11 April 2014 (UTC)
Permutation of cite journal authors
Hi! In this Boghog edit of the Bipolar disorder page, https://en.wikipedia.org/?title=Bipolar_disorder&curid=4531&diff=603896185&oldid=603893399 Bipolar disorder (Latest revision as of 11:38, 12 April 2014) the bot just permuted the author list of a "cite journal." It also didn't put them in numeric order so it seems a little pointless.
The request for approval appears to say that it only would affect 'cite doi' 'cite pmid' and 'cite isbn.' Is the Bipolar disorder change an accident or mistake? SesquiZed ( talk) 02:31, 13 April 2014 (UTC)
- Sorry.... I meant Bogbot 5 edit not Boghog edit ( /info/en/?search=Wikipedia:Bots/Requests_for_approval/BogBot_5) SesquiZed ( talk) 02:38, 13 April 2014 (UTC)
- Thanks for your note. That was an error. There was part of the bot script that was supposed to be a second pass over the substituted cite templates, not all of the templates. The error in the bot script has now been fixed. Boghog ( talk) 07:31, 13 April 2014 (UTC)
Bioinformatic analysis course Wikipedia pages on human genes of unknown function
Hi Boghog. I've tried to contact you before but don't see a reply here -- so maybe didn't get through. I'm the professor at U of M who has students write Wikipedia articles on little-known genes and have heard through my students that we should talk. I've given a formatting guide to my students based on what you sent a few of them last year but if there is more I can do to make things easier on your end, please let me know. My userID is Wikid25 and though I know you've made clear that I should come back here for a response to a question asked here, would you consider pasting any answer you might make at my talk page this once? Thanks! By the way, the following is what I give students based on what I've heard from you before:
• Please check out User:Diberri's Wikipedia template filling tool (instructions). Given a PubMed ID, one can quickly produce a full citation that can be copied and pasted into a Wikipedia article. This tool can save you a lot of work and ensure that the citations are displayed in a consistent manner. • For consistency with other Gene Wiki articles, the An Error has occurred retrieving Wikidata item for infobox infobox should normally not be included directly in the article but rather transcluded using the template. User:ProteinBoxBot is setup to maintain transcluded templates, but not templates directly contained in articles. The GeneWikiGenerator will automatically create both the template and the article that transcludes the template. • Concerning the lead sentence, as discussed here and here, we have tried to make clear that these Gene Wiki articles are not only about the human gene/protein, but also orthologs that exist in other species. The wording that was reached through consensus is perhaps a little awkward, but it is both accurate and concise: • <recommended UniProt name> is a protein that in humans is encoded by the <approved HUGO gene symbol> gene. The "that" in the above sentence is non-limiting implying that the protein (and gene) exists in other species besides human. • Per WP:HEAD, sentence case should be used for section headings. • Per WP:REFPUNC, punctuation before citations.
- Hi @ Wikid25: Thanks for your note. I did reply, but my reply has now been archived but still can be read here. If your students follow the recommendations above, that would be a great help. However, as stated in my previous reply, I the bigger concern I have is about the notability of many of these obscure genes whose function is not know. Boghog ( talk) 14:44, 13 April 2014 (UTC)
Thanks. I'll shift all future projects to genes/proteins that have at least one article in which they gene is the primary subject of the article. I'm thinking of the situation of a recently characterized gene such as SARAF/TMEM66. I will also make it clear to students that synthesis and other forms of original research are not permitted. I have discussed at length with them how all their sources must be provided such that any reasonably informed person could verify claims made. Hopefully this addresses the majority of the issues that have created work or concern for you in your editing role in the past years. — Preceding unsigned comment added by Wikid25 ( talk • contribs) 15:43, 13 April 2014 (UTC)
- Perfect! That should take care most of my concerns. I look forward to seeing your students contributions. Cheers. Boghog ( talk) 04:38, 14 April 2014 (UTC)
Disambiguation link notification for April 14
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited FGF4, you added a link pointing to the disambiguation page SHH ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:53, 14 April 2014 (UTC)
SETDB1
Hello,
About the article on Setdb1: http://en.wikipedia.org/wiki/SETDB1
"Model organisms have been used in the study of SETDB1 function" I found no article using this knock out mice.
The "abnormalities" have never been investigated and you can't conclude that there is a lymphocyte or bone issue in these mice on only one test carried out. Furthermore, this have never been reported in the literature yet. Saying that SETDB1 is implicated in bone strength and lymphocyte biology only according to the data obtained from a pre-phenotyping of a KO mice is (to my opinion) non scientifical comments. The links provided on this subject all leads to the "Wellcome Trust Sanger Institute" (that's half of the citation). It looks like more an advertisement from Eucomm that is selling the mice.
This article should be enriched with the already known roles of SETDB1 in the literature and not be an excuse reason for private companies to get a free advertisement.
I don't doubt that animal models are being widely used in research as I am a researcher myself. — Preceding unsigned comment added by 195.221.129.220 ( talk) 08:28, 16 April 2014 (UTC)
- The main problem is with your original
edit summary that stated "Deleted non scientifical comments". Edit summaries do not need to be long, but if you are deleted sourced material, you need to provide more rationale for your deletion. Even your more lengthy explanation above is problematic:
- "I found no article using this knock out mice" – see SETDB1 knockout
- "not be an excuse reason for private companies to get a free advertisement" – neither the Sanger Institute not the EUCOMM is a private company, both are a non-profit organizations.
- On the other hand, the citations provided were not specific to this gene. Hence I would agree that the deleted section as written is not appropriate for this article. Boghog ( talk) 17:58, 16 April 2014 (UTC)
Rictor page
Boghog:
Just wanted to let you know that I am working on the Rictor page for a class assignment. I would really appreciate it if you would hold off on your edits (especially the stylistic ones) until I submit this assignment for grading.
Thanks for your interest in the page and your current contributions.
kldextraze — Preceding unsigned comment added by Kldextraze ( talk • contribs) 16:43, 17 April 2014 (UTC)
WikiProject Genetics in the Signpost
The WikiProject Report would like to focus on WikiProject Genetics for a Signpost article. This is an excellent opportunity to draw attention to your efforts and attract new members to the project. Would you be willing to participate in an interview? If so, here are the questions for the interview. Just add your response below each question and feel free to skip any questions that you don't feel comfortable answering. Multiple editors will have an opportunity to respond to the interview questions, so be sure to sign your answers. If you know anyone else who would like to participate in the interview, please share this with them. Have a great day. –Mabeenot ( talk) 18:53, 19 April 2014 (UTC)
Bot for ref formatting
Where are we at? Doc James ( talk · contribs · email) (if I write on your page reply on mine) 03:31, 21 April 2014 (UTC)
- I just noticed the bot transcluded cite templates. Thanks. Blue Rasberry (talk) 19:07, 21 April 2014 (UTC)
A barnstar for you!

|
The Technical Barnstar |
| For fixing the refs you have :-) Looking forwards to the larger run. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 19:37, 21 April 2014 (UTC) |
- By the way you rock! So looking forwards to not having to fix these by hand :-) Doc James ( talk · contribs · email) (if I write on your page reply on mine) 19:37, 21 April 2014 (UTC)
Thanks, but I am not quite done. I should be able to finish this soon after I fix a few bugs in the bot script. BTW, it turns out that there are only about ~300 of the 1500 MED articles that have transcluded cite pmid/doi/isbn templates. Boghog ( talk) 19:42, 21 April 2014 (UTC)
- Ah good to know. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 20:09, 21 April 2014 (UTC)
Refs for preg cat
IMO we should have refereces for the preg cat listed in the info boxes. Could we get a bot to add these? Doc James ( talk · contribs · email) (if I write on your page reply on mine) 09:22, 24 April 2014 (UTC)
- Sure, adding references using a bot is no problem. But where do the references come from? Is there a generic reference to some database that we can use? Or are there drug specific references?
Boghog (
talk)
10:37, 24 April 2014 (UTC)
- So this lists the USA
http://www.drugs.com/monograph/isosorbide-dinitrate-mononitrate.html and this lists the Australian
http://www.tga.gov.au/hp/medicines-pregnancy.htm#.U1jqHRc3tqx
Doc James (
talk ·
contribs ·
email) (if I write on your page reply on mine)
10:54, 24 April 2014 (UTC)
- Could we format the monograph as <ref name=AHFS2014>{{cite web|title=Cephalexin|url=http://www.drugs.com/monograph/cephalexin.html|publisher=The American Society of Health-System Pharmacists|accessdate=Apr 21, 2014}}</ref> as I use it frequently for other stuff. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 10:55, 24 April 2014 (UTC)
- So this lists the USA
http://www.drugs.com/monograph/isosorbide-dinitrate-mononitrate.html and this lists the Australian
http://www.tga.gov.au/hp/medicines-pregnancy.htm#.U1jqHRc3tqx
Doc James (
talk ·
contribs ·
email) (if I write on your page reply on mine)
10:54, 24 April 2014 (UTC)
Invitation join the new Physiology Wikiproject!

Based on the long felt gap for categorization and improvization of WP:MED articles relating to the field of physiology, the new WikiProject Physiology has been created. WikiProject Physiology is still in its infancy and needs your help. On behalf of a group of editors striving to improve the quality of physiology articles here on Wikipedia, I would like to invite you to come on board and participate in the betterment of physiology related articles. Help us to jumpstart this WikiProject.
- Feel free to leave us a message at any time on the WikiProkect Physiology talk page. If you are interested in joining the project yourself, there is a participant list where you can sign up. Please leave a message on the talk page if you have any problems, suggestions, would like review of an article, need suggestions for articles to edit, or would like some collaboration when editing!
- You can tag the talk pages of relevant articles with {{WikiProject Physiology|class=|importance=}} with your assessment of the article class and importance alongwith. Please note that WP:Physiology, WP:Physio, WP:Phy can be used interchangeably.
- You will make a big difference to the quality of information by adding reliable sources. Sourcing physiology articles is essential and makes a big difference to the quality of articles. And, while you're at it, why not use a book to source information, which can source multiple articles at once!
- We try and use a standard way of arranging the content in each article. That layout is here. These headings let us have a standard way of presenting the information in anatomical articles, indicate what information may have been forgotten, and save angst when trying to decide how to organise an article. That said, this might not suit every article. If in doubt, be bold!
- Why not try and strive to create a good article! Physiology related articles are often small in scope, have available sources, and only a limited amount of research available that is readily presentable!
- Your contributions to the WikiProject page, related categories and templates is also welcome.
- To invite other editors to this WikiProject, copy and past this template (with the signature):
{{ subst:WP Physiology–invite}}~~~~
- To welcome editors of physiology articles, copy and past this template (with the signature):
{{ subst:WP Physiology–welcome}}~~~~
- You can feel free to contact us on the WikiProkect Physiology talk page if you have any problems, or wish to join us. You can also put your suggestions there and discuss the scope of participation.
Hoping for your cooperation! Diptanshu Talk 12:27, 27 April 2014 (UTC)
EDITS
It's my understanding that edit disagreements are best discussed on the talk page of the site where the edits are occurring. Are you OK with that approach? If so I'll leave you a note on the Ghrelin talk page for your response. Thanks.
Regards -
IiKkEe ( talk) 05:57, 5 May 2014 (UTC)
- Hi. Thanks for your message and for the impressive work you have done on the Ghrelin and related articles. In answer to your question, it is generally best to have article specific discussions on the article's talk page. That keeps the discussion in one place so that it easier to follow. Cheers. Boghog ( talk) 16:13, 5 May 2014 (UTC)
Edit to Tramadol
This edit appears to use some kind of automation to "fix" citations. However, it does not follow Help: Citation Style 1 (CS1). At least one difference is that CS1 separates authors with a semicolon, while you separated them with a comma. Please repair your automation before using it again. Jc3s5h ( talk) 14:20, 8 May 2014 (UTC)
- My edit does follow {{ cite journal}} (see Vancouver system). Also this was a manual edit that included a lot of other fixes. Boghog ( talk) 14:26, 8 May 2014 (UTC)
I noticed another error; authors who apparently have western European names have their names given in the format "Doe J" when it should be "Doe, J". Jc3s5h ( talk) 14:30, 8 May 2014 (UTC)
- That is not an error. That is intentional (see Vancouver system). Boghog ( talk) 14:31, 8 May 2014 (UTC)
- No, I will not look at Vancouver system. The article does not use Vancouver system. Consider this citation:
- Nelson, G.K.; Lombardi, M.A.; Okayama, D.T. (2005).
"NIST Time and Frequency Radio Stations: WWV, WWVH, and WWVB" (PDF).
National Institute of Standards and Technology. (Special Publication 250-67).
{{ cite web}}: Invalid|ref=harv( help) dead link
- Nelson, G.K.; Lombardi, M.A.; Okayama, D.T. (2005).
"NIST Time and Frequency Radio Stations: WWV, WWVH, and WWVB" (PDF).
National Institute of Standards and Technology. (Special Publication 250-67).
- No, I will not look at Vancouver system. The article does not use Vancouver system. Consider this citation:
- The authors are specified with the perfectly valid syntax
"| last1=Nelson |first1=G.K. |last2=Lombardi |first2=M.A. |last3=Okayama |first3=D.T."The results are last names separated from first names with commas, and authors separated by semicolons. You must follow this format or you are in violation of WP:CITEVAR. Jc3s5h ( talk) 14:37, 8 May 2014 (UTC)
- The authors are specified with the perfectly valid syntax
- That is not what WP:CITEVAR says. Please re-read it. The predominate style was initially the Vancouver system and I am restoring it. Boghog ( talk) 14:40, 8 May 2014 (UTC)
- Help:Citation Style 1 sets forth the style for templates in the Citaion Style 1 family (which includes {{ cite journal}}. If you believe the article uses Vancouver style, ask on the talk page of the article to change the templates to Help:Citation Style Vancouver. It is not reasonable to use templates in a way that legal variations on ways of expressing parameters result in different rendered citations.
- If you insist on this idiosyncratic interpretation of WP:CITEVAR I will bring this to the discussion about your new bot; we can't have bot operators writing citation-related bots if their ideas about citation rules are different from everyone else's. Jc3s5h ( talk) 14:49, 8 May 2014 (UTC)
Consider this
older version of the article which show a clear preference for the
Vancouver system. What
WP:CITEVAR says is defer to the style used by the first major contributor
. Your revert of my edit is in clear violation of
WP:CITEVAR.
Boghog (
talk)
14:51, 8 May 2014 (UTC)
Also I repeat, this was not a bot edit. Boghog ( talk) 14:52, 8 May 2014 (UTC)
In addition, before my edit, there was a mixture of citation styles and many included in the Vancouver system. Your reversion re-introduced a mixture of styles as well as undone a number of other fixes. Boghog ( talk) 14:56, 8 May 2014 (UTC)
Finally per WP:CITEVAR there are no fixed "citation rules". Boghog ( talk) 15:00, 8 May 2014 (UTC)
- That version of the article predates the availability of Vancouver citation templates. It seems to me that citation templates have been changed from time to time. The template writers generally try to be backwards-compatible, but that is not entirely successful. This appears to be a case where the current CS1 and associated documentation is not compatible with this usage from 2006. The current version of WP:CITECONSENSUS states in part:
If citation templates are used in an article, the parameters should be accurate. It is inappropriate to set parameters to false values in order that the template will be rendered to the reader as if it were written in some style other than the style normally produced by the template (e.g., MLA style).
- This creates a situation where adding new citations or repairing old citations would force one to either produce inconsistent citations (as rendered) or use parameters in ways that violate the template documentation. The solution, in this case, would be to ask permission on the talk page of the article and then change the templates to vcite xxx templates. If the vcite xxx templates lack function that is needed in the article, then the article should be copyedited to consistently follow the current CS1. Jc3s5h ( talk) 15:07, 8 May 2014 (UTC)
I have started a thread at WT:CITE#"When citation style guides are updated" to confirm that others agree that citation templates in articles should be copy edited to conform to current template function and documentation. I would think the same principle would apply if a new edition of a printed publication guide is published. Jc3s5h ( talk) 15:20, 8 May 2014 (UTC)
- The Vancouver system is not a violation of {{ cite journal}} documentation nor does the Vancouver system require the use of {{ vcite journal}} template. The use of a single author parameter to store multiple authors in {{ cite journal}} has long been accepted and has not been deprecated. The primary goal for the vcite templates was speed in rendering, not how the citations were rendered. Since the lua based CS1 templates are much more efficient, the primary reason for using the vcite templates has disappeared. The best long term solution may be a modified version of {{ vcite2 journal}} that would produced clean author metadata by parsing the author parameter while avoiding the "first1, last1, first2, last2, ..." parameter clutter. Boghog ( talk) 15:23, 8 May 2014 (UTC)
- This stripping of journal names down to impenetrable crap like Expert Opin Pharmacother, and author names into run-together gibberish like "Jones HB, Smith TE, Garcia XJ", has to stop. This is not
your journal, and WP certainly does not have to follow the excessively compressed and
jargonistic style preferred by the journals you read. See the essay
WP:Specialist style fallacy for an analysis of why attempts to impose on Wikipedia some style quirk from specialist publications is usually both a bad idea and a failure. —
SMcCandlish ☺
☏
¢ ≽ʌⱷ҅ᴥⱷʌ≼
20:51, 8 May 2014 (UTC)
- It is not your journal either. Are you suggesting that we repeal WP:CITEVAR? And why is "Jones, H.B., Smith, T.E.; Garcia, X.J." necessarily better than "Jones HB, Smith TE, Garcia XJ"? The later is much cleaner and easier to read. Boghog ( talk) 20:59, 8 May 2014 (UTC)
May 2014
![]() Hello, I'm
BracketBot. I have automatically detected that
your edit to
Steroid may have broken the
syntax by modifying 1 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
Hello, I'm
BracketBot. I have automatically detected that
your edit to
Steroid may have broken the
syntax by modifying 1 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
- List of unpaired brackets remaining on the page:
- year = 2011 | publisher = Wiley | location = Hoboken | isbn = 978-0-470-66084-3 }}</ref>{rp|10–19}} and the [[anti-inflammatory]] drug [[dexamethasone]].<ref name="pmid16236742">{{cite journal |
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, BracketBot ( talk) 06:18, 12 May 2014 (UTC)
Your submission at Articles for creation: Iminosugar has been accepted

The article has been assessed as Start-Class, which is recorded on the article's talk page. You may like to take a look at the grading scheme to see how you can improve the article.
You are more than welcome to continue making quality contributions to Wikipedia. Note that because you are a logged-in user, you can create articles yourself, and don't have to post a request. However, you may continue submitting work to Articles for Creation if you prefer.
- If you have any questions, you are welcome to ask at the help desk.
- If you would like to help us improve this process, please consider .
Thank you for helping improve Wikipedia!
— Anne Delong ( talk) 03:25, 18 May 2014 (UTC)DTX2
I looked at homoloGene and Blast. It seemed to me that the Blast comparison of proteins was easier to read, and, perhaps, easier for a broader audience to interpret quickly. I was going to suggest that a direct link to protein Blast in the box would be useful. But I did not want to bother your infobox. Even though the page is about the DTX2 gene, I was looking at the human protein.
Also, I look at the page and see one sentence, and a bunch of "further readings". The page cries out for some explanation for the general user. What do all those references "mean"?
The experts are already using the specific databases and tools, they hardly need an encyclopedia article.
What could you say about specific proteins that would help a general user of Wikipedia, not an expert? Or, how does expert explain the importance and relevance of DTX2?
I think Wikipedia has a responsibility to make every page accessible to everyone.
Kind regards, RC711 ( talk) 19:08, 20 May 2014 (UTC)
- Hi. Thanks for your note. Concerning Blast Q86UW9 vs. HomoloGene, the former takes over a minute to load, while the later is almost instantaneous. Many users have a short attention span. Also the former is a mix of homo and paralogs (between and within species) and contains duplicates, whereas the later is restricted to homologous (between species). Ideally there should also be a paralog link. Perhaps the KEGG SSDB Database might work.
- The intention of the further reading sections in bot generated Gene Wiki articles was to provide notability and encourage human editors to expand the articles by providing background references some of which would hopefully be moved in-line like I just did in this .
- Finally a removed a bunch of further reading citations in the DTX2 that were not specific to that gene. Hope this helps. Boghog ( talk) 20:05, 20 May 2014 (UTC)
DTX2
Thanks for the quick update on DTX2. It is much clearer now. I could not have done it myself; those few extra words of explanation make it much better. I appreciate your hard work. RC711 ( talk) 20:01, 20 May 2014 (UTC)
- Thanks and thank you for the suggestions. Cheers. Boghog ( talk) 20:07, 20 May 2014 (UTC)
Diberri
Hi Boghog,
mw:User:Mvolz is a new community developer whose project is coding a citation auto-filler system for VisualEditor. You can see what she's planning at mw:Cite-from-id. I'm hoping to get Diberri-style PMID (etc.) filling in this as well, because I think it's important to our science-related editors. If you're interested in this, I'm sure she'd be happy to hear from you or to have someone else to bounce ideas off of. Whatamidoing (WMF) ( talk) 21:25, 20 May 2014 (UTC)
- Hi Whatamidoing. Thanks for the heads up. I definitely will follow the development of this project and participate in the discussion. It appears that the cite-from-id service will also be used by RefToolbar in addition to VisualEditor. Hence I would like to see is some flexibility in how the citations are formatted (e.g., support for "m(edical)cite" style templates). Cheers.
Boghog (
talk)
04:51, 21 May 2014 (UTC)
- My first goal is "working at all". ;-) WhatamIdoing ( talk) 05:29, 21 May 2014 (UTC)
Melanopsin
While looking at new edits to Melanopsin I noticed that the addition of info about researcher Samer Hattar was added by user:Shattar. Glancing down the history, I saw your edit summary: "enough history, return the focus back to the protein, not the researchers", Diff. The new edits are similar to the one(s) you reverted way back when. Perhaps you'd like to check this out. -- Hordaland ( talk) 01:30, 23 May 2014 (UTC)
-
 Fixed −
Seppi333 (
Insert 2¢ |
Maintained)
02:49, 23 May 2014 (UTC)
Fixed −
Seppi333 (
Insert 2¢ |
Maintained)
02:49, 23 May 2014 (UTC)
- Thank for the heads up Hordaland and Seppi333 for fixing the problem. WP:COATRACK is a good description of these types of edits. There also appears to be a WP:COI. Boghog ( talk) 04:32, 23 May 2014 (UTC)
BAGBot: Your bot request BogBot 5
Someone has marked Wikipedia:Bots/Requests for approval/BogBot 5 as needing your input. Please visit that page to reply to the requests. Thanks! AnomieBOT ⚡ 00:12, 26 May 2014 (UTC) To opt out of these notifications, place {{bots|optout=operatorassistanceneeded}} anywhere on this page.
AEBP2
Thank you for your kind assistance with improving and clarifying the AEBP2 article. Your help is most appreciated. PEGLEG3 ( talk) 12:09, 27 May 2014 (UTC)
Disambiguation link notification for June 3
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited Circadian clock, you added a link pointing to the disambiguation page ATP ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:51, 3 June 2014 (UTC)
Hi. I'm wondering if there is some reason for the various appearances: BMAL1, Bmal1 and Bmal1. I'm tempted to change them all to BMAL1, but there might be some reason for the variation. (?) Thanks, Hordaland ( talk) 16:46, 3 June 2014 (UTC)
- By convention, human genes are written with all letters in uppercase (BMAL1), non-human mammals, only the first letter is capitalized (Bmal1), all other species are in lowercase (bmal1). In general, gene symbols should be italicized. Hence it might be appropriate to have a mix of gene cases, if human diseases and animal studies are included in the same article. On the other hand, if the species is not specifically mentioned in an article, it is probably better to standardize the capitalization to all upper case (human). Cheers. Boghog ( talk) 17:16, 3 June 2014 (UTC)
Not touching anything at the Nat Product page
…not synthesis or anything, until the matter of the "course of the ship" is settled. And I heard and acknowledged Smoke, both halves of what he said. I see one ambiguous statement, leaning as much my way as yours. So there is no decision possible based on it. You are the one that wants to run with half a vote, in your prescribed direction. That, my friend defines self-serving. I am staying put until (given summer, and busy life) others have time to respond. The WP standard seems (with merger proposals, and other higher order proposed changes), to allow 30 days. That is at least what I am looking at. And sorry, no longer playing your WP:THIS and WP:THAT game. You do not seem to care to hear that you, perhaps unconsciously but nevertheless self-servingly, limit your purview when you do this, so I've ceased arguing in such ways with you. Good news, no more walls of text. Le Prof Leprof 7272 ( talk) 08:45, 12 June 2014 (UTC)
Too much to ask
but I will do it to avoid confusion when folks start responding. I would appreciate it if you would, in good faith, sandbox the last three edits that you did, since they directly bear on the questions of the discussion that were ongoing, and then revert to the version that I linked to and discussed (immediately prior to your last three edits). If you are confident in your case, you should be confident without any changes, with the form that stood for the 6 mos. of your absence. Reverting will ensure that people see what I was describing at the time, and not some muddle of old and new. Don't expect it, but in integrity, and to give you the chance, I have to ask. If you do not, say by end of day Saturday, I will have to review my opening description of the history and argument, and make sure it still makes sense (fixing links and anything else that your edit will have muddied). But as I said, the simplest is to return the matter, in good faith, to what I was talking about before those three edits. Le Prof Leprof 7272 ( talk) 08:56, 12 June 2014 (UTC)
- I really do not see how my recent edits would fundamentally change the course of the discussion. If you disagree, you can always provide a link to the previous version on the article's talk page. In the meantime, this article is like any other, and unless there is persistent edit warring or vandalism, anyone should feel free to edit it. Boghog ( talk) 20:34, 12 June 2014 (UTC)
- Thank you for the clarity of your response. I will do what I have to do, to make this work. Le Prof Leprof 7272 ( talk) 04:09, 13 June 2014 (UTC)
PK pathway enzymes
Hey Boghog, I figured it wouldn't hurt to put this in the pharmacokinetics section of the amph page, but I'm not sure what the common abbreviations are for these enzymes. I'd rather not put the full names in the drugbox.
Benzoic acid is metabolized by butyrate-CoA ligase into an intermediate product, benzoyl-CoA, [1] which is then metabolized by glycine N-acyltransferase into hippuric acid. [2]
I'm guessing the 2nd is GLYAT, but I have no clue about the ligase. Would you happen to know of a contraction or where to find one for it? Seppi333 ( Insert 2¢ | Maintained) 01:21, 8 May 2014 (UTC)
References
-
^
"butyrate-CoA ligase". BRENDA. Technische Universität Braunschweig. Retrieved 7 May 2014.
{{ cite web}}:|section=ignored ( help) -
^
"glycine N-acyltransferase". BRENDA. Technische Universität Braunschweig. Retrieved 7 May 2014.
{{ cite web}}:|section=ignored ( help)
- Hi Seppi. PMID 12616642 refers to this enzyme as a xenobiotic/medium-chain fatty acid:CoA ligase or XM-ligase for short. Benzoic acid in this context is a xenobiotic, hence I believe XM-ligase would be an appropriate abbreviation. For CoA:glycine N-acyltransferase, ACGNAT ( PMID 7833837) would be an appropriate abbreviation. Boghog ( talk) 20:00, 8 May 2014 (UTC)
- Hey again Boghog; I just updated the articles with that information, but I noticed glycine N-acyltransferase has a duplicate at GLYAT while making redirects from the shortened names you provided. Should we merge these two? Seppi333 ( Insert 2¢ | Maintained) 23:28, 21 May 2014 (UTC)
- Sorry for not responding sooner but I have been really busy in real life. It is appropriate to merge protein and enzyme articles if there is a one-to-one correspondence (i.e., only one human gene that encodes an enzyme that catalyzes a specific rxn as designated by an EC number). Digging into this further, there are the following enzyme/protein correspondences:
- Whereas the UniProt entry for GLYAT (
Q6IB77) lists EC numbers 2.3.1.13 and 2.3.1.71 as entries. Because there is not a one-to-one correspondence, I think it would be better not to merge
glycine N-acyltransferase (EC 2.3.1.13) with
GLYAT.
Boghog (
talk)
16:28, 26 May 2014 (UTC)
- Ah, alright. Thanks for pointing that out. Seppi333 ( Insert 2¢ | Maintained) 00:48, 28 May 2014 (UTC)
- One other thing - since you've made major contributions to the article (half the images were made by you) and have already done a ton of work to help address issues at the FAC, would you be interested in being a co-nominator at the next FAC for
amphetamine? I figure you deserve credit ({{
FA user topicon}}) for all you've done to help attain the
 rating. In any event, I'm going to open the 3rd FAC for it in 2-3 days since I need to do copyedits to address Shudde's issues first.
rating. In any event, I'm going to open the 3rd FAC for it in 2-3 days since I need to do copyedits to address Shudde's issues first. - Best, Seppi333 ( Insert 2¢ | Maintained) 03:17, 23 May 2014 (UTC)
- Thanks for asking but I think you are overstating my contribution ;-). In any case, I might be interested in co-nominating the article, but this would obligate me to devote a significant amount of time to address reviewers comments, time that I do not have at the moment. If you could hold off on the nomination for a week or two, I would be able spend significantly more time on improving it. Cheers.
Boghog (
talk)
16:39, 26 May 2014 (UTC)
- I seriously mean it - your help has been indispensable in copyediting and expanding the chemistry portions of the article. Thanks to you, it puts chem sections in other drug articles to shame.
 The best I could have hoped for had I written it myself is to make it merely not suck, so I mean it when I say you've done a lot for the page!
The best I could have hoped for had I written it myself is to make it merely not suck, so I mean it when I say you've done a lot for the page!
- In any event, I actually wouldn't need or expect you to help out at the FAC with anything but technical issues related to the physical/chemical properties sections, since you're familiar with those sections and way more knowledgeable than me in that area. I'm perfectly fine with handling all the basic copyediting and technical issues in the remainder of the article myself. That said, feel free to address any issues outside phys/chem if feel comfortable editing the section and have the time. Seppi333 ( Insert 2¢ | Maintained) 00:48, 28 May 2014 (UTC)
- I seriously mean it - your help has been indispensable in copyediting and expanding the chemistry portions of the article. Thanks to you, it puts chem sections in other drug articles to shame.
- Thanks for asking but I think you are overstating my contribution ;-). In any case, I might be interested in co-nominating the article, but this would obligate me to devote a significant amount of time to address reviewers comments, time that I do not have at the moment. If you could hold off on the nomination for a week or two, I would be able spend significantly more time on improving it. Cheers.
Boghog (
talk)
16:39, 26 May 2014 (UTC)
Hey again Boghog. I've been off Wikipedia for nearly 2 weeks due to a broken laptop; I have a working one now though, so I intend to renominate the article at FAC very soon. Are you still interested in being a co-nominator? Seppi333 ( Insert 2¢ | Maintained) 22:08, 13 June 2014 (UTC)
- Sorry to hear about your lap top but I am glad your connected again. I have a bit more time now for the co-nomination, so bring it on! The biggest unsolved issue as I see it is to make some of the more technical sections more accessible. I will see what I can do to help. Boghog ( talk) 22:49, 13 June 2014 (UTC)
My closing word
Not engaging with you further at NP, sorry. In your most recent spate of unscrupulous actions, you have shredded any last basis here for AGF from me. I will work around you, if possible alongside you, but any chance for real discourse or collegiality is gone. Please do not post to me, or refer to me anywhere that you can help it. Cheers, best of fortunes in our shared, difficult day to day, as we try to do something real. Le Prof Leprof 7272 ( talk) 05:37, 14 June 2014 (UTC)
- @ Leprof 7272: unscrupulous actions? Is asking a response to a simple question unscrupulous? Boghog ( talk) 05:48, 14 June 2014 (UTC)
Hello, Boghog. Some time ago you made some improvements to the draft article above, and it turned out that it was a copy-paste remnant. Was there anything that you wanted to move over to the mainspace article before the draft is deleted? — Anne Delong ( talk) 01:37, 16 June 2014 (UTC)
- Hi Anne. Thanks for the heads up. I have compared the draft with the mainspace article and it doesn't look like there is anything additional that needs to be moved over. Please go ahead and delete the draft. Boghog ( talk) 05:59, 16 June 2014 (UTC)
NFkB in neurons
Hi Boghog,
In April of 2011 I updated the the section on NFkB in neurons and I see that user Neuroglider has made substantial changes to the section that I don't agree with. Neuroglider states that s/he talked with you on 11/23/2013, though I couldn't find the talk on either of your talk pages, about removal of synaptic components as gene targets of NFkB. Could you fill me in, or point me to the correct talk page? I feel that these should be in the article as the support the idea that NFkB is important in neurons, not just glia. Which brings me to my main point. I believe that the NFkB field in neuroscience does not hold to Neuroglider's views that NFkB is primarily important in glia and not in neurons. In 2011 I specificially removed sections stating these points as I believed them to not be representative of the thoughts of the field. Specifically Meffert et.al. 2003 (citation #7 in NFkB article) [1] demonstrated that NFkB could be activated in biochemically isolated synapses (synaptoneurosomes) and was unlikely to be confounded by glia. Additionally, there are many studies that support the idea that NFkB function is important in neurons, such as studies that show cell autonomous roles for NFkB in neurons related to synaptic function and regulation of genes that are also important to synaptic function. While Neuroglider does make a good point that we should all think about heterogeneity in our systems, I believe s/he has given too much weight to this idea and the paragraph is now larger than the rest of the section. I would like to rebut Neuroglider's addition to the section, but do not wish to turn the article into a discussion of the merits of Neuroglider's v.s. other perspective. While I do not agree with Neuroglider, I'd be okay with his/her opinions to remain, but with more balance. If you have any thoughts on the best way to proceed, please let me know.
P.S. Thank you for your time and effort looking after these wikipedia articles.
-
^ Meffert MK, Chang JM, Wiltgen BJ, Fanselow MS, Baltimore D (October 2003). "NF-kappa B functions in synaptic signaling and behavior". Nat. Neurosci. 6 (10): 1072–8.
doi:
10.1038/nn1110.
PMID
12947408.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link)
16:24, 16 June 2014 (UTC)Aventre3
- Hi @ Aventre3:. I believe that the link to the relevant discussion is here. I must admit that I am not an expert on neuronal molecular biology nor have I read these citations very carefully. My interest in NFkB mainly relates to its cross talk with nuclear receptors. I suggest that you continue the discussion with @ Neuroglider: at the above link. In cases where there may be controversy, it is especially important to cite secondary sources (please note PMID 12947408 is a primary source). I will follow the discussion and jump in if I think I have anything constructive to add. Cheers. Boghog ( talk) 18:26, 16 June 2014 (UTC)
Simple matter in re steroids
For you to consider. Colleague looked at WP steroids on an iphone, and said the article opened with three structures in a row, and it seemed off-putting as a way to open an article for general readers. I checked his report, and it is true. Because of the way the image markups are inserted relative to the lede, the article opens on small screen handhelds with the three images appearing 1-2-3 that show up to the right of the lede for a standard laptop or PC viewing. Your call, but perhaps the markups could be moved down a bit, with no change of appearance for computer viewing, but resolving the negative reported by this handheld user. No RSVP necessary. Completely your call. Cheers. Le Prof Leprof 7272 ( talk) 02:10, 17 June 2014 (UTC)
I am reverting you at NP Talk because you changed more than the shading
You also changed my words in several of MY TALK ENTRIES.
If you want to revert something in particular, it is YOUR responsibility to do it selectively, and not change my words. You have no right to change my words.
And if you had a modicum of decency, you would also leave the shading in place, as it is intended to call attention to the words of individual other than ours.
If you revert again, I will ask an admin to adjudicate.
Le Prof Leprof 7272 ( talk) 15:03, 19 June 2014 (UTC)
- @
Leprof 7272: First of all, you
redactedrefactored other peoples comments in the same edit that you changed your own comments. If you don't want edits to your own comments reverted, don't combine edits to your own comments with edits to other editors comments. Better still, don'tredactrefactor other editors comment to begin with. Second, I did restore the edits to your comments in the next edit. Please more carefully look at the edit history before you jump to conclusions and assume good faith. Third, your excessively long talk page post are drowning out other voices. If you posts were more concise, it would not be necessary to highlight text. Finally mature editors try to work out differences between themselves first. Only in extreme cases is admin intervention required. Boghog ( talk) 15:24, 19 June 2014 (UTC)
- Redact means to "edit (text) for publication. censor or obscure (part of a text) for legal or security purposes." I redacted noone's text, and in shading, censored and obscured nothing. If you look closely, you did not capture all of my changes to my own text. I did look carefully at the edit history, and changed it back as I had it the first time. Go ask Pad and Smokefoot, et al, if they object. It is not your right to object for them. Leprof 7272 ( talk) 15:27, 19 June 2014 (UTC)
- @ Leprof 7272: You have it completely backwards. It is your responsibility to first ask if it is OK, not my responsibility to ask for you. Boghog ( talk) 15:31, 19 June 2014 (UTC)
- Neither Smoke nor Pad have issues, and I have asked Doc for his permission. Stop warring with me. The shading (highlighting) of other editors' comments serves a clear positive purpose. Your response is knee-jerk negative against me. Plese, stop warring. As you say, focus on content. Leprof 7272 ( talk) 15:34, 19 June 2014 (UTC)
- @ Leprof 7272: I stand behind what I state above. Stop highlighting other editor's comments without their permission. Boghog ( talk) 15:38, 19 June 2014 (UTC)
- And I by mine. The only thing approaching an edit of another's text that can be asserted, is my adding another editor's name in square brackets, where a less than careful interspersed reply from you had made unclear who had authored the original comment. Otherwise, apart from shading (highlighting), there is no change, and highlighting of a whole section, without change in content or emphasis, and indeed with elevation of attention, cannot be considered a redaction. This is a redaction. Permissions are in place as they need be. Others see merit and AGF. Le Prof Leprof 7272 ( talk) 15:49, 19 June 2014 (UTC)
- And I by mine. The only thing approaching an edit of another's text that can be asserted, is my adding another editor's name in square brackets, where a less than careful interspersed reply from you had made unclear who had authored the original comment. Otherwise, apart from shading (highlighting), there is no change, and highlighting of a whole section, without change in content or emphasis, and indeed with elevation of attention, cannot be considered a redaction. This is a
- @ Leprof 7272: The only significant change that I missed was Pharmagognosy to Pharmacogn abbreviation. Stop highlighting other editor's comments without their permission. Boghog ( talk) 15:53, 19 June 2014 (UTC)
- Reread what I just said. If you revert again, you will be subverting not only my will, but others, and I will involve adjudication. There is a page of content issues in play. Deal with the real matter at hand. Le Prof Leprof 7272 ( talk) 15:58, 19 June 2014 (UTC)
- @ Leprof 7272: One editor already objected to your highlighting of their comments. Did you receive permission from the other editors? Your selective highlighting of other user comments (unless you quote them in your own post) is bad practice and needs to stop. Boghog ( talk) 16:04, 19 June 2014 (UTC)
- Leprof, rather than
WP:REDACT, I think what Boghog means is
WP:REFACTOR.
Flyer22 (
talk)
16:24, 19 June 2014 (UTC)
- Thanks Flyer22, that is indeed what I meant. Boghog ( talk) 16:58, 19 June 2014 (UTC)
- Leprof, rather than
WP:REDACT, I think what Boghog means is
WP:REFACTOR.
Flyer22 (
talk)
16:24, 19 June 2014 (UTC)
- You're welcome. Wikipedia:Talk page guidelines#Others' comments states: " Refactoring and archiving are still appropriate, but should be done with courtesy and reversed on protest." Flyer22 ( talk) 18:41, 19 June 2014 (UTC)
Got your note and
first, no problem about it being the weekend before a response to the outline. But, did it not strike you that I might find a bit odd, so many Talk edits fitting schedule, but no "the outline looks fine, go get started"? And, for all that, in future, don't presume regarding your refactoring either (or whatever it's called). Whatever my sins, your rejiggering the whole of the Talk — appearance, order of entries, what appears on the current page, etc.—was not appreciated mate. For one, I've several incoming links, now broken, to the archived Talk. (For another, your perceived "I'm in charge here" persona is still an issue.) Effectively reading one another's minds—a point we're clearly not at yet. Nor is empathy a natural product between us. Ask first, still. (And so, to be clear, I expect to see Talk about the NP outline/changes, before any editing starts.) No reply, please. Respectful actions, enough. Le Prof Leprof 7272 ( talk) 03:33, 21 June 2014 (UTC)
It is recommended to archive or refactor a page either when it exceeds 75 KB, or has more than 10 main sections.
— WP:TALKCOND- At the time that I had archived the talk page, it had grown to 133 KB and had 15 main sections. Based on length alone, it was time to archive. Furthermore TenOfAllTrades had specifically stated that (s)he thought that the talk page was a mess and needed to be cleaned up. Padillah agreed and indicated that (s)he would archive some of the discussion. If I had not archived the discussion, Padillah probably would have done it for me. Breaking links is an unfortunate side effect of archiving talk page discussions. However one of the reasons there has been very limited engagement from other editors was it was becoming impossible for anyone to follow the discussion. Creating links into this complex discussion is not likely to generate a much of a response. I was also having trouble reading the talk page and as a consequence, it was interfering with our recent productive discussions and improving the article. It is time to put the past acrimonious discussion behind us and move on to improving the article.
Boghog (
talk)
06:46, 21 June 2014 (UTC)
- You seem particulary incapable of perceiving what sort of treatment of others will put an end to acrimony. Try respect. Asking prior to acting significantly is respectful. Acting without asking, is not. Wrong impetus mate. I have not another word to say on this. Le Prof Leprof 7272 ( talk) 00:32, 22 June 2014 (UTC)
If my mind serves me
here are some ketone-containing drugs: acetohexamide, befunolol, daunorubicin, ethacrynic acid, haloperidol, ketoprofen, metyrapone, loxoprofen, naloxone, and naltrexone. Le Prof Leprof 7272 ( talk) 04:27, 21 June 2014 (UTC)
- Not sure what you are getting at here or if this message was meant for me. Are you suggesting we create a new Category:ketone containing drugs or mention these in the ketone article? Boghog ( talk) 08:19, 21 June 2014 (UTC)
- Misplaced, and since moved. You can delete this entry if you wish. Cheers. Le Prof Leprof 7272 ( talk) 00:33, 22 June 2014 (UTC)
IL-23 subunit alpha
I think that the IL-23 subunit alpha page needs to be reduced. Right now it is a mixture of pages about IL-23 subunit alpha and the IL-23 heterodimeric cytokine. There should be a separate page for IL-23 subunit alpha, and for the cytokine itself. The page re the subunit will by definition be shorter because little is known about its function, outside of functioning as a heterodimer. What do you think? That's how it is now set up for IL-12, you can read about the cytokine IL-12 and then about each of the subunits separately. — Preceding unsigned comment added by Exiledscientist ( talk • contribs) 19:33, 26 June 2014 (UTC)
- Agreed. Interleukin 23 which is now a redirect to Interleukin 23 subunit alpha should be converted into its own article. BTW, thanks for your contributions. You clearly know the subject and I do appreciate your contributions. Cheers. Boghog ( talk) 19:43, 26 June 2014 (UTC)
- Per your suggestion, I went ahead and converted Interleukin 23 to an stand alone article. Please edit and expand as you see fit. Cheers. Boghog ( talk) 20:00, 26 June 2014 (UTC)
Disambiguation link notification for June 27
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited Interleukin 23, you added a link pointing to the disambiguation page IL-12 ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:49, 27 June 2014 (UTC)
CASK article
Hi, You removed a lot of my entries from this page. /info/en/?search=CASK This information is critical for children (like mine) who suffer from CASK gene defects. How can we manage that this information reaches the general public and the parents of the children affected with this rare disease. Can you provide me with some tips on how to integrate this information more wisely into the article ? — Preceding unsigned comment added by Reachme A ( talk • contribs) 14:59, 2 July 2014 (UTC)
- Hi. Thanks for your note. Simply stating a journal article is about the subject of a Wikipedia article is generally not sufficient reason to include it. One also needs to understand what the journal article says about the subject and write some prose to go along with the citation. To start with, I will re-add the citations into the further reading section. As I have time, I will try to integrate a few of these papers to better illustrate what I mean. Cheers. Boghog ( talk) 15:36, 2 July 2014 (UTC)
Once again, no response from you at Talk, no apparent time taken to appreciate and discuss the point of another's edit
…no questioning, just your large scale reversion and so significant further wasted time trying to interact with you. The point of my edit, as stated, was to leave in place an actual outline that we both and others can view and discuss—and not an amalgam of one editor's writing, and the remainder a partial outline it overlays.
By reverting, you nullified/broke a link in Talk to this common outline, so I and others could easily find it. IF THIS POSTED AMALGAM OF TEXT AND OUTLINE WAS INTENDED AS YOUR SANDBOX, YOU SHOULD HAVE LEFT IT IN YOUR SANDBOXES. My edit was a good faith effort to create the joint outline I have asked you for, for weeks. IF MY DERIVATION OF OUTLINE BULLET POINTS FROM YOUR TEXT, THERE, WAS NOT PERFECT, YOU COULD HAVE EDITED IT. ONCE AGAIN, WITH SWEEP OF HAND YOU SET ASIDE AN HOUR OF WORK WITHOUT REGARD. (Note, I moved your text to talk, carefully, so nothing in terms of time or effort would be lost.)
If you simply cannot work collegially, allowing others to make changes, in this case, leaving somewhere an outline of the NP article that all can see and work from—if it always has to be entirely your way, then count me out on any collaborative editing with you.
I have nothing more to say on the subject. If you cannot meet halfway, if you cannot leave an outline in place, we will proceed fully independently, and contentiously as necessary, with adjudication at every turn, to continue working on the article we each deem important. No more wasted time with you, or blatant disrespect from you. Last straw. Le Prof Leprof 7272 ( talk) 05:06, 15 July 2014 (UTC)
- @ Leprof 7272: With all due respect, you created an incomprehensible mess on the Talk:Natural product page. Because of the length and complexity of the outline that was starting to overwhelming the talk page, it is much cleaner to move the outline to a subdirectory leaving behind a link. The advantage of the subdirectory is that we can use the normal section heading syntax and a single {{ reflist}} tag to display the citations. The section headings make it much easier to navigate and to edit the outline. Furthermore it is much easier to track the revision history if the outline is separated from the discussion. Boghog ( talk) 05:30, 15 July 2014 (UTC)
- I simply can no longer tell if you are completely rude, or near completely communicatively incapable. PLEASE, reread the foregoing, noting the following terms: " …leave in place an actual outline… …not an amalgam … IF THIS POSTED AMALGAM … leaving somewhere in place an outline … ...IF MY DERIVATION OF OUTLINE BULLET POINTS FROM YOUR TEXT, THERE, WAS NOT PERFECT, YOU COULD HAVE EDITED IT… "
- Your reply ignores the thrust, and responds with matters not contested (and in fact granted). Nowhere did I contest the move of the outline. INDEED, I linked to the outline you relocated, so I clearly have affirmed the step you took in creating the new location. I contest the amalgamation of a general outline intended for all, with your individual writing and you must know this, because this is what you reverted. PLEASE, READ AND REPLY TO WHAT I ACTUALLY WROTE, AND NOT TO A THOUGHT THAT COMES INTO YOUR MIND THE SPLIT SECOND YOU BEGIN TO READ. BL: Un-amalgamated outline—where? Le Prof Leprof 7272 ( talk) 05:49, 15 July 2014 (UTC)
- @ Leprof 7272: Why do you make things so complicated? Sometimes outlines become overly abstract and in these situations it helps to flesh out the outline to determine if the outline is workable. Hence my creation of the amalgam was intentional. Why can't we continue to work on the amalgam? Boghog ( talk) 06:00, 15 July 2014 (UTC)
- That an amalgam is simpler than the outline is nonsense; easier for you perhaps, better for you perhaps, yes. Simpler, no. Considerate of others, certainly not. So if you cannot see how ones personal editing would intrude upon an outline document intended for general, overall, unbiased, all-editor friendly discussion, there is nothing to say. I ask again, as I have for weeks now, where is the un-amalgamated outline that all can discuss and work on together (leaving your editing in your sandbox)? No more argument about this, if you cannot yield, you cannot, and I will not waste another moment of either of our time. Le Prof Leprof 7272 ( talk) 06:07, 15 July 2014 (UTC)
- @ Leprof 7272: Your outline was also an amalgam with comments interspersed throughout. The prose essentially is just a more extensive comment. And how is prose any less friendly than comments? I added the prose in direct response to your comment:
Clearly, later full sections of information cannot be subsumed into the brief, early, organizational, bullet-style Classes section
- I wanted to demonstrate one can write this section without bullet points. One last point: There is a completely clean outline minus comments and prose at the top of the Talk:Natural_product/Outline page that is labeled "contents". Boghog ( talk) 06:46, 15 July 2014 (UTC)
- Since you seem incapable of grasping the plain distinction I am making, and of taking the real temperature of this interaction, let me end it here: feel free to delete all explanatory comments in the earlier draft outline that I offered, to make it fulfill your perception of a simple bare, decimal type outline that all users can add to or edit. (The autogenerated TOC is not such.) Please let me know where this simple outline resides, or tell me you will not do so. (As simple as that; "I will not do so.") I will not comment further here, or there, unless the decision is to work together. Choose one place at the Talk page, to report the location of the simple outline you create (or report that you are not willing to work with me). Done here. Le Prof
Leprof 7272 (
talk)
07:05, 15 July 2014 (UTC)
- I am working with my nephew on Latin. It is interesting how very descriptive and revealing old languages can be. For instance, L. subterfugio means literally, to flee or escape under…
Leprof 7272 (
talk)
07:09, 15 July 2014 (UTC)
- @
Leprof 7272: You have not explained why editors cannot work on
Talk:Natural_product/Outline. There is no difference in the mechanics of editing this page as opposed to the talk page. In contrast, interactively working on a large complex outline on a talk page is a really bad idea. It amount to refactoring other edit's comments and complicates the revision history. Talk pages were simply not designed for this. It is much better to place the draft outline in a subdirectory which separates comments from the collaboratively worked on proposal. Furthermore separating the outline from comments and draft prose is inefficient. One gets a birds eye view from the "contents" outline at the top with more details below.
Boghog (
talk)
07:25, 15 July 2014 (UTC)
- quam subterfugium. Change of location long granted, as you well know. Le Prof
Leprof 7272 (
talk)
07:32, 15 July 2014 (UTC)
- @
Leprof 7272: OK, good, we can move the location of the outline. We can also
collapse the prose if you like.
Boghog (
talk)
07:40, 15 July 2014 (UTC)
- Reproduced from article Talk page, please reply only there Le Prof
Leprof 7272 (
talk)
07:25, 15 July 2014 (UTC)
- Replied there. Boghog ( talk) 14:10, 15 July 2014 (UTC)
- Reproduced from article Talk page, please reply only there Le Prof
Leprof 7272 (
talk)
07:25, 15 July 2014 (UTC)
- @
Leprof 7272: OK, good, we can move the location of the outline. We can also
collapse the prose if you like.
Boghog (
talk)
07:40, 15 July 2014 (UTC)
- quam subterfugium. Change of location long granted, as you well know. Le Prof
Leprof 7272 (
talk)
07:32, 15 July 2014 (UTC)
- @
Leprof 7272: You have not explained why editors cannot work on
Talk:Natural_product/Outline. There is no difference in the mechanics of editing this page as opposed to the talk page. In contrast, interactively working on a large complex outline on a talk page is a really bad idea. It amount to refactoring other edit's comments and complicates the revision history. Talk pages were simply not designed for this. It is much better to place the draft outline in a subdirectory which separates comments from the collaboratively worked on proposal. Furthermore separating the outline from comments and draft prose is inefficient. One gets a birds eye view from the "contents" outline at the top with more details below.
Boghog (
talk)
07:25, 15 July 2014 (UTC)
- I am working with my nephew on Latin. It is interesting how very descriptive and revealing old languages can be. For instance, L. subterfugio means literally, to flee or escape under…
Leprof 7272 (
talk)
07:09, 15 July 2014 (UTC)
- Since you seem incapable of grasping the plain distinction I am making, and of taking the real temperature of this interaction, let me end it here: feel free to delete all explanatory comments in the earlier draft outline that I offered, to make it fulfill your perception of a simple bare, decimal type outline that all users can add to or edit. (The autogenerated TOC is not such.) Please let me know where this simple outline resides, or tell me you will not do so. (As simple as that; "I will not do so.") I will not comment further here, or there, unless the decision is to work together. Choose one place at the Talk page, to report the location of the simple outline you create (or report that you are not willing to work with me). Done here. Le Prof
Leprof 7272 (
talk)
07:05, 15 July 2014 (UTC)
August 2014
![]() Hello, I'm
BracketBot. I have automatically detected that
your edit to
TAS1R2 may have broken the
syntax by modifying 2 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
Hello, I'm
BracketBot. I have automatically detected that
your edit to
TAS1R2 may have broken the
syntax by modifying 2 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
- List of unpaired brackets remaining on the page:
- 3 | pages = 293–301 | year = 2003 | pmid = 12581520 | doi = 10.1016/S0092-8674(03)00071-0 }}</ref>}}
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, BracketBot ( talk) 09:20, 3 August 2014 (UTC)
Reference Errors on 3 August
![]() Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
- On the 5-HT2A receptor page, your edit caused a DOI error ( help) and a cite error ( help). ( Fix | Ask for help)
Please check this page and fix the errors highlighted. If you think this is a false positive, you can report it to my operator. Thanks, ReferenceBot ( talk) 00:25, 4 August 2014 (UTC)
Minor typo in image
Hey Boghog, I noticed a small typo in an image you uploaded: File:Inverse_agonist_2.png (activty - in blue text). It's not really a big deal since the intended word is obvious, but I figured I should point it out. Best, Seppi333 ( Insert 2¢ | Maintained) 18:28, 10 August 2014 (UTC)
-
 Fixed Hi Seppi. You have sharp eyes. I have fixed the typo and created a higher quality svg version here:
File:Inverse agonist 3.svg. Thanks for the heads up.
Boghog (
talk)
19:47, 10 August 2014 (UTC)
Fixed Hi Seppi. You have sharp eyes. I have fixed the typo and created a higher quality svg version here:
File:Inverse agonist 3.svg. Thanks for the heads up.
Boghog (
talk)
19:47, 10 August 2014 (UTC)
- Looks great - noticeably sharper than the png version. Seppi333 ( Insert 2¢ | Maintained) 02:14, 12 August 2014 (UTC)
Image for signaling cascade in addiction
References
- ^
a
b
c Renthal W, Nestler EJ (September 2009).
"Chromatin regulation in drug addiction and depression". Dialogues in Clinical Neuroscience. 11 (3): 257–268.
doi:
10.31887/DCNS.2009.11.3/wrenthal.
PMC
2834246.
PMID
19877494.
[Psychostimulants] increase cAMP levels in striatum, which activates protein kinase A (PKA) and leads to phosphorylation of its targets. This includes the cAMP response element binding protein (CREB), the phosphorylation of which induces its association with the histone acetyltransferase, CREB binding protein (CBP) to acetylate histones and facilitate gene activation. This is known to occur on many genes including fosB and c-fos in response to psychostimulant exposure. ΔFosB is also upregulated by chronic psychostimulant treatments, and is known to activate certain genes (eg, cdk5) and repress others (eg, c-fos) where it recruits HDAC1 as a corepressor. ... Chronic exposure to psychostimulants increases glutamatergic [signaling] from the prefrontal cortex to the NAc. Glutamatergic signaling elevates Ca2+ levels in NAc postsynaptic elements where it activates CaMK (calcium/calmodulin protein kinases) signaling, which, in addition to phosphorylating CREB, also phosphorylates HDAC5.
Figure 2: Psychostimulant-induced signaling events -
^ Broussard JI (January 2012).
"Co-transmission of dopamine and glutamate". The Journal of General Physiology. 139 (1): 93–96.
doi:
10.1085/jgp.201110659.
PMC
3250102.
PMID
22200950.
Coincident and convergent input often induces plasticity on a postsynaptic neuron. The NAc integrates processed information about the environment from basolateral amygdala, hippocampus, and prefrontal cortex (PFC), as well as projections from midbrain dopamine neurons. Previous studies have demonstrated how dopamine modulates this integrative process. For example, high frequency stimulation potentiates hippocampal inputs to the NAc while simultaneously depressing PFC synapses (Goto and Grace, 2005). The converse was also shown to be true; stimulation at PFC potentiates PFC–NAc synapses but depresses hippocampal–NAc synapses. In light of the new functional evidence of midbrain dopamine/glutamate co-transmission (references above), new experiments of NAc function will have to test whether midbrain glutamatergic inputs bias or filter either limbic or cortical inputs to guide goal-directed behavior.
-
^ Kanehisa Laboratories (10 October 2014).
"Amphetamine – Homo sapiens (human)". KEGG Pathway. Retrieved 31 October 2014.
Most addictive drugs increase extracellular concentrations of dopamine (DA) in nucleus accumbens (NAc) and medial prefrontal cortex (mPFC), projection areas of mesocorticolimbic DA neurons and key components of the "brain reward circuit". Amphetamine achieves this elevation in extracellular levels of DA by promoting efflux from synaptic terminals. ... Chronic exposure to amphetamine induces a unique transcription factor delta FosB, which plays an essential role in long-term adaptive changes in the brain.
- ^ Cadet JL, Brannock C, Jayanthi S, Krasnova IN (2015). "Transcriptional and epigenetic substrates of methamphetamine addiction and withdrawal: evidence from a long-access self-administration model in the rat". Molecular Neurobiology. 51 (2): 696–717 ( Figure 1). doi: 10.1007/s12035-014-8776-8. PMC 4359351. PMID 24939695.
- ^
a
b
c Robison AJ, Nestler EJ (November 2011).
"Transcriptional and epigenetic mechanisms of addiction". Nature Reviews Neuroscience. 12 (11): 623–637.
doi:
10.1038/nrn3111.
PMC
3272277.
PMID
21989194.
ΔFosB serves as one of the master control proteins governing this structural plasticity. ... ΔFosB also represses G9a expression, leading to reduced repressive histone methylation at the cdk5 gene. The net result is gene activation and increased CDK5 expression. ... In contrast, ΔFosB binds to the c-fos gene and recruits several co-repressors, including HDAC1 (histone deacetylase 1) and SIRT 1 (sirtuin 1). ... The net result is c-fos gene repression.
Figure 4: Epigenetic basis of drug regulation of gene expression - ^
a
b
c Nestler EJ (December 2012).
"Transcriptional mechanisms of drug addiction". Clinical Psychopharmacology and Neuroscience. 10 (3): 136–143.
doi:
10.9758/cpn.2012.10.3.136.
PMC
3569166.
PMID
23430970.
The 35-37 kD ΔFosB isoforms accumulate with chronic drug exposure due to their extraordinarily long half-lives. ... As a result of its stability, the ΔFosB protein persists in neurons for at least several weeks after cessation of drug exposure. ... ΔFosB overexpression in nucleus accumbens induces NFκB ... In contrast, the ability of ΔFosB to repress the c-Fos gene occurs in concert with the recruitment of a histone deacetylase and presumably several other repressive proteins such as a repressive histone methyltransferase
-
^ Nestler EJ (October 2008).
"Transcriptional mechanisms of addiction: Role of ΔFosB". Philosophical Transactions of the Royal Society B: Biological Sciences. 363 (1507): 3245–3255.
doi:
10.1098/rstb.2008.0067.
PMC
2607320.
PMID
18640924.
Recent evidence has shown that ΔFosB also represses the c-fos gene that helps create the molecular switch—from the induction of several short-lived Fos family proteins after acute drug exposure to the predominant accumulation of ΔFosB after chronic drug exposure
-
^ (Text color) Transcription factors
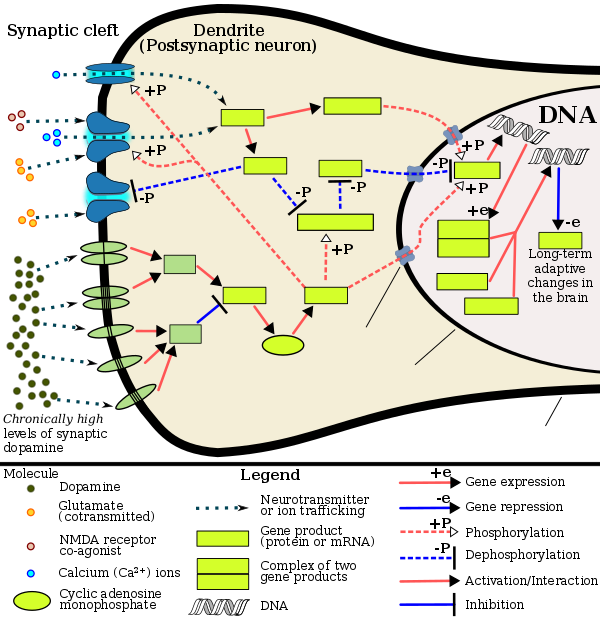 |
Hey Boghog. I was wondering if you'd happen to know of any any information resources (e.g., databases) that would have content on on signaling cascades / communication networks between receptors and gene transcription factors.
I'm working on an image ( File:ΔFosB.svg - a DA neuron/synapse in the nucleus accumbens) for the addiction section in amphetamine at the moment to illustrate how the NDMA receptors, calcium channels, and ΔFosB are related, since it's not entirely apparent in the text. This KEGG diagram (the bottom pathway) and my molecular neuropharmacology textbook are the only 2 resources I'm working off of at the moment. Unlike the KEGG pathway, I'm just doing the synaptic cleft and postsynaptic neuron since the schematic for the presynaptic half is exactly what's shown in pharmacodynamics. I know all 5 DA receptors are present in accumbens DA neurons, so I'm planning to add the 4 other DA receptors into the KEGG schematic that I recreate in order to show their influence on the cAMP cascade. Seppi333 ( Insert 2¢ | Maintained) 02:50, 17 June 2014 (UTC)
- Also if you have any suggestions or feedback for me on the diagram itself, feel free to chime in.
 Seppi333 (
Insert 2¢ |
Maintained)
03:42, 17 June 2014 (UTC)
Seppi333 (
Insert 2¢ |
Maintained)
03:42, 17 June 2014 (UTC)
- Update: I just realized that I saved a few papers on this topic on my website in a subpage that I forgot I made around 2 months ago. I unfortunately lost a large number of research papers when my laptop died, but at least I still have these signaling cascade models. In any event, I probably don't need more information resources for the model, but I'm always open to feedback! Seppi333 ( Insert 2¢ | Maintained) 00:42, 18 June 2014 (UTC)
- Hi Seppi. Sorry for not responding sooner. Your graphic looks great! One minor issue. Shouldn't the arrow between ΔFosB and DNA be reversed? According to the KEGG diagram, CREB regulates the expression of ΔFosB (i.e., CREB transcription factor → DNA (ΔFosB gene promoter) → ΔFosB protein). Also the transcription factor NFKB is a heterodimer of Fos and Jun. The KEGG diagram suggests that Fos (through the NFKB heterodimer) up regulates the expression of c-Fos. It appears that part of the diagram needs to be adjusted as well. Boghog ( talk) 17:27, 21 June 2014 (UTC)
- Ahh, alright. I'll make those tweaks when I work on the image within the next couple of days; several of the resources I used as a cross reference had a differing schematic inside the nucleus, so I wasn't really sure how to model that part. I don't expect to be done for another week or 2 since I'd like to look for additional reference models in reviews and on reactome first - assuming I can find anything relevant.
Thanks for the feedback by the way. Seppi333 ( Insert 2¢ | Maintained) 11:44, 22 June 2014 (UTC)
- Ahh, alright. I'll make those tweaks when I work on the image within the next couple of days; several of the resources I used as a cross reference had a differing schematic inside the nucleus, so I wasn't really sure how to model that part. I don't expect to be done for another week or 2 since I'd like to look for additional reference models in reviews and on reactome first - assuming I can find anything relevant.
@ Boghog: I've revised the image; I believe it's now consistent with the two KEGG diagrams and two literature reviews that I've listed in the description on the file page - File:ΔFosB.svg. The image legend is the same as KEGG's, with exception for the "+exp" and "-exp" terms which I use to denote gene expression and gene repression respectively. I'm hoping that I can make it more accessible to the average reader by removing all the SVG text and replacing it with annotated wikilinks in the finished version (similar to the amphetamine metabolites diagram). I'm also going to get some external feedback on the image's accessibility once I've finished drawing and annotating it. Consequently, it'll probably be at least another two weeks before I put it in an article.
In any event, you're more the expert in this area than I, so if there's anything you think I should add, edit, or remove from the current version of the image, please let me know! – Seppi333 ( Insert 2¢ | Maintained) 00:07, 12 July 2014 (UTC)
- It is looking even better! Two minor suggestions: Change GRIN3A → NMDAR and GRIA1 → AMPAR. GRIN3A and GRIA1 are the genes that encode subunits of NMDAR and AMPAR. The individual gene products themselves are not functional in isolation but require another subunit encoded by a distinct gene to form functional heterotetramers. Cheers. Boghog ( talk) 07:48, 12 July 2014 (UTC)
- @ Boghog: Thanks again for the feedback. More or less done now - just need to make cosmetic tweaks (the synaptic cleft and some minor tweaks) and replace the labels with a template after I remove them from the image. I might shrink the legend too, since some of those are redundant. Anyway, is there anything you think should be modified? Seppi333 ( Insert 2¢ | Maintained) 09:05, 18 July 2014 (UTC)
- @
Seppi333: Looks fantastic! As far as I can tell, it is technically correct. One further suggestion is to color code the function of the various proteins/factors. For example (using backgrounds instead of colored text):
- DRD1-5, Gs, Gi/o – GPCRs and G proteins
- ion channels
- AMPAR and NMDAR – ligand gated ion channels
- Cav1.2 – voltage gated ion channel
- cAMP, CaM – second messengers
- enzymes
- CaMKII – kinase
- DARPP-32. PP1, PP2B – phosphatases
- HDAC, SIRT1 – deacetylases
- CREB, Fos, Jun, c-Fos – transcription factors
- The following tool seems like a good way to select a color pallet. Also you might want to submit the diagram as a featured picture candidate. A few examples of diagrams that have been designated feature pictures:
- Cheers.
Boghog (
talk)
07:09, 20 July 2014 (UTC)
- Ah, that's a pretty good idea. I'll work on that sometime tomorrow, There's not much room to work with listing the color descriptions on the image itself, but I can use an annotation transclude a note/reference which contains the list you just wrote with the appropriate colors and has the group parameter "Color code". Then - assuming the transcluded page has a color code references group below the refs section - it'll transclude the named ref group along with the list which will appear (hopefully) appear in a reference tooltip whenever people mouse over it, like below. Seppi333 ( Insert 2¢ | Maintained) 09:20, 20 July 2014 (UTC)
- @
Seppi333: Looks fantastic! As far as I can tell, it is technically correct. One further suggestion is to color code the function of the various proteins/factors. For example (using backgrounds instead of colored text):
I tried a variety of color combinations with this image to see how things looked. I think I'd want to use at most 1 additional background color in the boxes (4 color groups total); it just looks too busy/complicated with more than that. Should I just identify the transcript factors, GPCRs, and ion channels, or is there a better grouping? Seppi333 ( Insert 2¢ | Maintained) 01:11, 21 July 2014 (UTC)
- @
Seppi333: I agree that adding too many colors can be counter productive (both for aesthetics and clarity). Highlighting the GPCRs, G-proteins, ion channels (membrane associated) and transcription factors (nuclear) in one color and everything else (enzymes and second messengers) in another color helps. Membrane associated proteins and the transcription factors are at least on opposite sides of the diagram and therefore already spatially distinguished. Two additional suggestions: (1) Using two different shapes for DRD-1,5 (spirals) and DRD-2,3,4 (ellipses) is confusing. Wouldn't it be better to use single ellipses for DRD-2,3,4 and retain double ellipses for DRD-1,5? That way it is immediately clear that the former are monomers while the latter are dimers. (2) The shapes for the ion channels could be altered so they are a bit closer to their actual low resolution shape (see for example
File:Ion channels.svg). This would make clearer the distinction between the ion conduction pore and the ligand binding sites.
Boghog (
talk)
05:38, 21 July 2014 (UTC)
- Hey Boghog - I updated the dopamine receptors and LGIC's, although I was actually already using the voltage gated ion channel structure in the graphic for the nuclear pore (I more or less duplicated the structure of the nuclear pore and DNA from this svg derivation of your image - File:NF-κB.svg. I improvised and created a new structure that reflected the channel structure of the LGICs for the VGIC. Hope that looks ok. I recolored them for consistency and to differentiate them from the nuclear pore; I think it's completely obvious what part constitutes the ion channel now though. I figure I can just use the multicol template to indicate color relationships.
- How's it look? Seppi333 ( Insert 2¢ | Maintained) 23:37, 21 July 2014 (UTC)
Edit: I'm still fairly certainAs is currently evident on FosB, I can transclude the color code in on an annotated reference to take advantage of the tooltip. Seppi333 ( Insert 2¢ | Maintained) 23:39, 21 July 2014 (UTC)- 2nd edit - I captioned the image with enough generality to hopefully make it reasonably accessible. Feel free to tweak it; the parts of the 3rd and 4th paper I used are in the quote parameter.
Seppi333 (
Insert 2¢ |
Maintained)
02:41, 22 July 2014 (UTC)
- @
Seppi333: Thanks for incorporating the suggested changes above. The figure and the caption are now close to perfect. Great job! Cheers.
Boghog (
talk)
15:06, 22 July 2014 (UTC)
- I may nominate this for EN-wiki FP soon, though I have no clue how people will respond to the annotation template.
File:ΔFosB.svg doesn't exactly reflect
template:psychostimulant addiction very well, even with the Commons annotations.
 In any event, I dropped by to ask you if you knew what the compound in the center of the
synthesis diagram on Bupropion is. I've turned the file into an annotated image in order to add the compound names and fix some formatting issues with that section.
Seppi333 (
Insert 2¢ |
Maintained)
23:49, 20 August 2014 (UTC)
In any event, I dropped by to ask you if you knew what the compound in the center of the
synthesis diagram on Bupropion is. I've turned the file into an annotated image in order to add the compound names and fix some formatting issues with that section.
Seppi333 (
Insert 2¢ |
Maintained)
23:49, 20 August 2014 (UTC)
- A name for the compound is 2-bromo-1-(3-chlorophenyl)propan-1-one. Beyond that, the compound probably has no special significance other than it is a synthetic intermediate. I am not sure how people will respond to the template either. From a technical standpoint, I think it is excellent. But to be featured, it also has to have aesthetic appeal. File:Steroidogenesis.svg demonstrates that it is possible to get a technical diagram designated as a feature picture. It will be interesting to see how others respond to it. Cheers. Boghog ( talk) 19:23, 21 August 2014 (UTC)
- I may nominate this for EN-wiki FP soon, though I have no clue how people will respond to the annotation template.
File:ΔFosB.svg doesn't exactly reflect
template:psychostimulant addiction very well, even with the Commons annotations.
- @
Seppi333: Thanks for incorporating the suggested changes above. The figure and the caption are now close to perfect. Great job! Cheers.
Boghog (
talk)
15:06, 22 July 2014 (UTC)
Disambiguation link notification for August 24
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited REG1B, you added a link pointing to the disambiguation page Regeneration. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 09:17, 24 August 2014 (UTC)
ANKK1 and the mistitlted ANNK1 and addictive behaviors article
Hey Boghog, I figured I should probably run this by you before doing it. Is there any reason the latter article shouldn't be merged into the former? Seppi333 ( Insert 2¢ | Maintained) 00:36, 4 September 2014 (UTC)
- @
Seppi333: Thanks for the heads up. Yes, I agree that
ANNK1 and addictive behaviors should be merged into
ANKK1.
Boghog (
talk) 06:32, 6 September 2014 (UTC)
 DoneI was bold and went ahead and merged the two article.
Boghog (
talk)
07:28, 6 September 2014 (UTC)
DoneI was bold and went ahead and merged the two article.
Boghog (
talk)
07:28, 6 September 2014 (UTC)
Disambiguation link notification for October 4
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited Morphogen, you added links pointing to the disambiguation pages Posterior and Smad. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 09:12, 4 October 2014 (UTC)
Lactase
Greetings, and thank you for your contributions. I am referring to your edit to the Lactase article, which you edited on March 3rd 2010. You changed the optimal temperature to 25 degrees Celcius, and added a reference for this claim. However, I cannot find this in the referenced paper, and a friend who is more used to chemistry publications also could not help me with that. I'm not saying you are wrong, but I kindly ask you to re-check the reference, and if possible hint me at where the temperature optimum of 25 degrees Celcius is mentioned in it. Thank you C-Otto ( talk) 21:57, 9 October 2014 (UTC)
A barnstar for you!

|
The Original Barnstar |
| Appreciate the template work you do :-) Doc James ( talk · contribs · email) (if I write on your page reply on mine) 09:34, 12 October 2014 (UTC) |
There you go, barnstar boy
Hi Boghog. Thanks for the message. Until yesterday, the page regarding PCP4 was totally empty, and I have tryied to add relevant information to it. However, I saw no editions of yours previous to that in this page. Would you possibly be more interested in deleting than adding value? About original research, that would mean research that has never been published. And the immunohistochemistry of Purkinje cell protein 4 in Purkinje cells, as well as the Western blot of PCP4 in H295R cells are well known and have been previously published. If you have access to the actual scientific journals you will be able to check that out yourself. But hey, why not just Google it? Google images id great finding scientific material of that sort sometimes. I know deleting apparently irrelevant material and adding tags to new pages can help adding up a badge of honor as a cure something-something award of Wikipedia to one's user talk page. But I see no reason to hide scientific information that has been published (and therefore does no longer constitute original research) from the public. About the content being too complex for featuring in an encyclopaedia, may I ask what would you mean by that? What I see in Wikipedia right now is some articles containing medical information that is wrong, although those are about topics that are well known since the 50's. Nevertheless, no one with a badge goes there to fix them. And no, I am not going to list these articles. I am sure you can do a good job looking for and fixing all of them. By the way, why don't you add some information about PCP4 to its page? You seem to know the topic well, I mean, you seem to know what you are talking about, right? Please do make sure to add your contribution. I will be looking forward to read it. Cheers. Herizora ( talk) 22:30, 17 October 2014 (UTC)
- Thanks for your response, however you still have not provided a clear answer to the question I asked above. Namely, where did these figures come from? Above you state that the figures are not original research and imply that they were obtained from some published source. If in fact you did obtain these images from a published source, then we have a larger problem. Unless you have obtained permission from the publisher to reuse these images, then it is a
copyright violation and the these images must be removed from Wikipedia. On the other hand, in the image summaries you state that:
-
File:PCP4 immunohistochemistry in human cerebellum.jpg:
I have performed the immunohistochemical analysis of PCP4 in 3 human cerebellum specimens
-
File:PCP4 Western blot with knock-down and control H295R adrenocortical cells.jpg:
I have performed Western blotting of PCP4 and beta-actin
-
File:PCP4 immunohistochemistry in human cerebellum.jpg:
- So which is it? Did you create these figures yourself or did you copy them from somewhere? Please answer the question.
- One of the first things I in fact did was search for the source of these images using Google Image search, and here is what I found:
- Finally what you should be doing is paraphrasing the discussion/conclusion parts of papers and not the results section. Even better would be to paraphrase relevant secondary source (e.g., review articles). If anyone is interested in experimental details, they will read directly the primary literature, not a Wikipedia article. What a Wikipedia article should do is to give a general overview of the subject. Including experimental details (e.g.,
after 6h of angiotensin-II treatment in H295R cells
) is almost never appropriate. Boghog ( talk) 14:26, 18 October 2014 (UTC)
- Hi.
- I have no obligation of answering your questions what-so-ever. But let me tell you. I have published the researched in a peer reviewed journal, and the pictures you deleted are from my PhD thesis, which was subject to peer review for several months from 4 independent peers, which makes it a secondary source.
- I thought of making those files available, both because of the good quality of the content and because my thesis is not be available online and therefore those figures cannot be accessed from the internet.
- So there you have your answer. I HAVE made those figures, and I HAVE copied them from my own peer-reviewed PUBLISHED material.
- At any rate, I will be deleting those files.
- Take your Wikipedia and stick it. I won't make material available to you any more.
- Herizora ( talk) 14:52, 18 October 2014 (UTC)
- BTW, let's keep the discussion at one place, here in your talk page. Hope it won't spoil your shiny badge. Herizora ( talk) 16:20, 18 October 2014 (UTC)
- @
Herizora: I started the discussion on your talk page, so why do you keep moving it back here? Also if someone questions the copyright status of material that you include in Wikipedia, you do have an obligation to answer to the question or agree to the removal of the material. In any case, thank you for answering my question. However your answer raises some additional concerns. While you claim your research has been published, you need to prove it (e.g., by including a filled out {{
cite thesis}} template). Otherwise your figures and associated text must be considered
WP:OR. Completed theses or dissertations may be considered reliable sources but it is preferable to cite journal articles which have undergone external peer review (see
WP:SCHOLARSHIP). Wikipedia is not a good place to publish what are essentially unpublished parts of your thesis. There is also the potential for
WP:COI. While not prohibited, including sources that you have authored in a Wikipedia article is generally frowned upon. Finally there is the appropriateness of the figures. The figures may be appropriate (e.g., to illustrate that PCP4 is expressed in Purkinje cells), but the experimental details of how the figures were obtained are not appropriate for Wikipedia. Including a citation to the published work should be sufficient.
Boghog (
talk)
19:17, 18 October 2014 (UTC)
peer review for several months from 4 independent peers, which makes it a secondary source
– No, it is still primary. Please read secondary.
- Thanks for trimming the PCP4 article. While significantly shorter, IMHO it is now much improved. Boghog ( talk) 21:32, 18 October 2014 (UTC)
- @
Herizora: I started the discussion on your talk page, so why do you keep moving it back here? Also if someone questions the copyright status of material that you include in Wikipedia, you do have an obligation to answer to the question or agree to the removal of the material. In any case, thank you for answering my question. However your answer raises some additional concerns. While you claim your research has been published, you need to prove it (e.g., by including a filled out {{
cite thesis}} template). Otherwise your figures and associated text must be considered
WP:OR. Completed theses or dissertations may be considered reliable sources but it is preferable to cite journal articles which have undergone external peer review (see
WP:SCHOLARSHIP). Wikipedia is not a good place to publish what are essentially unpublished parts of your thesis. There is also the potential for
WP:COI. While not prohibited, including sources that you have authored in a Wikipedia article is generally frowned upon. Finally there is the appropriateness of the figures. The figures may be appropriate (e.g., to illustrate that PCP4 is expressed in Purkinje cells), but the experimental details of how the figures were obtained are not appropriate for Wikipedia. Including a citation to the published work should be sufficient.
Boghog (
talk)
19:17, 18 October 2014 (UTC)
- My pleasure about the PCP4 article. Hope it will stay that way for the next few years.
- Also, thanks for the explanation about the proceedings regarding the {{ cite thesis}} template. I wish I could use the figures from the peer-reviewed article, if that was the case, but then there would be a copyright conflict with the publishing company, as I only have the legal rights to self-archive in one repository previously agreed at the time of publishing.
- While the figures that were uploaded are not unpublished, my thesis won't be available online and I have no intention of filling the {{ cite thesis}} template at present.
- Please flag the two photographs for deletion, if you find appropriate.
- And please make sure to do a more useful job. There are several supposedly serious medical articles in Wikipedia that are full of inappropriate information and pictures (ie. months ago I had to exclude a figure from a major article in medical sciences. Basically, a guy uploaded his own abdominal X-ray with the objective to show his kidneys to the world. And it had been there for months, maybe years). Maybe it's time to start addressing that kind of situation in order for Wikipedia to gain reliability.
-
Herizora (
talk)
22:10, 18 October 2014 (UTC)
- I tend to focus more on articles within the scope of WP:MCB rather than WP:MED. Furthermore there are approximately 19,000 articles in the former and 32,000 articles in the later with significant overlap between the two projects. Given the enormous number of articles and the limited number of active editors, it is not realistic that that all errors will be corrected. The best we can do right now is to concentrate on the most accessed articles. Wikipedia clearly needs to recruit and retain more editors so that we can do a better job of maintaining articles. If you see errors, please continue to correct them. We need all the help we can get! Cheers. Boghog ( talk) 08:05, 19 October 2014 (UTC)
Citation policy
Thanks very much for your comments at the Chemistry project in response to my query. I hope that you will also comment on my follow-up question. In many cases, say an article about a chemical compound or a drug etc, there are hundreds or thousands of papers. Your insights/opinions about how to select from these? -- Smokefoot ( talk) 22:53, 22 October 2014 (UTC)
Class assignment
Good afternoon, I saw you delate my contribution to the article of the pancreatic elastase because it was too much exprimental, I agree with you, but I did it for a university project. The project is due on Wenesday, after that I will delate it. So can you leave it just for now. I will really apreciate it.
Thank you so much. Have a nice week end.
BQUB14 Vmourre — Preceding unsigned comment added by BQUB14 Vmourre ( talk • contribs) 14:47, 24 October 2014 (UTC)
- While I am sympathetic to your situation, Wikipedia does not solely exist for the purpose of class assignments. Your instructor should be aware that added material must conform to Wikipedia standards, particularly with respective to WP:NOTJOURNAL (no experimental details) and WP:Verifiability (any material added must be supported by reliable sources) and communicated this to your class. Your contributions have not been deleted and can still be found in the history of the article. Your instructor should already have been in contact with the people at Wikipedia:Education program. If not, I strongly encourage your instructor to do so. Boghog ( talk) 19:54, 24 October 2014 (UTC)
FERMT3
Dear Boghog, I would really appreciate it if you stopped deleting my work from Wikipedia. The article on FERMT3 is an unfinished university assignment. We are in the process of adding online citations and references but it doesn't help that every time we work on it something has been deleted. Thank you. — Preceding unsigned comment added by Marinabuene ( talk • contribs) 10:20, 26 October 2014 (UTC)
- @ Marinabuene: Thanks for your message. If you carefully take a look at the history of the FERMT3, I have actually restored some of the content your class mates have added with WP:MEDRS compliant sourcing. I and several other editors were becoming very frustrated that our requests to your class mates to provide in-line citations were being ignored (see for example here). The only way seem to get your attention was by deleting unsourced text. I am delighted that you and your class mates are now doing so. Good luck on the completion of your assignment. Cheers. Boghog ( talk) 10:42, 26 October 2014 (UTC)
Thanks a lot for your help. We're new in Wikipedia so we don't know how it really works. Some of our references are repeated and I don't know how to delete one (there are two equal references, the same article). I would be glad if you show me how. Our work is almost done. In your opinion, is it good? — Preceding unsigned comment added by BQUB14-pdelamata ( talk • contribs) 11:55, 26 October 2014 (UTC)
- @ BQUB14-pdelamata:. Actually, it looks very good for a first effort. It needs some additional wikification and copyedit, but all in all, not too bad. You and your classmates are to be congratulated. Concerning duplicate citations, these can be combined by using named ref tags. For example, if your first citation looks like "<ref name="Doe_2014"> {{cite journal | author = Doe J | year = 2014 }}</ref>", then you can cite this source again by inserting the named ref tag "<ref name="Doe_2014"/>" (note the "/" character that is in the reused citation and not in the original). Boghog ( talk) 13:23, 26 October 2014 (UTC)
Wikipedia template filling - ISBN bug
There is no form named "frmconvert" at /data/project/citation-template-filling/perl5/lib/perl5/WWW/Mechanize.pm line 1930. at
[5]
Ihaveacatonmydesk (
talk)
14:10, 26 October 2014 (UTC)
- a lot of other examples don't work either. Ihaveacatonmydesk ( talk) 17:24, 26 October 2014 (UTC)
- I know. I took over the maintenance of the tool from
User:Diberri, and I have never been able to get this to work either. I was just happy to get the PMID and PMC searches to work and these are the most frequent type of searches. I am not a Perl expert, but I tried debugging it, but was not successful. The source code is located
here is you wanted to look into it yourself.
Boghog (
talk)
17:45, 26 October 2014 (UTC)
- Maybe put a banner on it with a notice, or remove the non-working options. Ihaveacatonmydesk ( talk) 17:17, 27 October 2014 (UTC)
- I know. I took over the maintenance of the tool from
User:Diberri, and I have never been able to get this to work either. I was just happy to get the PMID and PMC searches to work and these are the most frequent type of searches. I am not a Perl expert, but I tried debugging it, but was not successful. The source code is located
here is you wanted to look into it yourself.
Boghog (
talk)
17:45, 26 October 2014 (UTC)
Dear Boghog, sorry for any inconvenience. The deadline for the project is today, therefore we won't be making any more modifications. Thanks a lot for your kindness and for helping us with this. — Preceding unsigned comment added by BQUB14-pdelamata ( talk • contribs) 19:05, 26 October 2014 (UTC)
Citation tools
Hi Boghog. Am I correct in thinking the Cite doi and cite pmid templates have been deprecated? They don't seem to work any more and I saw that you added full references to some I'd done as cite doi, and had linked to some discussion about the citation templates. I understand you have resurrected that old template filler tool that works with pmid, but do you know if there is an equivalent for doi? Just as these days I often find myself editing on a mobile device and copy pasting the full citation is much more fiddly than just the doi. Any suggestions would be appreciated. Meodipt ( talk) 08:54, 30 October 2014 (UTC)
- Hi Meodipt. Yes, there was a consensus reached here that transcluded cite templates should no longer be used. For the top 1000 medical articles, I have gone through and substituted transcluded cite templates. I was planning to create a bot to go through all articles within the scope of the WP:MED, WP:PHARMA, and WP:MCB projects and substitute these templates, but I have not gotten around to it yet. Substitution is easy, but respecting WP:CITEVAR can be tricky. In theory, Diberri's citation tool could be extended to create citations from a DOI, but Diberri no longer maintains the tool and it is beyond my limited perl scripting abilities. With a concerted effort, I might be able to figure out how to do this, but I don't have much time to devote to this at the moment. As an alternative, you could use the RefToolbar. I personally don't like that solution because of the number of firstn, lastn parameters it will generate if there is a long author list. But it is still better than creating these by hand. Also you could try searching PubMed using the DOI as a query. This sometimes returns a PMID, especially for more recent citations. Boghog ( talk) 08:40, 1 November 2014 (UTC)
Thanks for your help
Thanks for correcting me over here, I didn't realize that NCBI stuff was public domain. Knowing that will help in the future, so I appreciate it :) Kyle Barbour 00:19, 10 November 2014 (UTC)
nuklear
In my view Nulkear is WP:NOTHERE and WP:DISRUPTIVE - it is not clear to me what your point is with the comments you are making at the SPI discussion. You seem to be trying to lay the ground for unblocking him. Is that your goal? Thanks. Jytdog ( talk) 13:49, 13 November 2014 (UTC)
- The point of my comments at the SPI discussion is that the discussion should be focused on issues relevant to an SPI. Nulkear is a serial sock puppeteer. That reason alone is sufficient for blocking confirmed socks of Nulkear and I do not dispute that for a millisecond. I was responding to Doc James' narrower question of whether the
chemical diagrams are copyright violations. If they are copyright violations, they should be removed from Wikimedia commons. For the reasons I have already stated at the SPI discussion, I think it is pretty clear that these diagrams are not copyright violations. Furthermore in the proper context with proper sources, these diagrams could legitimately be included in a Wikipedia article.
Boghog (
talk)
21:54, 13 November 2014 (UTC)
- thanks, that is very clear! and thanks for all your great work, all the time. 22:22, 13 November 2014 (UTC)
Please reinstate my fixes to Nuclear receptor
This edit reverted my fix of displayauthors errors along with the Citation bot edits to which you are objecting. You should have been able to undo just the Citation bot edits while leaving my fixes in place. Please reinstate my fixes and be more selective next time. Thanks. – Jonesey95 ( talk) 00:17, 15 November 2014 (UTC)
- @ Jonesey95: What on earth is this Help:CS1_errors#.7Cdisplayauthors.3D_suggested non-sense and why should I possibly care? The error documentation is complete gibberish. The documentation needs to clearly explain why there is a problem. Boghog ( talk) 00:33, 15 November 2014 (UTC)
The new Lua-based templates cannot know if editors created citations with exactly nine authors because there were only nine authors or, because old-style citations were limited to nine authors
– huh ??? This seems like a made up problem. In short, there is no problem. Boghog ( talk) 00:37, 15 November 2014 (UTC)
-
"Fixed" in this edit. (
|displayauthors=9) There were exactly 9 authors, not 8. Boghog ( talk) 00:54, 15 November 2014 (UTC)
- In answer to your "What on earth" question, here's a step-by-step explanation, as far as I understand it:
- The old versions of the cite templates had a limit of nine authors.
- Because of those limits, many editors and citation-creation tools created citations with only nine authors, even if the cited source had more than nine authors. Eight authors were displayed, followed by "et al."
- When the cite templates were modernized and converted to Lua, the nine-author limit was eliminated.
- Citations created before the cite templates were modernized, and that had exactly nine authors displayed, were marked with an error message, since there was no way to know whether eight authors and "et al." was correct, or whether exactly nine authors should be displayed.
- I, along with a few other gnome editors, have been comparing these citations with the source articles to determine whether more authors should be added, or if all nine authors should be displayed. In some cases, it is difficult to locate the source article, so
|displayauthors=8is added to match the old citation format. From a total of over 10,000 articles, we are down to 175. Once these 175 are fixed, the error category and error message can be eliminated from the citation template system, and the number of authors displayed will be controlled by the number in the citation, unless|displayauthors=is added intentionally by an editor. – Jonesey95 ( talk) 06:31, 15 November 2014 (UTC)
- In answer to your "What on earth" question, here's a step-by-step explanation, as far as I understand it:
- OK, thanks for the explanation. It seems like a lot of effort is going into fixing a minor problem that most readers would never notice, but at least it will completely disappear once your sweep is over.
Boghog (
talk)
09:26, 15 November 2014 (UTC)
- "a lot of effort is going into fixing a minor problem that most readers would never notice": That's my chosen vocation on WP. To each his own.... – Jonesey95 ( talk) 15:31, 15 November 2014 (UTC)
- OK, thanks for the explanation. It seems like a lot of effort is going into fixing a minor problem that most readers would never notice, but at least it will completely disappear once your sweep is over.
Boghog (
talk)
09:26, 15 November 2014 (UTC)
Reference Errors on 16 November
![]() Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
- On the Anabolic steroid page, your edit caused an unsupported parameter error ( help). ( Fix | Ask for help)
Please check this page and fix the errors highlighted. If you think this is a false positive, you can report it to my operator. Thanks, ReferenceBot ( talk) 00:39, 17 November 2014 (UTC)
Copyright checks when performing AfC reviews
Hello Boghog. This message is part of a mass mailing to people who appear active in reviewing articles for creation submissions. First of all, thank you for taking part in this important work! I'm sorry this message is a form letter – it really was the only way I could think of to covey the issue economically. Of course, this also means that I have not looked to see whether the matter is applicable to you in particular.
The issue is in rather large numbers of copyright violations ("copyvios") making their way through AfC reviews without being detected (even when easy to check, and even when hallmarks of copyvios in the text that should have invited a check, were glaring). A second issue is the correct method of dealing with them when discovered.
If you don't do so already, I'd like to ask for your to help with this problem by taking on the practice of performing a copyvio check as the first step in any AfC review. The most basic method is to simply copy a unique but small portion of text from the draft body and run it through a search engine in quotation marks. Trying this from two different paragraphs is recommended. (If you have any question about whether the text was copied from the draft, rather than the other way around (a "backwards copyvio"), the Wayback Machine is very useful for sussing that out.)
If you do find a copyright violation, please do not decline the draft on that basis. Copyright violations need to be dealt with immediately as they may harm those whose content is being used and expose Wikipedia to potential legal liability. If the draft is substantially a copyvio, and there's no non-infringing version to revert to, please mark the page for speedy deletion right away using {{db-g12|url=URL of source}}. If there is an assertion of permission, please replace the draft article's content with {{subst:copyvio|url=URL of source}}.
Some of the more obvious indicia of a copyvio are use of the first person ("we/our/us..."), phrases like "this site", or apparent artifacts of content written for somewhere else ("top", "go to top", "next page", "click here", use of smartquotes, etc.); inappropriate tone of voice, such as an overly informal tone or a very slanted marketing voice with weasel words; including intellectual property symbols (™,®); and blocks of text being added all at once in a finished form with no misspellings or other errors.
I hope this message finds you well and thanks again you for your efforts in this area. Best regards-- Fuhghettaboutit ( talk) 02:20, 18 November 2014 (UTC).
Sent via-- MediaWiki message delivery ( talk) 02:20, 18 November 2014 (UTC)
Disambiguation link notification for November 20
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited HRG (gene), you added a link pointing to the disambiguation page Rosette. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 14:21, 20 November 2014 (UTC)
Boghog -- thanks for your feedback on my Quercetin edits. Respectfully, however, biochemistry is not a natural science but rather is a major field of health science at every medical school. On the WP:SCIRS page is stated "The scope of this page is limited to the natural sciences, including astronomy, biology, chemistry, geoscience, physics, and interdiscliplinary fields. For articles about medicine, see Wikipedia:Reliable sources (medicine-related articles)."
Is it a matter of interpretation or definition? The edits I removed were made on the topic of mechanisms of action of quercetin in vitro. By my experience, I'd say this is biochemistry, so WP:MEDRS applies. All Best. -- Zefr ( talk) 23:41, 4 December 2014 (UTC)
- Respectfully, biochemistry is a natural science, not a medical science. While biochemistry is taught in medical school, biochemistry departments are normally part of colleges of liberal arts and not in medical schools. I double majored in biochemistry as an undergraduate and never attended medical school. More importantly, a biochemical mechanism of action falls within the scope of WP:MCB, not WP:MED. Boghog ( talk) 23:49, 4 December 2014 (UTC)
- "The scope of this page is limited to ... biology, chemistry". biology ∩ chemistry = biochemistry. Boghog ( talk) 23:55, 4 December 2014 (UTC)
Thank you for being one of Wikipedia's top medical contributors!
- please help translate this message into the local language

|
The Cure Award |
| In 2013 you were one of the top 300 medical editors across any language of Wikipedia. Thank you so much for helping bring free, complete, accurate, up-to-date medical information to the public. We really appreciate you and the vital work you do! |
We are wondering about the educational background of our top medical editors. Would you please complete a quick 5-question survey? (please only fill this out if you received the award)
Thanks again :) -- Ocaasi, Doc James and the team at Wiki Project Med Foundation
Deleting the message is fine
Your Talk page is yours to do with as you please. Message was intended just for you, in any case. Sad that you can see only nonsense, and nothing worth your consideration, but that is the state of things. Cheers. Le Prof
In reply to your talk my page also appearing immediately above
I have no clue as to who Flyer22 is in relation to the question of highlighting Talk text to make minority voices clearer, or what "refactoring" or "archiving" are, and what they have to do with highlighting. Here, brevity is your enemy; I am not chasing possible meanings down for you. This you need to clarify, significantly, to simply be understood. As for Padillah's statement, I have asked him to make clear what he wants, and when he does, I will act. As for your "protest", you can assuredly speak to what I do with your comments, but—as far as I see in the WPs, and as far as I have been counseled—not about what I do with other others' comments. I have highlighted none of your comments. I have queried all others, and will respond to their requests. It is, I understand, up to them, individually. (There is no overriding, firm policy, whatever you might think; this is what I am told.) So for now, I will await Padillah et al's replies, and yours if you care to say more. (But no action taken until.) Cheers. Le Prof
Disambiguation link notification for December 7
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited IGFBP3, you added a link pointing to the disambiguation page Esophageal. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:57, 7 December 2014 (UTC)
Wikipedia study- Thank you
Hello Boghog, I hope you remember speaking to me in the summer of 2012 about your motivations for contributing to the health-related pages on Wikipedia. The great news is that the study got published this Wednesday in JMIR (Journal of Medical Internet Research). You can read it here: http://www.jmir.org/2014/12/e260 This would not have been possible without your contributions so once again, I would like to thank you for taking the time and sharing your experiences with me. I also wrote an entry about my own experience with the study, about additional observations and how I plan to further extend my research - published in the WMF blog today: https://blog.wikimedia.org.uk/2014/12/who-writes-wikipedias-health-and-medical-pages-and-why/If you have any comments or questions please get in touch.Perhaps see you at the next Wikimania conference in Mexico! Best Wishes Hydra Rain ( talk) 21:03, 7 December 2014 (UTC)
Need to update template:Amphetamine pharmacodynamics...
Hey Boghog. I noticed in the recently updated KEGG pathway as well as in a recently published paper that presynaptic CaMKIIα signaling plays a role in amphetamine-induced dopamine efflux. Unfortunately, the mechanism of this isn't published because the KEGG and the DAT researchers working on this somehow have never heard of trace amine-associated receptors...
Notes from my sandbox
|
|---|
|
–Amphetamine causes
EAAT3 endocytosis (presynaptic, possibly also postsynaptic) and presynaptic CaMKIIα signaling in DA neurons -
KEGG pathway and
PMID
25033183
References
|
|
After reading all the papers, it seems intuitive to me how this works, but I wanted to run these steps by you before I update the diagram:
- Since TAAR1 is Gq-coupled, its activation causes an influx of calcium via IP3R in the presynaptic neuron (as noted in PMID 16831861 and IUPHAR's TAAR1 page).
- From this point, the intermediate pathway steps between the calcium channel and CaMKIIα are analogous to the pathway in template:Psychostimulant addiction
- CaMKIIα-dependent DAT phosphorylation occurs and results in dopamine efflux/reverse transport
In a nutshell, I'd basically be drawing the following connections: TAAR1 → IP3R → CaM → CaMKIIα → DAT (I'd need to draw IP3R). That said, is there anything missing, or does that sound right?
Seppi333 (
Insert 2¢ |
Maintained)
02:55, 12 November 2014 (UTC)
- I can't even find a review that includes amphetamine and CAMKII, so I'm just going to ignore this until someone decides to write one. The only primary source I found that mentioned both TAAR1-mediated PKC and CAMKII as mechanisms of amphetamine efflux didn't draw the conclusion I jumped to either, so I probably shouldn't do this anyway. I still think that a bit odd how there isn't a paper covering all three kinases yet.
Seppi333 (
Insert 2¢ |
Maintained)
08:15, 8 December 2014 (UTC)
- @
Seppi333: Sorry for not responding sooner. I did take a look at this a couple of weeks ago and could not find any sources that made the connection either. While your reasoning appears to be sound, unfortunately we cannot include the connection without a reliable source to support the connection, otherwise it become OR. Cheers.
Boghog (
talk)
21:23, 8 December 2014 (UTC)
- Yeah... that was my main reason for not wanting to follow up on this. At best, I can point out the associations between the three as I did in the text of
amphetamine, but I don't have any sources to adequately cite that causal chain of events. Besides that, there could even be another intracellular receptor that amph binds to which promotes CAMKII, which was my other reservation. In any event, I moved a lot of the CAMK articles around because
Ca2+/calmodulin-dependent protein kinase was originally going to the article that I moved to
Ca2+/calmodulin-dependent protein kinase II. I turned The former into a redirect to
CAMK, though I'm not sure if "Ca2+/calmodulin-dependent protein kinase" or "CAMK" should be the set index for these kinases. There doesn't appear to be any general article written that includes an overview of all subtypes.
CAMKI, the CAMKII family, and CAMKIV at the moment (I believe that's all the CAMKs, based upon BRENDA at least).Seppi333 ( Insert 2¢ | Maintained) 02:34, 9 December 2014 (UTC)
- Yeah... that was my main reason for not wanting to follow up on this. At best, I can point out the associations between the three as I did in the text of
amphetamine, but I don't have any sources to adequately cite that causal chain of events. Besides that, there could even be another intracellular receptor that amph binds to which promotes CAMKII, which was my other reservation. In any event, I moved a lot of the CAMK articles around because
Ca2+/calmodulin-dependent protein kinase was originally going to the article that I moved to
Ca2+/calmodulin-dependent protein kinase II. I turned The former into a redirect to
CAMK, though I'm not sure if "Ca2+/calmodulin-dependent protein kinase" or "CAMK" should be the set index for these kinases. There doesn't appear to be any general article written that includes an overview of all subtypes.
- @
Seppi333: Sorry for not responding sooner. I did take a look at this a couple of weeks ago and could not find any sources that made the connection either. While your reasoning appears to be sound, unfortunately we cannot include the connection without a reliable source to support the connection, otherwise it become OR. Cheers.
Boghog (
talk)
21:23, 8 December 2014 (UTC)
References
- ^ a b c d e Miller GM (January 2011). "The emerging role of trace amine-associated receptor 1 in the functional regulation of monoamine transporters and dopaminergic activity". J. Neurochem. 116 (2): 164–176. doi: 10.1111/j.1471-4159.2010.07109.x. PMC 3005101. PMID 21073468.
- ^
a
b Eiden LE, Weihe E (January 2011).
"VMAT2: a dynamic regulator of brain monoaminergic neuronal function interacting with drugs of abuse". Ann. N. Y. Acad. Sci. 1216 (1): 86–98.
Bibcode:
2011NYASA1216...86E.
doi:
10.1111/j.1749-6632.2010.05906.x.
PMC
4183197.
PMID
21272013.
VMAT2 is the CNS vesicular transporter for not only the biogenic amines DA, NE, EPI, 5-HT, and HIS, but likely also for the trace amines TYR, PEA, and thyronamine (THYR) ... [Trace aminergic] neurons in mammalian CNS would be identifiable as neurons expressing VMAT2 for storage, and the biosynthetic enzyme aromatic amino acid decarboxylase (AADC). ... AMPH release of DA from synapses requires both an action at VMAT2 to release DA to the cytoplasm and a concerted release of DA from the cytoplasm via "reverse transport" through DAT.
-
^ Sulzer D, Cragg SJ, Rice ME (August 2016).
"Striatal dopamine neurotransmission: regulation of release and uptake". Basal Ganglia. 6 (3): 123–148.
doi:
10.1016/j.baga.2016.02.001.
PMC
4850498.
PMID
27141430.
Despite the challenges in determining synaptic vesicle pH, the proton gradient across the vesicle membrane is of fundamental importance for its function. Exposure of isolated catecholamine vesicles to protonophores collapses the pH gradient and rapidly redistributes transmitter from inside to outside the vesicle. ... Amphetamine and its derivatives like methamphetamine are weak base compounds that are the only widely used class of drugs known to elicit transmitter release by a non-exocytic mechanism. As substrates for both DAT and VMAT, amphetamines can be taken up to the cytosol and then sequestered in vesicles, where they act to collapse the vesicular pH gradient.
-
^ Ledonne A, Berretta N, Davoli A, Rizzo GR, Bernardi G, Mercuri NB (July 2011).
"Electrophysiological effects of trace amines on mesencephalic dopaminergic neurons". Front. Syst. Neurosci. 5: 56.
doi:
10.3389/fnsys.2011.00056.
PMC
3131148.
PMID
21772817.
Three important new aspects of TAs action have recently emerged: (a) inhibition of firing due to increased release of dopamine; (b) reduction of D2 and GABAB receptor-mediated inhibitory responses (excitatory effects due to disinhibition); and (c) a direct TA1 receptor-mediated activation of GIRK channels which produce cell membrane hyperpolarization.
-
^
"TAAR1". GenAtlas. University of Paris. 28 January 2012. Retrieved 29 May 2014.
• tonically activates inwardly rectifying K(+) channels, which reduces the basal firing frequency of dopamine (DA) neurons of the ventral tegmental area (VTA)
-
^ Underhill SM, Wheeler DS, Li M, Watts SD, Ingram SL, Amara SG (July 2014).
"Amphetamine modulates excitatory neurotransmission through endocytosis of the glutamate transporter EAAT3 in dopamine neurons". Neuron. 83 (2): 404–416.
doi:
10.1016/j.neuron.2014.05.043.
PMC
4159050.
PMID
25033183.
AMPH also increases intracellular calcium (Gnegy et al., 2004) that is associated with calmodulin/CamKII activation (Wei et al., 2007) and modulation and trafficking of the DAT (Fog et al., 2006; Sakrikar et al., 2012). ... For example, AMPH increases extracellular glutamate in various brain regions including the striatum, VTA and NAc (Del Arco et al., 1999; Kim et al., 1981; Mora and Porras, 1993; Xue et al., 1996), but it has not been established whether this change can be explained by increased synaptic release or by reduced clearance of glutamate. ... DHK-sensitive, EAAT2 uptake was not altered by AMPH (Figure 1A). The remaining glutamate transport in these midbrain cultures is likely mediated by EAAT3 and this component was significantly decreased by AMPH
-
^ Vaughan RA, Foster JD (September 2013).
"Mechanisms of dopamine transporter regulation in normal and disease states". Trends Pharmacol. Sci. 34 (9): 489–496.
doi:
10.1016/j.tips.2013.07.005.
PMC
3831354.
PMID
23968642.
AMPH and METH also stimulate DA efflux, which is thought to be a crucial element in their addictive properties [80], although the mechanisms do not appear to be identical for each drug [81]. These processes are PKCβ– and CaMK–dependent [72, 82], and PKCβ knock-out mice display decreased AMPH-induced efflux that correlates with reduced AMPH-induced locomotion [72].
Merge proposal
You have proposed that Environmental fate of TNT be merged to TNT#Bio-degradation, but you have not opened a discussion regarding said merger. I have an opinion on the matter, but in the interest of maintaining a proper chronolgy of the discussion, I'd like you to open a discussion first. WikiDan61 ChatMe! ReadMe!! 21:48, 11 December 2014 (UTC)
 Done OK, per your request, I have opened a merge discussion
here. Sorry for not doing this sooner.
Boghog (
talk)
05:06, 12 December 2014 (UTC)
Done OK, per your request, I have opened a merge discussion
here. Sorry for not doing this sooner.
Boghog (
talk)
05:06, 12 December 2014 (UTC)
Disambiguation link notification for December 20
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited BCAR1, you added a link pointing to the disambiguation page Hypoxia. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:56, 20 December 2014 (UTC)
Thanks for your input.
As you may have gathered, this article has a new anonymous editor, who posted a question today...
Wikipedia:Help_desk#Help:Cite_errors.2FCite_error_ref_no_input. I didn’t want to frighten her off right at the start by mentioning COI (she is one of the authors of the paper she posted). However, as she first was trying to place it in Further Reading, then later included it in Ref 8, the former is now superfluous. Do you suggest I delete it now or wait till she comes back? I'm thinking that it can be off-putting for a any new editor to have loads of other editors making changes before one has had a chance to get a hang of our wiki-ways. She appears to have in depth knowledge of this subject so I welcomed her
here.--
Aspro (
talk)
00:54, 21 December 2014 (UTC)
- Hi. Thanks for you diligence in following up on this and for encouraging this new editor. Yes, clearly the editor is an expert in the field as she is a senior author on the paper she cited. While that constitutes a conflict of interest, the paper itself is both relevant and notable (it has been highly cited by independent reliable sources, see for example Google Scholar, Cited by 37 and Cited by 16 PubMed Central articles). Hence I am less concerned about the conflict of interest. Of bigger concern however is that the source is primary. Per WP:MEDRS secondary sources are needed to support medical claims. Hence I suggest that this source be replaced by a relevant review article. Boghog ( talk) 10:15, 21 December 2014 (UTC)
Disambiguation link notification for December 27
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that you've added some links pointing to disambiguation pages. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
-
ARHGEF7
- added a link pointing to Rho GTPase
-
Ecological impact of explosives
- added a link pointing to Environment
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 09:32, 27 December 2014 (UTC)
science the endless frontier
did you ever read this? Jytdog ( talk) 15:24, 29 December 2014 (UTC)
- I was aware of the Vannevar Bush report, but never read it in detail. It was enormously influential and lead to a major expansion of Federal support for research after World War II. I am not exactly sure why you brought this up, but the following quote from the report may be relevant to this recent MEDMOS discussion:
Further [medical] progress requires that the entire front of medicine and the underlying sciences of chemistry, physics, anatomy, biochemistry, physiology, pharmacology, bacteriology, pathology, parasitology, etc., be broadly developed.
Boghog (
talk)
02:31, 30 December 2014 (UTC)
| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 5 | Archive 6 | Archive 7 | Archive 8 | Archive 9 | Archive 10 | → | Archive 15 |
The Chemistry Bond Star
- Thanks and thank you for the fantastic work you are doing on amphetamine and related drug articles. Boghog ( talk) 15:56, 3 January 2014 (UTC)
Disambiguation link notification for January 4
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited C-Met, you added a link pointing to the disambiguation page Clear cell carcinoma ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:59, 4 January 2014 (UTC)
Drug box help: Created a new page
Hi - Yesterday I created a new page on Obeticholic acid which was in the news. I wanted to add a Drugbox (and further chemical details) but couldn't find Help on doing this. Are there template /bots to help with this? The CAS number is 459789-99-2. Thanks for any help. Jrfw51 ( talk) 10:03, 10 January 2014 (UTC)
- Hi. Nice work with the
Obeticholic acid article. Unfortunately there are no bots that I am aware of that can create a filled out {{
infobox drug}} template. I went ahead and manually created it. Cheers.
Boghog (
talk)
14:41, 10 January 2014 (UTC)
- Thanks for creating the Drug infobox. I see you had to reformat many of the references -- and these look better now. Why does the Cite | journal template produce such garbled input? Usually I cut and paste the PMID # but the formating then always needs changing. I see this was discussed above and I would support any extra help on this common feature for us less-than-daily contributors.
Jrfw51 (
talk)
15:31, 10 January 2014 (UTC)
- There are several ways of producing {{ cite journal}} templates. I presume that you were using the RefToolbar. I prefer the User:Diberri's Wikipedia template filling tool ( instructions). While the later requires one extra step (a copy and paste), it produces cleaner and sometimes more complete output. The RefToolbar really needs to be updated, but unfortunately there are no active maintainers of this code. Boghog ( talk) 15:56, 10 January 2014 (UTC)
- Thanks for creating the Drug infobox. I see you had to reformat many of the references -- and these look better now. Why does the Cite | journal template produce such garbled input? Usually I cut and paste the PMID # but the formating then always needs changing. I see this was discussed above and I would support any extra help on this common feature for us less-than-daily contributors.
Jrfw51 (
talk)
15:31, 10 January 2014 (UTC)
Help with citation tool
Hi, Boghog, I work for NCBI, and as discussed here on WP:NIH, I would like to get started working on a way to make it easier for editors to cite PubMed sources. I've read some of your discussions above, and I think you could probably help me scope out the task. I see that you are partial to Diberri's tool, and I'm thinking that maybe that among the things we might do is to host that at NCBI (no promises -- it's just brainstorming at this point). Maybe we could also hook it in with the ref toolbar, although I'm not familiar at all with how that works. I have a bad tendency to try to fix everything at once, and then never get anything done, so if we could scope out something reasonable, I think we could make some real progress. Let me know if you have ideas about this. Klortho ( talk) 07:08, 16 January 2014 (UTC)
Phytoestrogens
Hi Boghog, I don't usually edit pages and don't really know what's customary, but aren't readers of an article like this interested in knowing funders' agendas? I was thinking that people who are concerned about their health would want to know that a study was funded by an organization whose mission is to make sure people keep eating soy. — Preceding unsigned comment added by 75.73.43.96 ( talk) 18:28, 11 January 2014 (UTC)
- Hi. Thanks for your note. I agree who funded the study may have an impact on the reliability of the source. However if there are serious concerns about the source, I think a better course of action is to delete the source and associated text entirely from the article. The particular source in question ( PMID 19524224) appears to be WP:MEDRS compliant (secondary source published in a peer reviewed journal). Editors should not be questioning the reliability of a reliable sources directly in a Wikipedia article unless there is another reliable source to support that doubt. If there is a second reliable source that question the first source, then by all means, include that source in the article. Also:
The text of the article should not needlessly duplicate the names, dates, titles, and other information about the source that you list in the citation.
— WP:MEDMOS
- Details of who funded the study would seem to be excessive. Boghog ( talk) 20:12, 11 January 2014 (UTC)
- Thanks for your response. Now I have a much better idea of how to proceed with this article and how to deal with similar concerns in the future. — Preceding unsigned comment added by 2607:EA00:104:3C00:DC22:EE16:38E0:4E71 ( talk) 18:07, 17 January 2014 (UTC)
Favor to ask
If you're not too busy, could I ask you to look through my reply to Hamiltonstone (the 3 indented bullets) in
Wikipedia:Featured article candidates/Amphetamine/archive1#Boghog) really quickly to make sure I didn't say something stupid in my explanation on the chemistry? I'd appreciate it if you can!
![]() Seppi333 (
Insert 2¢)
17:06, 19 January 2014 (UTC)
Seppi333 (
Insert 2¢)
17:06, 19 January 2014 (UTC)
Hey again Boghog, I'm making a trace amine biosynthesis image that I plan to annotate like template:amphetamine pharmacokinetics, but I have no idea how to construct/label adrenaline consistently with the format I used for the structures - it wasn't included in the papers I used as a reference. Would you be able to give me some guidance on that? Best, Seppi333 ( Insert 2¢) 09:38, 26 January 2014 (UTC)
- Hi Seppi333. Nice work with the biosynthesis graphic. I am not sure exactly what you are asking here. One potential problem is that the R stereochemistry of the benzyl alcohol in norepinephrine is not currently specified in the figure. The absolute stereochemistry of both norepinephrine and epinephrine should be included in the figure. An additional trivial issue is the orientation of the phenyl group. The hydroxyl groups are pointed to the left in the File:Catecholamine and trace amine biosynthesis.png where they are pointed down in File:Norepinephrine structure with descriptor.svg and File:Epinephrine structure.svg. Ideally they should all be drawn in the same orientation. Or are you asking whether to label the structure as adrenaline or epinephrine? (My suggestion is the later since that what the epinephrine article is titled.) Or did you have some other concern? Boghog ( talk) 10:25, 26 January 2014 (UTC)
Ahh alright, I'll update it accordingly. I was confused by the structure difference between the paper I was using as a reference (Fig 2) and wikipedia's articles on the compounds. Seppi333 ( Insert 2¢) 11:03, 26 January 2014 (UTC)
Not done annotating the figure or tweaking the image yet, but did I address the all the chem issues with my changes?
- Yes, looks good. No major issues. There is a minor issue of the bonds not lining up exactly with the labeled atoms to which they are attached. What software are you using to produce these figures? Most chemical drawing software aligns the labeled atoms automatically. Cheers. Boghog ( talk) 17:49, 26 January 2014 (UTC)
- Not that I consider it a chem editor, but Mspaint gets the job done for me, hehe. I actually skewed the side labels slightly to crop it more, but I'll just expand it a little and center them before I use the image in the mainspace (although I just noticed I didn't center NH2 at the top). Seppi333 ( Insert 2¢) 20:54, 26 January 2014 (UTC)
Preferred format for Gene Wiki articles
Last year (13May2003) I heard from a student who was editing the c10orf76 gene page as a project in my bioinformatics analysis course that you would like me to contact you so that I could give my students formatting instructions for the Wikipedia pages on human genes of unknown function that they work on. He suggested that my class (of 44 students) was giving you and others a lot of extra work due to formatting errors in student pages. He forwarded to me some common issues you'd observed, which I'll put into my assignment for this year. If there were other formatting resources you had in mind, please let me know.
Each student wants to make a meaningful contribution to Wikipedia even if it is not a notable one. Our goal is to find sources of information where there are few obvious ones (that is their challenge) so contributing to the page on beta-globin, for instance, wouldn't work.
I look forward to hearing from you. — Preceding unsigned comment added by Wikid25 ( talk • contribs) 20:35, 29 January 2014 (UTC)
- Hi Wikid25. Thank you for contacting me and for adding my formatting requests to your class assignment. To expand on what I previously wrote, there are two problems with creating articles about obscure genes. As I previously mentioned, the first is notability. In order for an article to meet the notability requirement, the subject should have "significant coverage" (i.e., "more than a passing mention") in a reliable source. The second is that there is a high risk that these articles will contain original research, which is not permitted in Wikipedia articles. Straight forward calculations are allowed (see WP:CALC) but synthesis is not. For example, speculation as to function (protein X contains domain Y and domain Y in other proteins has function Z therefore protein X may have function Z) is synthesis. Unless this speculation (protein X may have function Z) is directly support by a reliable source, it should not be include in a Wikipedia article. Unfortunately some of the articles created by your former students are border line in terms of notability and original research. Both of these problems could be avoid if your students were to choose less obscure genes. Boghog ( talk) 04:34, 31 January 2014 (UTC)
few questions again
Hey Boghog, I noticed recently while looking up information on alpha-methylated biogenic amines that there's been some new developments in research on TAARs and their ligands - specifically TAAR5 and TAAR2. I'll probably update the stubs for those articles within the next week or so, but I'm not entirely sure what can be concluded about TAAR2 based upon that paper (its paywalled, but downloadable here). I get that the paper is asserting that TAAR1/TAAR2 are co-expressed in white blood cells and they're "involved" in chemotaxic trafficking of those cells to amines, but does that just suggest or did it actually show that PEA/TYR are hTAAR2 agonists?
- Interesting question. What the paper demonstrates is that TAAR1 and TAAR2 are coexpressed in PMN cells and likely physically interact with each other (e.g., form heterodimers). The paper also demonstrates that trace amines (TAs) are functional in PMN cells (trigger chemotactic migration and cytokine secretion) and through siRNA knockdown experiments, these activities are dependent on the presence of both TAAR1 and TAAR2. TA activity is greatly reduced if either TAAR1 or TAAR2 is knocked down. This suggests that TAAR1/TAAR2 heterodimers are required for full activity. What the paper did not demonstrate is direct binding of TAs to either TAAR1 or TAAR2. It is possible that TAs bind only to TAAR1 or TAAR2 or both partners in the TAAR1/TAAR2 complex. Alternatively TAs could bind to a third protein that signals through the TAAR1/TAAR2 complex. The bottom line is that TAs are probably agonists of TAAR2, but this has not been conclusively proven. What can be said is that TAAR2 is required for full activity of TAs in PMN cells. Boghog ( talk) 07:05, 15 February 2014 (UTC)
My other question is on N-methylhistamine and alpha-methylhistamine. I noticed some databases, like Pubchem,
list the two terms together in the synonyms, but
IUPHAR's HRH4 ligand list contains a mention of N, alpha, and 2 methylhistamine (see agonists section) around 10 times and none are indicated as being endogenouus. Aren't these different enantiomers/racemic forms of the same endogenous compound (the one made by
histamine N-methyltransferase)? Sorry for all the questions; I'm
working on something related that I'm considering on moving to article space at some point.
No rush in responding to this - I know you're busy, so take your time. Regards,
![]() Seppi333 (
Insert 2¢ |
Maintained)
03:56, 13 February 2014 (UTC)
Seppi333 (
Insert 2¢ |
Maintained)
03:56, 13 February 2014 (UTC)
- alpha-Methylhistamine (methyl group attached to the carbon adjacent to the amine group) is definitely not the same molecule as N-Methylhistamine (methyl group attached to the amino group or one of the histidine nitrogen atoms). These two molecules (or actually four molecules since N-methylhistamine is ambiguous) could be described as regio- but definitely not stereoisomers of each other. The Pubchem list should not list N-Methylhistamine as a synonym of alpha-Methylhistamine and therefore is in error. Histamine N-methyltransferase methylates the epsilon histidine nitrogen atom to produce N-Methylhistamine ( Kegg reaction R02155) which is distinct from alpha-methylhistamine. Boghog ( talk) 07:42, 15 February 2014 (UTC)
- Ah, ok that makes sense. Thanks for the explanations - they were really helpful! Best, Seppi333 ( Insert 2¢ | Maintained) 04:03, 16 February 2014 (UTC)
- Sorry to bother you with this again; I need your input in the amph FA again whenever you get a chance - WP:Featured article candidates/Amphetamine/archive2#Boghog2 - just in the 2nd bullet in the synthesis section comments. Seppi333 ( Insert 2¢ | Maintained) 02:21, 19 February 2014 (UTC)
- 2 quick questions - is this correct so far? User:Seppi333/sandbox2
- And I'm showing my chem ignorance by asking this, but what are the 2 remaining reactions for phenylacetone -> benzoic -> hippuric? Thanks for your help with the FAC btw!
Seppi333 (
Insert 2¢ |
Maintained)
10:45, 22 February 2014 (UTC)
- No problem. The graphic so far looks good. Actually what to call the phenylacetone → benzoic transformation is not trivial. The sources that you supply say it is an oxidization, but say nothing about the route. I can find no specifics about the oxidation of phenylacetone in humans. Certain bacteria have a phenylacetone monooxygenase which catalyzes the Baeyer–Villiger oxidation of phenylacetone to benzylacetate. (Warning: what follows is original research) Perhaps humans have a similar enzyme. Once formed benzylacetate can be hydrolyzed to benzyl alchohol which in turn can be oxidized into benzoic acid. Alternatively the benzylic position of phenylacetone can be oxidized to the ketone and the resulting 1,2-diketone could undergo oxidative cleavage to yield benzoic acid (End of OR warning). All you can say is that phenylacetone is (somehow) oxidized into benzoic acid. The second reaction is a conjugation of benzoic acid with glycine to yield hippuric acid (see for example PMID 1811921). I am still working on the expanding the synthesis section for the FAC. Boghog ( talk) 14:19, 22 February 2014 (UTC)
February 2014
Warning, do not remove useful information from articles, like the months of date parameters. See my revert. Debresser ( talk) 16:58, 22 February 2014 (UTC)
- Warning, do not undo constructive edits by other editors. Think before you revert. Boghog ( talk) 18:37, 22 February 2014 (UTC)
- If you make a whole set of edits, it will be your own problem if some of them are not in accordance with Wikipedia guidelines. I reverted to the last version before your removal of citation parameters, and all subsequent edits you will simply have to restore manually. I would like to point out that I seem to remember we have had this discussion before, so you were warned! Debresser ( talk) 19:05, 22 February 2014 (UTC)
- Precisely which of my edits are not in accordance with Wikipedia guidelines? What guideline states that the date parameter must specify the month? In the previous
discussion in which you "warned" me, you simply stated that I was removing useful information which I disputed and you never gave an adequate reply. This
revert with the edit summary
Perhaps this will teach him a lesson
is incredibly arrogant. Editors are supposed to resolve conflicts through discussion, not scolding. Boghog ( talk) 19:35, 22 February 2014 (UTC)
- Precisely which of my edits are not in accordance with Wikipedia guidelines? What guideline states that the date parameter must specify the month? In the previous
discussion in which you "warned" me, you simply stated that I was removing useful information which I disputed and you never gave an adequate reply. This
revert with the edit summary
- The phrase you quote was only part of my edit summary, and the whole edit summary makes better sense than you credit me for.
- Please also post the same "edit warring" warning you were so kind to paste on my talk page on your own talk page, since it is you who is the second party to this "edit war".
- The claim "you never answered my question", or as you say here "you never gave an adequate reply" is the stereotype reply of a person who does not listen to the objective arguments that have been brought forth. I have explained myself adequately, both this time and in the past. Do not remove month parameters from citation templates! Debresser ( talk) 20:55, 22 February 2014 (UTC)
- No, you have not explained yourself adequately and you have still not answered my question. What guideline states that the date parameter must specify the month? I am still waiting for an answer. Boghog ( talk) 21:10, 22 February 2014 (UTC)
- You are asking the wrong question. The question you should ask is "why was I reverted"? And the answer is, because you removed helpful information from the article, namely the months from the citation parameters. There is no rule that these must be here, as there is no rule that any information whatsoever must be in an article. But once it is there, nobody should remove it, as in Wikipedia, we do not remove helpful information without a good reason. That is called "vandalism", among other terms. Even though in this case I understand it was not vandalism motivating you, still, the act of removing information from an article is detrimental to the project and will not be tolerated. Debresser ( talk) 21:47, 22 February 2014 (UTC)
- Thank you for your detailed reply. To reiterate, month is redundant with issue in scientific citations and all the citations in the article in question are scientific. Removing redundant information is not necessarily detrimental to the project. Boghog ( talk) 22:11, 22 February 2014 (UTC)
- I see you point, even though I disagree with it. What do you say we ask at Help talk:Citation Style 1 (which is where the talk page of Cite:Journal redirects to)? Debresser ( talk) 21:12, 23 February 2014 (UTC)
Amphetamine Problems
Hey Boghog. Seppi is being pretty aggressive again, reverting edits solely because I made them. He chopped the entire toxicity section down to two short sentences (which is far too short for the nuanced topic) just to piss me off. He's also strangely, subtly vandalizing a talk page section I added. I correctly guessed the reason for his (admittedly mild) POV, which can be seen in a section of my talk page. He seems to have gone off the deep end, though, and we may need mediation. Exercisephys ( talk) 00:51, 27 February 2014 (UTC)
- I have not been following this too closely, but I have noticed some friction. This is a bit outside my area of expertise, but I will take a look and try to offer some suggestions that might help resolve the dispute. Boghog ( talk) 08:57, 27 February 2014 (UTC)
Hi, can you check the above article - a complete section has been blanked - is it ok? Regards Denisarona ( talk) 16:10, 5 March 2014 (UTC)
- If you have time, maybe also look at Amphiregulin - I don't know much about these subjects. Denisarona ( talk) 16:12, 5 March 2014 (UTC)
- Hi. Thanks for the heads up. I have mixed feelings about these edits. The intention of the further reading sections in bot generated Gene Wiki articles was to provide notability and encourage human editors to expand the articles by providing background references some of which would hopefully be moved in-line. The closest relevant guideline I could find is this:
Some editors list sources that they hope to use in the future to build the article in Further reading. This is neither encouraged nor prohibited.
— Wikipedia:Further_reading#Relation_to_reference_sections
- I myself occasionally remove these further reading sections, especially if there are large numbers of in-line citations. These particular deletions are borderline. These articles have undergone minor expansion and only contain a few in-line citations. In addition, the edit summary Cite in footnote if important for the article ( diff) is a bit condescending. If this editor only deletes a few of these sections, I would tend to ignore it. However if this editor continues these further reading section deletions on large numbs of articles, particularly if the articles have few in-line citations, I would strongly oppose it. 16:52, 5 March 2014 (UTC) Boghog ( talk)
Some stroopwafels for you!

|
For generally continuous high-level editing of articles in the medical and pharmaceutical science area. Keep it up! JFW | T@lk 17:50, 9 March 2014 (UTC) |
- Thanks for the stroopwafels! Have had tasted the delicious cheese from the same region but never before the wafels. Yum, yum. I have always thought MEDRS was a bit over the top, but I am starting to appreciate why MEDRS is necessary. Boghog ( talk) 18:48, 9 March 2014 (UTC)
- Oi, I like these so much too! Debresser ( talk) 19:05, 9 March 2014 (UTC)
Autofixing CS1 cites
Hello, User:Wikid77 here. I noticed you have been discussing ways to correct cite parameters. I am currently writing extra Lua script to autofix (auto-correct) typos when using the wp:CS1 cite templates. When I helped to develop those Lua-based cite templates, during October 2012 to April 2013, the intent was to auto-correct for typos, not issue numerous error messages, and I never imagined Lua would be used to issue thousands of red-error messages when simple auto-correction would have been quite easy. Well, after waiting all year, I have returned to re-focus on autofixing typos in cite templates, and suppress most of the red-error messages. Across Wikipedia editing, many editors are just too busy to nitpick the details and so, autofixing of cite parameters provides a rapid way to solve the problems and make many cite templates almost trivial to use. Last year, I estimated the hand-correction of cites to require over 3 years of manual, hand-crafted edits, and now after another whole year, the backlog is still about 3-5 years if hand-fixed. Although several users are diligently hand-editing the pages to fix cites, many other users are actively inserting invalid cite parameters into almost as many dozens of pages each week. The past year (of tedious cite work) has proven how autofixing is the only hope to rapidly correct the 10,000 pages in the backlog categories. For example:
During early 2014, the unsupported parameters have been fixed at only 100-200 pages per month, as meaning more years of backlog work. I wrote new essay, " wp:Autofixing cites" to explain some simple ways to autofix the major cite parameters and hope people might discuss issues about the autofixing in the talk-page there, " WT:Autofixing cites" where all the complex tactics of fixing URLs and dates could be discussed, in more detail. - Wikid77 ( talk) 21:48, 12 March 2014 (UTC)
Bot
Hey Boghog. Would you be interested in making this bot [1]? Would be happy to send you the reward :-) Doc James ( talk · contribs · email) (if I write on your page reply on mine) 06:33, 15 March 2014 (UTC)
- By the way also agree that it would be nice to move refs over one line rather than many lines. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 11:17, 15 March 2014 (UTC)
Okay it appears we may have consensus. Do you want to start the bot request or should I? What we proposing to do, if I understand correctly is:
- Replacing "cite PMID" and "cite DOI" with "cite journal"
- Replacing "cite ISBN" with "cite book"
- Placing all refs that occur over many lines to over one line
- Probably not a good idea to address the author issue yet
Doc James ( talk · contribs · email) (if I write on your page reply on mine) 00:17, 16 March 2014 (UTC)
- I recommend that you advertise more widely before deciding that you have consensus to make a substantive change that would affect tens of thousands of pages. –
Jonesey95 (
talk)
01:30, 16 March 2014 (UTC)
- The current proposal is to substitute {{
cite pmid}} with equivalent {{
cite journal}} templates, not in tens of thousands of pages, but rather in the "1500 most viewed articles" that are within the scope of
WikiProject Medicine. The proper venue for this discussion is therefore the project's talk page and the
consensus established there is overwhelmingly in favor of this change.
Boghog (
talk)
05:58, 16 March 2014 (UTC)
- Yes there is consensus among the editors who edit this content most. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 08:40, 16 March 2014 (UTC)
- The current proposal is to substitute {{
cite pmid}} with equivalent {{
cite journal}} templates, not in tens of thousands of pages, but rather in the "1500 most viewed articles" that are within the scope of
WikiProject Medicine. The proper venue for this discussion is therefore the project's talk page and the
consensus established there is overwhelmingly in favor of this change.
Boghog (
talk)
05:58, 16 March 2014 (UTC)
Looks like there is agreement for both. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 06:07, 20 March 2014 (UTC)
Is the the citation-template-filling tool on wmflabs working for you?
I get it trying to save index.cgi rather than running the script. Does it need to be authorized again or something? RDBrown ( talk) 12:26, 24 March 2014 (UTC)
I am not quite sure what you mean by "trying to save index.cgi". I just got the citation template filling tool working today by following these instructions, and in particular this solution. Boghog ( talk) 18:53, 24 March 2014 (UTC)
Template data
Is the data stored in a db for citePMID and citeISBN? Or is it a bunch of templates that somehow get polled by the individual articles when they use the cite-call? Ian Furst ( talk) 18:06, 17 March 2014 (UTC)
- I am not sure that I completely understand your question. Which data are you referring to? Currently Wikidata does not store citation data. It is planned for the future, but it has not yet been implemented. There are of course other sources of citation data. For example,
PubMed is a database that stores citations that can be retrieved using a
PMID (e.g.,
PMID
12345). There are several book databases that can be searched by
ISBN and return citation data (e.g.,
0-321-73975-2). Finally there are web tools that will return citation data given a
doi (e.g.,
CrossRef). The long term plan is to capture citation data stored in the individual Wikipedias in Wikidata and check that data against each other and against external databases. Again, I am not exactly sure how this will be implemented since it still is in the planning stage.
Boghog (
talk)
18:34, 17 March 2014 (UTC)
- I think I'm asking the question the wrong way. I've been using citeISBN in some articles. If I place citeISBN into articleA, the data is stored on another page with a title that includes that ISBN number (same with PMID, I think). Is that data just being stored on the template page or does it get entered into a relational database?
Ian Furst (
talk)
18:40, 17 March 2014 (UTC)
- Oh sorry, now I understand. The {{
cite pmid}} and {{
cite isbn}} templates stores the citation data in another template that is
transcluded back into the Wikipedia article. For example, {{cite pmid|10367338}} produces the following citation:
- Collins, A. S.; Sumner, S. C.; Borghoff, S. J.; Medinsky, M. A. (1999). "A physiological model for tert-amyl methyl ether and tert-amyl alcohol: Hypothesis testing of model structures". Toxicological sciences : an official journal of the Society of Toxicology. 49 (1): 15–28. doi: 10.1093/toxsci/49.1.15. PMID 10367338.
- The data is not stored on my talk page. If you click on the small "edit" button at the end of the citation above, it will take you to the template where the data is actually stored.
Boghog (
talk)
18:54, 17 March 2014 (UTC)
- That part I understand. Where I'm unsure is how the article {{cite pmid|}} function calls the data. Are the individual fields (name, title, etc...) that were entered into the template part of a larger backend db (e.g. is the template just a form that I'm seeing and the data is actually stored as individual fields behind the scenes and the cite function is a query. Or does the {{cite pmid|}} trigger a bot that searches the necessary fields. I'm guess I'm trying to figure out how the backend of the cite function works.
Ian Furst (
talk)
19:01, 17 March 2014 (UTC)
- Interesting question. I believe there is no database as such. Rather the data is pulled in from the template on-the-fly as needed. After the data is read from the template, it is inserted directly into the article as if it were there from the beginning. Of course there is a performance penalty for doing this. To minimize the delay in rendering the page, Wikipedia caches the fully composed Web page using Squid (software) so that it can quickly be displayed. Boghog ( talk) 19:15, 17 March 2014 (UTC)
- That part I understand. Where I'm unsure is how the article {{cite pmid|}} function calls the data. Are the individual fields (name, title, etc...) that were entered into the template part of a larger backend db (e.g. is the template just a form that I'm seeing and the data is actually stored as individual fields behind the scenes and the cite function is a query. Or does the {{cite pmid|}} trigger a bot that searches the necessary fields. I'm guess I'm trying to figure out how the backend of the cite function works.
Ian Furst (
talk)
19:01, 17 March 2014 (UTC)
- Oh sorry, now I understand. The {{
cite pmid}} and {{
cite isbn}} templates stores the citation data in another template that is
transcluded back into the Wikipedia article. For example, {{cite pmid|10367338}} produces the following citation:
- I think I'm asking the question the wrong way. I've been using citeISBN in some articles. If I place citeISBN into articleA, the data is stored on another page with a title that includes that ISBN number (same with PMID, I think). Is that data just being stored on the template page or does it get entered into a relational database?
Ian Furst (
talk)
18:40, 17 March 2014 (UTC)
Found my way to some documentation on it and you're right (sorry - should have looked for the solution myself, thought it would be a simple question). It says that it's transcluded from the template. I've done lots of db work but never used a page to act as a record source like this. My suspicion is that the parameter fields in the template are easier to screw up than a source field in a db (not to mention the performance hit). I wonder what causes problems across languages? Also, if a wiki solution is appropriate for data like this with discrete fields. It might be better managed in a db. 19:29, 17 March 2014 (UTC)
- Boghog; I was thinking more about the problems with citations and their lack of portability. My assumption is if a citations template is on a particular languages wiki it's not easily transportable or accessible from another languages wiki. What do you think about beginning a discussion on the hosting of the citation data in a database (that would be accessible to all language wiki's). A particular citation then would have a single uniqueID (PMID/ISBN/DOI or whatever) which would allow enough fields to satisfy all languages. E.g. a PMID entry might be made once in english with the field "article_name_english" but could then be translated to all 248 languages (and the same with other fields). Because it's indexable the call from the article to populate the data would be quick. Hopefully, entry is done once and the translation automated.
Ian Furst (
talk)
23:51, 17 March 2014 (UTC)
- I am not certain how Wikidata works under the hood, but I believe Wikidata is essentially a database. Furthermore Wikidata already contains language specific fields (see for example Q5633421). So in principle, Wikidata could store translated citation titles and journal names for each language. In practice, I am not sure how this would work. Perhaps Google translate could be used to provide first pass translations that could be tweaked by human editors if necessary. In any case, before support for citations in Wikidata is rolled out, these issues will need to be taken into account. See Wikidata road map for current status and future plans. Since citations are not yet on the road map, I think it will be some time before citations are added to Wikidata. Boghog ( talk) 07:56, 26 March 2014 (UTC)
Metabolism of amph (again)
Hey Boghog, I found this information of a toxicity data site: http://www.acutetox.eu/ (PDF: http://www.acutetox.eu/pdf_human_short/35-Amphetamine%20revised.pdf)
Metabolism and excretion
Metabolism of amphetamine generally includes aromatic hydroxylation, aliphatic hydroxylation, and n-dealkylation, resulting in inactive and active metabolites.
Main metabolites of amphetamine are: hippuric acid (16 to 28%), phenylacetone (0.9%), benzoylglucuronide (4%), norephedrine (2%), conjugated p-hydroxyamphetamine (2-4%) and conjugated p-hydroxynorephedrine (0.3%). One of metabolites, p-hydroxyamphetamine is a potent hallucinogen implicated in the development of amphetamine-induced psychosis [1].
The excretion of unchanged amphetamine is dependent on the urine pH. In an acidic urine (pH less than 6.6) 67 to 73% of an ingested dose is excreted as amphetamine. When pH is higher than 6.7, 17 to 43% is excreted unchanged in the urine [1].
Do you think its worth updating the metabolic pathway diagram to include "conjugated p-hydroxyamphetamine"?
Also, what program do you use to generate SVG structure diagrams? I think I may end up redoing the ones I made in that format.
Best, Seppi333 ( Insert 2¢ | Maintained) 02:35, 26 March 2014 (UTC)
- PS: Your synthesis diagrams look really nice.
- Thanks. I am not sure that it really necessary to add conjugation since it is a relatively minor metabolic pathway for amphetamine and the source you supplied does not specify what kind of conjugation is taking place (I assume it is glucuronidation). I use ChemDraw to produce the graphics, save as pdf and use pdf2svg to convert the pdf to svg format. Boghog ( talk) 12:23, 26 March 2014 (UTC)
Please tell why you reverted my paragraph
On the myostatin article. The article was from Nasdaq.com. If youre going to revert someone's edit, please tell why.-- Wyn.junior ( talk) 15:04, 28 March 2014 (UTC)
- Sorry, I should have left an edit summary. The source you have supplied just states that MYO-T12 is just entering clinical trials. Furthermore it is claimed that this dietary supplement somehow reduces myostatin levels. There are no reliable sources that support this claim. Just press releases by the company that contain the disclaimer "These statements have not been evaluated by the Food and Drug Administration. This product is not intended to diagnose, treat, cure or prevent any disease". Per WP:MEDRS, what is need to support medical claims are reliable secondary sources that review the results of control clinical trials.
- Elsewhere in the myostatin article, it states:
- "Some athletes, eager to get their hands on such drugs, turn to the internet, where fake "myostatin blockers" are being sold.". MYO-T12 is being sold a dietary supplement, but without a shred of evidence that is actually increases muscle mass. Until it is proven that it works, MYO-T12 would appear to be just another fake "myostatin blocker". Boghog ( talk) 15:30, 28 March 2014 (UTC)
- The products on the market are all pill based. Is it possible that MYO-T12 is an injection?-- Wyn.junior ( talk) 00:10, 29 March 2014 (UTC)
- I don't think it is likely that MYO-T12 could be injected. One of the main ingredients is egg yoke powder which would case a severe immune reaction if injected directly into the blood stream without first being digested. Follistatin from chicken egg yoke is claimed to be the active ingredient of MYO-T12. One might be able to inject purified chicken follistatin, but I would be very worried about an immune response. Recombinant human follistatin would be safer, but would be expensive to produce.
- PMID 24627466 has demonstrated that native follistatin injected systematically has little effect on muscle regeneration because of short half-life. A re-engineered follistatin with longer half-life had a much greater effect. In summary, MYO-T12 taken orally would not work because the follistatin it contains would be digested like any protein before it reaches the blood stream. Injected re-engineered follistatin may work, but this is not what MYO-T12 is. Bottom line, don't waste your money. This stuff cannot possibly work. Boghog ( talk) 07:48, 29 March 2014 (UTC)
- Also the text that you introduced is internally inconsistent. How could the statements "MYOS Corp is conducting its first clinical trial" and "the world's first clinically proven natural myostatin inhibitor" both be true? Boghog ( talk) 15:39, 28 March 2014 (UTC)
citation-template-filling tool
Editors using
this tool are creating references containing the
deprecated |month= parameter (e.g. in PMID, maybe others).
Are there any plans to amend the output to instead use the |date= parameter? This parameter can take a variety of complete and partial dates. --
79.67.241.76 (
talk)
18:28, 28 March 2014 (UTC)
- Thanks for your message. No one has yet to provide me with a clear explanation for why it was necessary to deprecate this parameter. But yes, I was planning to take care of this. My preference is to skip the month and return only the year in the year parameter since (1) journals themselves rarely include month in reference sections of articles, (2) month is redundant with issue, and (3) it is cleaner and avoids date formatting issues. To date, the only thing have have done with the code is to port it to the new tool server. So it may take a bit of digging to figure out how to do this, but I am working on it. Boghog ( talk) 21:52, 28 March 2014 (UTC)
- Untested, but I think this should change month, year to date. I was doubtful about month, until I checked the IEEE journals I have. Month is given at the top of pages, not issue so month may help in finding articles in bound journals ... if I'm not showing my age.
--- WWW/Wikipedia/TemplateFiller/Source/PubmedId.pm.orig 2009-05-15 11:48:47.000000000 +1000
+++ WWW/Wikipedia/TemplateFiller/Source/PubmedId.pm 2014-04-07 22:10:10.203585151 +1000
@@ -79,6 +79,10 @@ sub template_basic_fields {
my $month = Decode_Month( $self->{month} ) if $self->{month};
$month = Month_to_Text( $month ) if $month;
+ if (defined($month) && defined($self->{year})) {
+ $month = $month . ' ' . $self->{year};
+ $self->{year} = undef;
+ }
tie( my %fields, 'Tie::IxHash' );
%fields = (
@@ -89,8 +93,8 @@ sub template_basic_fields {
volume => { value => $self->{volume} },
issue => { value => $self->{issue} },
pages => { value => $pages },
- year => { value => $self->{year} },
- month => { value => $month, show => 'if-filled' },
+ year => { value => $self->{year}, show => 'if-filled' },
+ date => { value => $month, show => 'if-filled' },
pmid => { value => $self->{pmid} },
pmc => { value => $self->{pmc_id}, show => 'if-filled' },
doi => { value => $self->{doi} },
- Hope this helps/works. RDBrown ( talk) 13:12, 7 April 2014 (UTC)
- For existing references, Monkbot currently checks
|day=|month=|year=(or|month=|year=) are valid and then merges them into|date=. If month data is available in your tool it may be better to return both month and year in|date=. I don't think there's any format other than|date=March 2014available for this. When day is also present, that's when the fun begins! My preference is 28 March 2014 for publication date (e.g. newspapers), and 2014-03-28 for archivedate. Likewise the latter for accessdate, but only when the publication date is not available. -- 79.67.241.76 ( talk) 22:48, 28 March 2014 (UTC)
- For existing references, Monkbot currently checks
- Very specific reasons as to why it is better to return the year in the year parameter and skip the month were presented above. In contrast you merely state that it "may be better to return both month and year in
|date=". Also IMHO, including the day on which an article was published is excessive and completely unnecessary. Finally it makes no sense to include an accessdate in a journal citation. Articles that are published in journals must by definition have been published on a specific date. If an article does not have a publication date (e.g., year + volume + issue), it wasn't published in a journal. Boghog ( talk) 23:28, 28 March 2014 (UTC)
- Very specific reasons as to why it is better to return the year in the year parameter and skip the month were presented above. In contrast you merely state that it "may be better to return both month and year in
- Totally agree about
|accessdate=. Many other editor tools blindly shove accessdate into every reference. Whenever|date=is present I remove accessdate. As for full dates, I agree they are not needed in journal references. I should have clarified my latter comments about Monkbot and dates were meant to be more general, i.e. used for newspaper references and other such things, and not specifically to do with journals. Glad to hear you're considering updating the tools at some point. -- 79.67.241.76 ( talk) 23:53, 28 March 2014 (UTC)
- Totally agree about
Mistake
Sorry, I made a mistake deleting this paragraph. Thank you for correcting it. Humpath ( talk) 09:15, 11 April 2014 (UTC)
- No worries. Cheers. Boghog ( talk) 10:07, 11 April 2014 (UTC)
Permutation of cite journal authors
Hi! In this Boghog edit of the Bipolar disorder page, https://en.wikipedia.org/?title=Bipolar_disorder&curid=4531&diff=603896185&oldid=603893399 Bipolar disorder (Latest revision as of 11:38, 12 April 2014) the bot just permuted the author list of a "cite journal." It also didn't put them in numeric order so it seems a little pointless.
The request for approval appears to say that it only would affect 'cite doi' 'cite pmid' and 'cite isbn.' Is the Bipolar disorder change an accident or mistake? SesquiZed ( talk) 02:31, 13 April 2014 (UTC)
- Sorry.... I meant Bogbot 5 edit not Boghog edit ( /info/en/?search=Wikipedia:Bots/Requests_for_approval/BogBot_5) SesquiZed ( talk) 02:38, 13 April 2014 (UTC)
- Thanks for your note. That was an error. There was part of the bot script that was supposed to be a second pass over the substituted cite templates, not all of the templates. The error in the bot script has now been fixed. Boghog ( talk) 07:31, 13 April 2014 (UTC)
Bioinformatic analysis course Wikipedia pages on human genes of unknown function
Hi Boghog. I've tried to contact you before but don't see a reply here -- so maybe didn't get through. I'm the professor at U of M who has students write Wikipedia articles on little-known genes and have heard through my students that we should talk. I've given a formatting guide to my students based on what you sent a few of them last year but if there is more I can do to make things easier on your end, please let me know. My userID is Wikid25 and though I know you've made clear that I should come back here for a response to a question asked here, would you consider pasting any answer you might make at my talk page this once? Thanks! By the way, the following is what I give students based on what I've heard from you before:
• Please check out User:Diberri's Wikipedia template filling tool (instructions). Given a PubMed ID, one can quickly produce a full citation that can be copied and pasted into a Wikipedia article. This tool can save you a lot of work and ensure that the citations are displayed in a consistent manner. • For consistency with other Gene Wiki articles, the An Error has occurred retrieving Wikidata item for infobox infobox should normally not be included directly in the article but rather transcluded using the template. User:ProteinBoxBot is setup to maintain transcluded templates, but not templates directly contained in articles. The GeneWikiGenerator will automatically create both the template and the article that transcludes the template. • Concerning the lead sentence, as discussed here and here, we have tried to make clear that these Gene Wiki articles are not only about the human gene/protein, but also orthologs that exist in other species. The wording that was reached through consensus is perhaps a little awkward, but it is both accurate and concise: • <recommended UniProt name> is a protein that in humans is encoded by the <approved HUGO gene symbol> gene. The "that" in the above sentence is non-limiting implying that the protein (and gene) exists in other species besides human. • Per WP:HEAD, sentence case should be used for section headings. • Per WP:REFPUNC, punctuation before citations.
- Hi @ Wikid25: Thanks for your note. I did reply, but my reply has now been archived but still can be read here. If your students follow the recommendations above, that would be a great help. However, as stated in my previous reply, I the bigger concern I have is about the notability of many of these obscure genes whose function is not know. Boghog ( talk) 14:44, 13 April 2014 (UTC)
Thanks. I'll shift all future projects to genes/proteins that have at least one article in which they gene is the primary subject of the article. I'm thinking of the situation of a recently characterized gene such as SARAF/TMEM66. I will also make it clear to students that synthesis and other forms of original research are not permitted. I have discussed at length with them how all their sources must be provided such that any reasonably informed person could verify claims made. Hopefully this addresses the majority of the issues that have created work or concern for you in your editing role in the past years. — Preceding unsigned comment added by Wikid25 ( talk • contribs) 15:43, 13 April 2014 (UTC)
- Perfect! That should take care most of my concerns. I look forward to seeing your students contributions. Cheers. Boghog ( talk) 04:38, 14 April 2014 (UTC)
Disambiguation link notification for April 14
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited FGF4, you added a link pointing to the disambiguation page SHH ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:53, 14 April 2014 (UTC)
SETDB1
Hello,
About the article on Setdb1: http://en.wikipedia.org/wiki/SETDB1
"Model organisms have been used in the study of SETDB1 function" I found no article using this knock out mice.
The "abnormalities" have never been investigated and you can't conclude that there is a lymphocyte or bone issue in these mice on only one test carried out. Furthermore, this have never been reported in the literature yet. Saying that SETDB1 is implicated in bone strength and lymphocyte biology only according to the data obtained from a pre-phenotyping of a KO mice is (to my opinion) non scientifical comments. The links provided on this subject all leads to the "Wellcome Trust Sanger Institute" (that's half of the citation). It looks like more an advertisement from Eucomm that is selling the mice.
This article should be enriched with the already known roles of SETDB1 in the literature and not be an excuse reason for private companies to get a free advertisement.
I don't doubt that animal models are being widely used in research as I am a researcher myself. — Preceding unsigned comment added by 195.221.129.220 ( talk) 08:28, 16 April 2014 (UTC)
- The main problem is with your original
edit summary that stated "Deleted non scientifical comments". Edit summaries do not need to be long, but if you are deleted sourced material, you need to provide more rationale for your deletion. Even your more lengthy explanation above is problematic:
- "I found no article using this knock out mice" – see SETDB1 knockout
- "not be an excuse reason for private companies to get a free advertisement" – neither the Sanger Institute not the EUCOMM is a private company, both are a non-profit organizations.
- On the other hand, the citations provided were not specific to this gene. Hence I would agree that the deleted section as written is not appropriate for this article. Boghog ( talk) 17:58, 16 April 2014 (UTC)
Rictor page
Boghog:
Just wanted to let you know that I am working on the Rictor page for a class assignment. I would really appreciate it if you would hold off on your edits (especially the stylistic ones) until I submit this assignment for grading.
Thanks for your interest in the page and your current contributions.
kldextraze — Preceding unsigned comment added by Kldextraze ( talk • contribs) 16:43, 17 April 2014 (UTC)
WikiProject Genetics in the Signpost
The WikiProject Report would like to focus on WikiProject Genetics for a Signpost article. This is an excellent opportunity to draw attention to your efforts and attract new members to the project. Would you be willing to participate in an interview? If so, here are the questions for the interview. Just add your response below each question and feel free to skip any questions that you don't feel comfortable answering. Multiple editors will have an opportunity to respond to the interview questions, so be sure to sign your answers. If you know anyone else who would like to participate in the interview, please share this with them. Have a great day. –Mabeenot ( talk) 18:53, 19 April 2014 (UTC)
Bot for ref formatting
Where are we at? Doc James ( talk · contribs · email) (if I write on your page reply on mine) 03:31, 21 April 2014 (UTC)
- I just noticed the bot transcluded cite templates. Thanks. Blue Rasberry (talk) 19:07, 21 April 2014 (UTC)
A barnstar for you!

|
The Technical Barnstar |
| For fixing the refs you have :-) Looking forwards to the larger run. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 19:37, 21 April 2014 (UTC) |
- By the way you rock! So looking forwards to not having to fix these by hand :-) Doc James ( talk · contribs · email) (if I write on your page reply on mine) 19:37, 21 April 2014 (UTC)
Thanks, but I am not quite done. I should be able to finish this soon after I fix a few bugs in the bot script. BTW, it turns out that there are only about ~300 of the 1500 MED articles that have transcluded cite pmid/doi/isbn templates. Boghog ( talk) 19:42, 21 April 2014 (UTC)
- Ah good to know. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 20:09, 21 April 2014 (UTC)
Refs for preg cat
IMO we should have refereces for the preg cat listed in the info boxes. Could we get a bot to add these? Doc James ( talk · contribs · email) (if I write on your page reply on mine) 09:22, 24 April 2014 (UTC)
- Sure, adding references using a bot is no problem. But where do the references come from? Is there a generic reference to some database that we can use? Or are there drug specific references?
Boghog (
talk)
10:37, 24 April 2014 (UTC)
- So this lists the USA
http://www.drugs.com/monograph/isosorbide-dinitrate-mononitrate.html and this lists the Australian
http://www.tga.gov.au/hp/medicines-pregnancy.htm#.U1jqHRc3tqx
Doc James (
talk ·
contribs ·
email) (if I write on your page reply on mine)
10:54, 24 April 2014 (UTC)
- Could we format the monograph as <ref name=AHFS2014>{{cite web|title=Cephalexin|url=http://www.drugs.com/monograph/cephalexin.html|publisher=The American Society of Health-System Pharmacists|accessdate=Apr 21, 2014}}</ref> as I use it frequently for other stuff. Doc James ( talk · contribs · email) (if I write on your page reply on mine) 10:55, 24 April 2014 (UTC)
- So this lists the USA
http://www.drugs.com/monograph/isosorbide-dinitrate-mononitrate.html and this lists the Australian
http://www.tga.gov.au/hp/medicines-pregnancy.htm#.U1jqHRc3tqx
Doc James (
talk ·
contribs ·
email) (if I write on your page reply on mine)
10:54, 24 April 2014 (UTC)
Invitation join the new Physiology Wikiproject!

Based on the long felt gap for categorization and improvization of WP:MED articles relating to the field of physiology, the new WikiProject Physiology has been created. WikiProject Physiology is still in its infancy and needs your help. On behalf of a group of editors striving to improve the quality of physiology articles here on Wikipedia, I would like to invite you to come on board and participate in the betterment of physiology related articles. Help us to jumpstart this WikiProject.
- Feel free to leave us a message at any time on the WikiProkect Physiology talk page. If you are interested in joining the project yourself, there is a participant list where you can sign up. Please leave a message on the talk page if you have any problems, suggestions, would like review of an article, need suggestions for articles to edit, or would like some collaboration when editing!
- You can tag the talk pages of relevant articles with {{WikiProject Physiology|class=|importance=}} with your assessment of the article class and importance alongwith. Please note that WP:Physiology, WP:Physio, WP:Phy can be used interchangeably.
- You will make a big difference to the quality of information by adding reliable sources. Sourcing physiology articles is essential and makes a big difference to the quality of articles. And, while you're at it, why not use a book to source information, which can source multiple articles at once!
- We try and use a standard way of arranging the content in each article. That layout is here. These headings let us have a standard way of presenting the information in anatomical articles, indicate what information may have been forgotten, and save angst when trying to decide how to organise an article. That said, this might not suit every article. If in doubt, be bold!
- Why not try and strive to create a good article! Physiology related articles are often small in scope, have available sources, and only a limited amount of research available that is readily presentable!
- Your contributions to the WikiProject page, related categories and templates is also welcome.
- To invite other editors to this WikiProject, copy and past this template (with the signature):
{{ subst:WP Physiology–invite}}~~~~
- To welcome editors of physiology articles, copy and past this template (with the signature):
{{ subst:WP Physiology–welcome}}~~~~
- You can feel free to contact us on the WikiProkect Physiology talk page if you have any problems, or wish to join us. You can also put your suggestions there and discuss the scope of participation.
Hoping for your cooperation! Diptanshu Talk 12:27, 27 April 2014 (UTC)
EDITS
It's my understanding that edit disagreements are best discussed on the talk page of the site where the edits are occurring. Are you OK with that approach? If so I'll leave you a note on the Ghrelin talk page for your response. Thanks.
Regards -
IiKkEe ( talk) 05:57, 5 May 2014 (UTC)
- Hi. Thanks for your message and for the impressive work you have done on the Ghrelin and related articles. In answer to your question, it is generally best to have article specific discussions on the article's talk page. That keeps the discussion in one place so that it easier to follow. Cheers. Boghog ( talk) 16:13, 5 May 2014 (UTC)
Edit to Tramadol
This edit appears to use some kind of automation to "fix" citations. However, it does not follow Help: Citation Style 1 (CS1). At least one difference is that CS1 separates authors with a semicolon, while you separated them with a comma. Please repair your automation before using it again. Jc3s5h ( talk) 14:20, 8 May 2014 (UTC)
- My edit does follow {{ cite journal}} (see Vancouver system). Also this was a manual edit that included a lot of other fixes. Boghog ( talk) 14:26, 8 May 2014 (UTC)
I noticed another error; authors who apparently have western European names have their names given in the format "Doe J" when it should be "Doe, J". Jc3s5h ( talk) 14:30, 8 May 2014 (UTC)
- That is not an error. That is intentional (see Vancouver system). Boghog ( talk) 14:31, 8 May 2014 (UTC)
- No, I will not look at Vancouver system. The article does not use Vancouver system. Consider this citation:
- Nelson, G.K.; Lombardi, M.A.; Okayama, D.T. (2005).
"NIST Time and Frequency Radio Stations: WWV, WWVH, and WWVB" (PDF).
National Institute of Standards and Technology. (Special Publication 250-67).
{{ cite web}}: Invalid|ref=harv( help) dead link
- Nelson, G.K.; Lombardi, M.A.; Okayama, D.T. (2005).
"NIST Time and Frequency Radio Stations: WWV, WWVH, and WWVB" (PDF).
National Institute of Standards and Technology. (Special Publication 250-67).
- No, I will not look at Vancouver system. The article does not use Vancouver system. Consider this citation:
- The authors are specified with the perfectly valid syntax
"| last1=Nelson |first1=G.K. |last2=Lombardi |first2=M.A. |last3=Okayama |first3=D.T."The results are last names separated from first names with commas, and authors separated by semicolons. You must follow this format or you are in violation of WP:CITEVAR. Jc3s5h ( talk) 14:37, 8 May 2014 (UTC)
- The authors are specified with the perfectly valid syntax
- That is not what WP:CITEVAR says. Please re-read it. The predominate style was initially the Vancouver system and I am restoring it. Boghog ( talk) 14:40, 8 May 2014 (UTC)
- Help:Citation Style 1 sets forth the style for templates in the Citaion Style 1 family (which includes {{ cite journal}}. If you believe the article uses Vancouver style, ask on the talk page of the article to change the templates to Help:Citation Style Vancouver. It is not reasonable to use templates in a way that legal variations on ways of expressing parameters result in different rendered citations.
- If you insist on this idiosyncratic interpretation of WP:CITEVAR I will bring this to the discussion about your new bot; we can't have bot operators writing citation-related bots if their ideas about citation rules are different from everyone else's. Jc3s5h ( talk) 14:49, 8 May 2014 (UTC)
Consider this
older version of the article which show a clear preference for the
Vancouver system. What
WP:CITEVAR says is defer to the style used by the first major contributor
. Your revert of my edit is in clear violation of
WP:CITEVAR.
Boghog (
talk)
14:51, 8 May 2014 (UTC)
Also I repeat, this was not a bot edit. Boghog ( talk) 14:52, 8 May 2014 (UTC)
In addition, before my edit, there was a mixture of citation styles and many included in the Vancouver system. Your reversion re-introduced a mixture of styles as well as undone a number of other fixes. Boghog ( talk) 14:56, 8 May 2014 (UTC)
Finally per WP:CITEVAR there are no fixed "citation rules". Boghog ( talk) 15:00, 8 May 2014 (UTC)
- That version of the article predates the availability of Vancouver citation templates. It seems to me that citation templates have been changed from time to time. The template writers generally try to be backwards-compatible, but that is not entirely successful. This appears to be a case where the current CS1 and associated documentation is not compatible with this usage from 2006. The current version of WP:CITECONSENSUS states in part:
If citation templates are used in an article, the parameters should be accurate. It is inappropriate to set parameters to false values in order that the template will be rendered to the reader as if it were written in some style other than the style normally produced by the template (e.g., MLA style).
- This creates a situation where adding new citations or repairing old citations would force one to either produce inconsistent citations (as rendered) or use parameters in ways that violate the template documentation. The solution, in this case, would be to ask permission on the talk page of the article and then change the templates to vcite xxx templates. If the vcite xxx templates lack function that is needed in the article, then the article should be copyedited to consistently follow the current CS1. Jc3s5h ( talk) 15:07, 8 May 2014 (UTC)
I have started a thread at WT:CITE#"When citation style guides are updated" to confirm that others agree that citation templates in articles should be copy edited to conform to current template function and documentation. I would think the same principle would apply if a new edition of a printed publication guide is published. Jc3s5h ( talk) 15:20, 8 May 2014 (UTC)
- The Vancouver system is not a violation of {{ cite journal}} documentation nor does the Vancouver system require the use of {{ vcite journal}} template. The use of a single author parameter to store multiple authors in {{ cite journal}} has long been accepted and has not been deprecated. The primary goal for the vcite templates was speed in rendering, not how the citations were rendered. Since the lua based CS1 templates are much more efficient, the primary reason for using the vcite templates has disappeared. The best long term solution may be a modified version of {{ vcite2 journal}} that would produced clean author metadata by parsing the author parameter while avoiding the "first1, last1, first2, last2, ..." parameter clutter. Boghog ( talk) 15:23, 8 May 2014 (UTC)
- This stripping of journal names down to impenetrable crap like Expert Opin Pharmacother, and author names into run-together gibberish like "Jones HB, Smith TE, Garcia XJ", has to stop. This is not
your journal, and WP certainly does not have to follow the excessively compressed and
jargonistic style preferred by the journals you read. See the essay
WP:Specialist style fallacy for an analysis of why attempts to impose on Wikipedia some style quirk from specialist publications is usually both a bad idea and a failure. —
SMcCandlish ☺
☏
¢ ≽ʌⱷ҅ᴥⱷʌ≼
20:51, 8 May 2014 (UTC)
- It is not your journal either. Are you suggesting that we repeal WP:CITEVAR? And why is "Jones, H.B., Smith, T.E.; Garcia, X.J." necessarily better than "Jones HB, Smith TE, Garcia XJ"? The later is much cleaner and easier to read. Boghog ( talk) 20:59, 8 May 2014 (UTC)
May 2014
![]() Hello, I'm
BracketBot. I have automatically detected that
your edit to
Steroid may have broken the
syntax by modifying 1 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
Hello, I'm
BracketBot. I have automatically detected that
your edit to
Steroid may have broken the
syntax by modifying 1 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
- List of unpaired brackets remaining on the page:
- year = 2011 | publisher = Wiley | location = Hoboken | isbn = 978-0-470-66084-3 }}</ref>{rp|10–19}} and the [[anti-inflammatory]] drug [[dexamethasone]].<ref name="pmid16236742">{{cite journal |
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, BracketBot ( talk) 06:18, 12 May 2014 (UTC)
Your submission at Articles for creation: Iminosugar has been accepted

The article has been assessed as Start-Class, which is recorded on the article's talk page. You may like to take a look at the grading scheme to see how you can improve the article.
You are more than welcome to continue making quality contributions to Wikipedia. Note that because you are a logged-in user, you can create articles yourself, and don't have to post a request. However, you may continue submitting work to Articles for Creation if you prefer.
- If you have any questions, you are welcome to ask at the help desk.
- If you would like to help us improve this process, please consider .
Thank you for helping improve Wikipedia!
— Anne Delong ( talk) 03:25, 18 May 2014 (UTC)DTX2
I looked at homoloGene and Blast. It seemed to me that the Blast comparison of proteins was easier to read, and, perhaps, easier for a broader audience to interpret quickly. I was going to suggest that a direct link to protein Blast in the box would be useful. But I did not want to bother your infobox. Even though the page is about the DTX2 gene, I was looking at the human protein.
Also, I look at the page and see one sentence, and a bunch of "further readings". The page cries out for some explanation for the general user. What do all those references "mean"?
The experts are already using the specific databases and tools, they hardly need an encyclopedia article.
What could you say about specific proteins that would help a general user of Wikipedia, not an expert? Or, how does expert explain the importance and relevance of DTX2?
I think Wikipedia has a responsibility to make every page accessible to everyone.
Kind regards, RC711 ( talk) 19:08, 20 May 2014 (UTC)
- Hi. Thanks for your note. Concerning Blast Q86UW9 vs. HomoloGene, the former takes over a minute to load, while the later is almost instantaneous. Many users have a short attention span. Also the former is a mix of homo and paralogs (between and within species) and contains duplicates, whereas the later is restricted to homologous (between species). Ideally there should also be a paralog link. Perhaps the KEGG SSDB Database might work.
- The intention of the further reading sections in bot generated Gene Wiki articles was to provide notability and encourage human editors to expand the articles by providing background references some of which would hopefully be moved in-line like I just did in this .
- Finally a removed a bunch of further reading citations in the DTX2 that were not specific to that gene. Hope this helps. Boghog ( talk) 20:05, 20 May 2014 (UTC)
DTX2
Thanks for the quick update on DTX2. It is much clearer now. I could not have done it myself; those few extra words of explanation make it much better. I appreciate your hard work. RC711 ( talk) 20:01, 20 May 2014 (UTC)
- Thanks and thank you for the suggestions. Cheers. Boghog ( talk) 20:07, 20 May 2014 (UTC)
Diberri
Hi Boghog,
mw:User:Mvolz is a new community developer whose project is coding a citation auto-filler system for VisualEditor. You can see what she's planning at mw:Cite-from-id. I'm hoping to get Diberri-style PMID (etc.) filling in this as well, because I think it's important to our science-related editors. If you're interested in this, I'm sure she'd be happy to hear from you or to have someone else to bounce ideas off of. Whatamidoing (WMF) ( talk) 21:25, 20 May 2014 (UTC)
- Hi Whatamidoing. Thanks for the heads up. I definitely will follow the development of this project and participate in the discussion. It appears that the cite-from-id service will also be used by RefToolbar in addition to VisualEditor. Hence I would like to see is some flexibility in how the citations are formatted (e.g., support for "m(edical)cite" style templates). Cheers.
Boghog (
talk)
04:51, 21 May 2014 (UTC)
- My first goal is "working at all". ;-) WhatamIdoing ( talk) 05:29, 21 May 2014 (UTC)
Melanopsin
While looking at new edits to Melanopsin I noticed that the addition of info about researcher Samer Hattar was added by user:Shattar. Glancing down the history, I saw your edit summary: "enough history, return the focus back to the protein, not the researchers", Diff. The new edits are similar to the one(s) you reverted way back when. Perhaps you'd like to check this out. -- Hordaland ( talk) 01:30, 23 May 2014 (UTC)
-
 Fixed −
Seppi333 (
Insert 2¢ |
Maintained)
02:49, 23 May 2014 (UTC)
Fixed −
Seppi333 (
Insert 2¢ |
Maintained)
02:49, 23 May 2014 (UTC)
- Thank for the heads up Hordaland and Seppi333 for fixing the problem. WP:COATRACK is a good description of these types of edits. There also appears to be a WP:COI. Boghog ( talk) 04:32, 23 May 2014 (UTC)
BAGBot: Your bot request BogBot 5
Someone has marked Wikipedia:Bots/Requests for approval/BogBot 5 as needing your input. Please visit that page to reply to the requests. Thanks! AnomieBOT ⚡ 00:12, 26 May 2014 (UTC) To opt out of these notifications, place {{bots|optout=operatorassistanceneeded}} anywhere on this page.
AEBP2
Thank you for your kind assistance with improving and clarifying the AEBP2 article. Your help is most appreciated. PEGLEG3 ( talk) 12:09, 27 May 2014 (UTC)
Disambiguation link notification for June 3
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited Circadian clock, you added a link pointing to the disambiguation page ATP ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:51, 3 June 2014 (UTC)
Hi. I'm wondering if there is some reason for the various appearances: BMAL1, Bmal1 and Bmal1. I'm tempted to change them all to BMAL1, but there might be some reason for the variation. (?) Thanks, Hordaland ( talk) 16:46, 3 June 2014 (UTC)
- By convention, human genes are written with all letters in uppercase (BMAL1), non-human mammals, only the first letter is capitalized (Bmal1), all other species are in lowercase (bmal1). In general, gene symbols should be italicized. Hence it might be appropriate to have a mix of gene cases, if human diseases and animal studies are included in the same article. On the other hand, if the species is not specifically mentioned in an article, it is probably better to standardize the capitalization to all upper case (human). Cheers. Boghog ( talk) 17:16, 3 June 2014 (UTC)
Not touching anything at the Nat Product page
…not synthesis or anything, until the matter of the "course of the ship" is settled. And I heard and acknowledged Smoke, both halves of what he said. I see one ambiguous statement, leaning as much my way as yours. So there is no decision possible based on it. You are the one that wants to run with half a vote, in your prescribed direction. That, my friend defines self-serving. I am staying put until (given summer, and busy life) others have time to respond. The WP standard seems (with merger proposals, and other higher order proposed changes), to allow 30 days. That is at least what I am looking at. And sorry, no longer playing your WP:THIS and WP:THAT game. You do not seem to care to hear that you, perhaps unconsciously but nevertheless self-servingly, limit your purview when you do this, so I've ceased arguing in such ways with you. Good news, no more walls of text. Le Prof Leprof 7272 ( talk) 08:45, 12 June 2014 (UTC)
Too much to ask
but I will do it to avoid confusion when folks start responding. I would appreciate it if you would, in good faith, sandbox the last three edits that you did, since they directly bear on the questions of the discussion that were ongoing, and then revert to the version that I linked to and discussed (immediately prior to your last three edits). If you are confident in your case, you should be confident without any changes, with the form that stood for the 6 mos. of your absence. Reverting will ensure that people see what I was describing at the time, and not some muddle of old and new. Don't expect it, but in integrity, and to give you the chance, I have to ask. If you do not, say by end of day Saturday, I will have to review my opening description of the history and argument, and make sure it still makes sense (fixing links and anything else that your edit will have muddied). But as I said, the simplest is to return the matter, in good faith, to what I was talking about before those three edits. Le Prof Leprof 7272 ( talk) 08:56, 12 June 2014 (UTC)
- I really do not see how my recent edits would fundamentally change the course of the discussion. If you disagree, you can always provide a link to the previous version on the article's talk page. In the meantime, this article is like any other, and unless there is persistent edit warring or vandalism, anyone should feel free to edit it. Boghog ( talk) 20:34, 12 June 2014 (UTC)
- Thank you for the clarity of your response. I will do what I have to do, to make this work. Le Prof Leprof 7272 ( talk) 04:09, 13 June 2014 (UTC)
PK pathway enzymes
Hey Boghog, I figured it wouldn't hurt to put this in the pharmacokinetics section of the amph page, but I'm not sure what the common abbreviations are for these enzymes. I'd rather not put the full names in the drugbox.
Benzoic acid is metabolized by butyrate-CoA ligase into an intermediate product, benzoyl-CoA, [1] which is then metabolized by glycine N-acyltransferase into hippuric acid. [2]
I'm guessing the 2nd is GLYAT, but I have no clue about the ligase. Would you happen to know of a contraction or where to find one for it? Seppi333 ( Insert 2¢ | Maintained) 01:21, 8 May 2014 (UTC)
References
-
^
"butyrate-CoA ligase". BRENDA. Technische Universität Braunschweig. Retrieved 7 May 2014.
{{ cite web}}:|section=ignored ( help) -
^
"glycine N-acyltransferase". BRENDA. Technische Universität Braunschweig. Retrieved 7 May 2014.
{{ cite web}}:|section=ignored ( help)
- Hi Seppi. PMID 12616642 refers to this enzyme as a xenobiotic/medium-chain fatty acid:CoA ligase or XM-ligase for short. Benzoic acid in this context is a xenobiotic, hence I believe XM-ligase would be an appropriate abbreviation. For CoA:glycine N-acyltransferase, ACGNAT ( PMID 7833837) would be an appropriate abbreviation. Boghog ( talk) 20:00, 8 May 2014 (UTC)
- Hey again Boghog; I just updated the articles with that information, but I noticed glycine N-acyltransferase has a duplicate at GLYAT while making redirects from the shortened names you provided. Should we merge these two? Seppi333 ( Insert 2¢ | Maintained) 23:28, 21 May 2014 (UTC)
- Sorry for not responding sooner but I have been really busy in real life. It is appropriate to merge protein and enzyme articles if there is a one-to-one correspondence (i.e., only one human gene that encodes an enzyme that catalyzes a specific rxn as designated by an EC number). Digging into this further, there are the following enzyme/protein correspondences:
- Whereas the UniProt entry for GLYAT (
Q6IB77) lists EC numbers 2.3.1.13 and 2.3.1.71 as entries. Because there is not a one-to-one correspondence, I think it would be better not to merge
glycine N-acyltransferase (EC 2.3.1.13) with
GLYAT.
Boghog (
talk)
16:28, 26 May 2014 (UTC)
- Ah, alright. Thanks for pointing that out. Seppi333 ( Insert 2¢ | Maintained) 00:48, 28 May 2014 (UTC)
- One other thing - since you've made major contributions to the article (half the images were made by you) and have already done a ton of work to help address issues at the FAC, would you be interested in being a co-nominator at the next FAC for
amphetamine? I figure you deserve credit ({{
FA user topicon}}) for all you've done to help attain the
 rating. In any event, I'm going to open the 3rd FAC for it in 2-3 days since I need to do copyedits to address Shudde's issues first.
rating. In any event, I'm going to open the 3rd FAC for it in 2-3 days since I need to do copyedits to address Shudde's issues first. - Best, Seppi333 ( Insert 2¢ | Maintained) 03:17, 23 May 2014 (UTC)
- Thanks for asking but I think you are overstating my contribution ;-). In any case, I might be interested in co-nominating the article, but this would obligate me to devote a significant amount of time to address reviewers comments, time that I do not have at the moment. If you could hold off on the nomination for a week or two, I would be able spend significantly more time on improving it. Cheers.
Boghog (
talk)
16:39, 26 May 2014 (UTC)
- I seriously mean it - your help has been indispensable in copyediting and expanding the chemistry portions of the article. Thanks to you, it puts chem sections in other drug articles to shame.
 The best I could have hoped for had I written it myself is to make it merely not suck, so I mean it when I say you've done a lot for the page!
The best I could have hoped for had I written it myself is to make it merely not suck, so I mean it when I say you've done a lot for the page!
- In any event, I actually wouldn't need or expect you to help out at the FAC with anything but technical issues related to the physical/chemical properties sections, since you're familiar with those sections and way more knowledgeable than me in that area. I'm perfectly fine with handling all the basic copyediting and technical issues in the remainder of the article myself. That said, feel free to address any issues outside phys/chem if feel comfortable editing the section and have the time. Seppi333 ( Insert 2¢ | Maintained) 00:48, 28 May 2014 (UTC)
- I seriously mean it - your help has been indispensable in copyediting and expanding the chemistry portions of the article. Thanks to you, it puts chem sections in other drug articles to shame.
- Thanks for asking but I think you are overstating my contribution ;-). In any case, I might be interested in co-nominating the article, but this would obligate me to devote a significant amount of time to address reviewers comments, time that I do not have at the moment. If you could hold off on the nomination for a week or two, I would be able spend significantly more time on improving it. Cheers.
Boghog (
talk)
16:39, 26 May 2014 (UTC)
Hey again Boghog. I've been off Wikipedia for nearly 2 weeks due to a broken laptop; I have a working one now though, so I intend to renominate the article at FAC very soon. Are you still interested in being a co-nominator? Seppi333 ( Insert 2¢ | Maintained) 22:08, 13 June 2014 (UTC)
- Sorry to hear about your lap top but I am glad your connected again. I have a bit more time now for the co-nomination, so bring it on! The biggest unsolved issue as I see it is to make some of the more technical sections more accessible. I will see what I can do to help. Boghog ( talk) 22:49, 13 June 2014 (UTC)
My closing word
Not engaging with you further at NP, sorry. In your most recent spate of unscrupulous actions, you have shredded any last basis here for AGF from me. I will work around you, if possible alongside you, but any chance for real discourse or collegiality is gone. Please do not post to me, or refer to me anywhere that you can help it. Cheers, best of fortunes in our shared, difficult day to day, as we try to do something real. Le Prof Leprof 7272 ( talk) 05:37, 14 June 2014 (UTC)
- @ Leprof 7272: unscrupulous actions? Is asking a response to a simple question unscrupulous? Boghog ( talk) 05:48, 14 June 2014 (UTC)
Hello, Boghog. Some time ago you made some improvements to the draft article above, and it turned out that it was a copy-paste remnant. Was there anything that you wanted to move over to the mainspace article before the draft is deleted? — Anne Delong ( talk) 01:37, 16 June 2014 (UTC)
- Hi Anne. Thanks for the heads up. I have compared the draft with the mainspace article and it doesn't look like there is anything additional that needs to be moved over. Please go ahead and delete the draft. Boghog ( talk) 05:59, 16 June 2014 (UTC)
NFkB in neurons
Hi Boghog,
In April of 2011 I updated the the section on NFkB in neurons and I see that user Neuroglider has made substantial changes to the section that I don't agree with. Neuroglider states that s/he talked with you on 11/23/2013, though I couldn't find the talk on either of your talk pages, about removal of synaptic components as gene targets of NFkB. Could you fill me in, or point me to the correct talk page? I feel that these should be in the article as the support the idea that NFkB is important in neurons, not just glia. Which brings me to my main point. I believe that the NFkB field in neuroscience does not hold to Neuroglider's views that NFkB is primarily important in glia and not in neurons. In 2011 I specificially removed sections stating these points as I believed them to not be representative of the thoughts of the field. Specifically Meffert et.al. 2003 (citation #7 in NFkB article) [1] demonstrated that NFkB could be activated in biochemically isolated synapses (synaptoneurosomes) and was unlikely to be confounded by glia. Additionally, there are many studies that support the idea that NFkB function is important in neurons, such as studies that show cell autonomous roles for NFkB in neurons related to synaptic function and regulation of genes that are also important to synaptic function. While Neuroglider does make a good point that we should all think about heterogeneity in our systems, I believe s/he has given too much weight to this idea and the paragraph is now larger than the rest of the section. I would like to rebut Neuroglider's addition to the section, but do not wish to turn the article into a discussion of the merits of Neuroglider's v.s. other perspective. While I do not agree with Neuroglider, I'd be okay with his/her opinions to remain, but with more balance. If you have any thoughts on the best way to proceed, please let me know.
P.S. Thank you for your time and effort looking after these wikipedia articles.
-
^ Meffert MK, Chang JM, Wiltgen BJ, Fanselow MS, Baltimore D (October 2003). "NF-kappa B functions in synaptic signaling and behavior". Nat. Neurosci. 6 (10): 1072–8.
doi:
10.1038/nn1110.
PMID
12947408.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link)
16:24, 16 June 2014 (UTC)Aventre3
- Hi @ Aventre3:. I believe that the link to the relevant discussion is here. I must admit that I am not an expert on neuronal molecular biology nor have I read these citations very carefully. My interest in NFkB mainly relates to its cross talk with nuclear receptors. I suggest that you continue the discussion with @ Neuroglider: at the above link. In cases where there may be controversy, it is especially important to cite secondary sources (please note PMID 12947408 is a primary source). I will follow the discussion and jump in if I think I have anything constructive to add. Cheers. Boghog ( talk) 18:26, 16 June 2014 (UTC)
Simple matter in re steroids
For you to consider. Colleague looked at WP steroids on an iphone, and said the article opened with three structures in a row, and it seemed off-putting as a way to open an article for general readers. I checked his report, and it is true. Because of the way the image markups are inserted relative to the lede, the article opens on small screen handhelds with the three images appearing 1-2-3 that show up to the right of the lede for a standard laptop or PC viewing. Your call, but perhaps the markups could be moved down a bit, with no change of appearance for computer viewing, but resolving the negative reported by this handheld user. No RSVP necessary. Completely your call. Cheers. Le Prof Leprof 7272 ( talk) 02:10, 17 June 2014 (UTC)
I am reverting you at NP Talk because you changed more than the shading
You also changed my words in several of MY TALK ENTRIES.
If you want to revert something in particular, it is YOUR responsibility to do it selectively, and not change my words. You have no right to change my words.
And if you had a modicum of decency, you would also leave the shading in place, as it is intended to call attention to the words of individual other than ours.
If you revert again, I will ask an admin to adjudicate.
Le Prof Leprof 7272 ( talk) 15:03, 19 June 2014 (UTC)
- @
Leprof 7272: First of all, you
redactedrefactored other peoples comments in the same edit that you changed your own comments. If you don't want edits to your own comments reverted, don't combine edits to your own comments with edits to other editors comments. Better still, don'tredactrefactor other editors comment to begin with. Second, I did restore the edits to your comments in the next edit. Please more carefully look at the edit history before you jump to conclusions and assume good faith. Third, your excessively long talk page post are drowning out other voices. If you posts were more concise, it would not be necessary to highlight text. Finally mature editors try to work out differences between themselves first. Only in extreme cases is admin intervention required. Boghog ( talk) 15:24, 19 June 2014 (UTC)
- Redact means to "edit (text) for publication. censor or obscure (part of a text) for legal or security purposes." I redacted noone's text, and in shading, censored and obscured nothing. If you look closely, you did not capture all of my changes to my own text. I did look carefully at the edit history, and changed it back as I had it the first time. Go ask Pad and Smokefoot, et al, if they object. It is not your right to object for them. Leprof 7272 ( talk) 15:27, 19 June 2014 (UTC)
- @ Leprof 7272: You have it completely backwards. It is your responsibility to first ask if it is OK, not my responsibility to ask for you. Boghog ( talk) 15:31, 19 June 2014 (UTC)
- Neither Smoke nor Pad have issues, and I have asked Doc for his permission. Stop warring with me. The shading (highlighting) of other editors' comments serves a clear positive purpose. Your response is knee-jerk negative against me. Plese, stop warring. As you say, focus on content. Leprof 7272 ( talk) 15:34, 19 June 2014 (UTC)
- @ Leprof 7272: I stand behind what I state above. Stop highlighting other editor's comments without their permission. Boghog ( talk) 15:38, 19 June 2014 (UTC)
- And I by mine. The only thing approaching an edit of another's text that can be asserted, is my adding another editor's name in square brackets, where a less than careful interspersed reply from you had made unclear who had authored the original comment. Otherwise, apart from shading (highlighting), there is no change, and highlighting of a whole section, without change in content or emphasis, and indeed with elevation of attention, cannot be considered a redaction. This is a redaction. Permissions are in place as they need be. Others see merit and AGF. Le Prof Leprof 7272 ( talk) 15:49, 19 June 2014 (UTC)
- And I by mine. The only thing approaching an edit of another's text that can be asserted, is my adding another editor's name in square brackets, where a less than careful interspersed reply from you had made unclear who had authored the original comment. Otherwise, apart from shading (highlighting), there is no change, and highlighting of a whole section, without change in content or emphasis, and indeed with elevation of attention, cannot be considered a redaction. This is a
- @ Leprof 7272: The only significant change that I missed was Pharmagognosy to Pharmacogn abbreviation. Stop highlighting other editor's comments without their permission. Boghog ( talk) 15:53, 19 June 2014 (UTC)
- Reread what I just said. If you revert again, you will be subverting not only my will, but others, and I will involve adjudication. There is a page of content issues in play. Deal with the real matter at hand. Le Prof Leprof 7272 ( talk) 15:58, 19 June 2014 (UTC)
- @ Leprof 7272: One editor already objected to your highlighting of their comments. Did you receive permission from the other editors? Your selective highlighting of other user comments (unless you quote them in your own post) is bad practice and needs to stop. Boghog ( talk) 16:04, 19 June 2014 (UTC)
- Leprof, rather than
WP:REDACT, I think what Boghog means is
WP:REFACTOR.
Flyer22 (
talk)
16:24, 19 June 2014 (UTC)
- Thanks Flyer22, that is indeed what I meant. Boghog ( talk) 16:58, 19 June 2014 (UTC)
- Leprof, rather than
WP:REDACT, I think what Boghog means is
WP:REFACTOR.
Flyer22 (
talk)
16:24, 19 June 2014 (UTC)
- You're welcome. Wikipedia:Talk page guidelines#Others' comments states: " Refactoring and archiving are still appropriate, but should be done with courtesy and reversed on protest." Flyer22 ( talk) 18:41, 19 June 2014 (UTC)
Got your note and
first, no problem about it being the weekend before a response to the outline. But, did it not strike you that I might find a bit odd, so many Talk edits fitting schedule, but no "the outline looks fine, go get started"? And, for all that, in future, don't presume regarding your refactoring either (or whatever it's called). Whatever my sins, your rejiggering the whole of the Talk — appearance, order of entries, what appears on the current page, etc.—was not appreciated mate. For one, I've several incoming links, now broken, to the archived Talk. (For another, your perceived "I'm in charge here" persona is still an issue.) Effectively reading one another's minds—a point we're clearly not at yet. Nor is empathy a natural product between us. Ask first, still. (And so, to be clear, I expect to see Talk about the NP outline/changes, before any editing starts.) No reply, please. Respectful actions, enough. Le Prof Leprof 7272 ( talk) 03:33, 21 June 2014 (UTC)
It is recommended to archive or refactor a page either when it exceeds 75 KB, or has more than 10 main sections.
— WP:TALKCOND- At the time that I had archived the talk page, it had grown to 133 KB and had 15 main sections. Based on length alone, it was time to archive. Furthermore TenOfAllTrades had specifically stated that (s)he thought that the talk page was a mess and needed to be cleaned up. Padillah agreed and indicated that (s)he would archive some of the discussion. If I had not archived the discussion, Padillah probably would have done it for me. Breaking links is an unfortunate side effect of archiving talk page discussions. However one of the reasons there has been very limited engagement from other editors was it was becoming impossible for anyone to follow the discussion. Creating links into this complex discussion is not likely to generate a much of a response. I was also having trouble reading the talk page and as a consequence, it was interfering with our recent productive discussions and improving the article. It is time to put the past acrimonious discussion behind us and move on to improving the article.
Boghog (
talk)
06:46, 21 June 2014 (UTC)
- You seem particulary incapable of perceiving what sort of treatment of others will put an end to acrimony. Try respect. Asking prior to acting significantly is respectful. Acting without asking, is not. Wrong impetus mate. I have not another word to say on this. Le Prof Leprof 7272 ( talk) 00:32, 22 June 2014 (UTC)
If my mind serves me
here are some ketone-containing drugs: acetohexamide, befunolol, daunorubicin, ethacrynic acid, haloperidol, ketoprofen, metyrapone, loxoprofen, naloxone, and naltrexone. Le Prof Leprof 7272 ( talk) 04:27, 21 June 2014 (UTC)
- Not sure what you are getting at here or if this message was meant for me. Are you suggesting we create a new Category:ketone containing drugs or mention these in the ketone article? Boghog ( talk) 08:19, 21 June 2014 (UTC)
- Misplaced, and since moved. You can delete this entry if you wish. Cheers. Le Prof Leprof 7272 ( talk) 00:33, 22 June 2014 (UTC)
IL-23 subunit alpha
I think that the IL-23 subunit alpha page needs to be reduced. Right now it is a mixture of pages about IL-23 subunit alpha and the IL-23 heterodimeric cytokine. There should be a separate page for IL-23 subunit alpha, and for the cytokine itself. The page re the subunit will by definition be shorter because little is known about its function, outside of functioning as a heterodimer. What do you think? That's how it is now set up for IL-12, you can read about the cytokine IL-12 and then about each of the subunits separately. — Preceding unsigned comment added by Exiledscientist ( talk • contribs) 19:33, 26 June 2014 (UTC)
- Agreed. Interleukin 23 which is now a redirect to Interleukin 23 subunit alpha should be converted into its own article. BTW, thanks for your contributions. You clearly know the subject and I do appreciate your contributions. Cheers. Boghog ( talk) 19:43, 26 June 2014 (UTC)
- Per your suggestion, I went ahead and converted Interleukin 23 to an stand alone article. Please edit and expand as you see fit. Cheers. Boghog ( talk) 20:00, 26 June 2014 (UTC)
Disambiguation link notification for June 27
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited Interleukin 23, you added a link pointing to the disambiguation page IL-12 ( check to confirm | fix with Dab solver). Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:49, 27 June 2014 (UTC)
CASK article
Hi, You removed a lot of my entries from this page. /info/en/?search=CASK This information is critical for children (like mine) who suffer from CASK gene defects. How can we manage that this information reaches the general public and the parents of the children affected with this rare disease. Can you provide me with some tips on how to integrate this information more wisely into the article ? — Preceding unsigned comment added by Reachme A ( talk • contribs) 14:59, 2 July 2014 (UTC)
- Hi. Thanks for your note. Simply stating a journal article is about the subject of a Wikipedia article is generally not sufficient reason to include it. One also needs to understand what the journal article says about the subject and write some prose to go along with the citation. To start with, I will re-add the citations into the further reading section. As I have time, I will try to integrate a few of these papers to better illustrate what I mean. Cheers. Boghog ( talk) 15:36, 2 July 2014 (UTC)
Once again, no response from you at Talk, no apparent time taken to appreciate and discuss the point of another's edit
…no questioning, just your large scale reversion and so significant further wasted time trying to interact with you. The point of my edit, as stated, was to leave in place an actual outline that we both and others can view and discuss—and not an amalgam of one editor's writing, and the remainder a partial outline it overlays.
By reverting, you nullified/broke a link in Talk to this common outline, so I and others could easily find it. IF THIS POSTED AMALGAM OF TEXT AND OUTLINE WAS INTENDED AS YOUR SANDBOX, YOU SHOULD HAVE LEFT IT IN YOUR SANDBOXES. My edit was a good faith effort to create the joint outline I have asked you for, for weeks. IF MY DERIVATION OF OUTLINE BULLET POINTS FROM YOUR TEXT, THERE, WAS NOT PERFECT, YOU COULD HAVE EDITED IT. ONCE AGAIN, WITH SWEEP OF HAND YOU SET ASIDE AN HOUR OF WORK WITHOUT REGARD. (Note, I moved your text to talk, carefully, so nothing in terms of time or effort would be lost.)
If you simply cannot work collegially, allowing others to make changes, in this case, leaving somewhere an outline of the NP article that all can see and work from—if it always has to be entirely your way, then count me out on any collaborative editing with you.
I have nothing more to say on the subject. If you cannot meet halfway, if you cannot leave an outline in place, we will proceed fully independently, and contentiously as necessary, with adjudication at every turn, to continue working on the article we each deem important. No more wasted time with you, or blatant disrespect from you. Last straw. Le Prof Leprof 7272 ( talk) 05:06, 15 July 2014 (UTC)
- @ Leprof 7272: With all due respect, you created an incomprehensible mess on the Talk:Natural product page. Because of the length and complexity of the outline that was starting to overwhelming the talk page, it is much cleaner to move the outline to a subdirectory leaving behind a link. The advantage of the subdirectory is that we can use the normal section heading syntax and a single {{ reflist}} tag to display the citations. The section headings make it much easier to navigate and to edit the outline. Furthermore it is much easier to track the revision history if the outline is separated from the discussion. Boghog ( talk) 05:30, 15 July 2014 (UTC)
- I simply can no longer tell if you are completely rude, or near completely communicatively incapable. PLEASE, reread the foregoing, noting the following terms: " …leave in place an actual outline… …not an amalgam … IF THIS POSTED AMALGAM … leaving somewhere in place an outline … ...IF MY DERIVATION OF OUTLINE BULLET POINTS FROM YOUR TEXT, THERE, WAS NOT PERFECT, YOU COULD HAVE EDITED IT… "
- Your reply ignores the thrust, and responds with matters not contested (and in fact granted). Nowhere did I contest the move of the outline. INDEED, I linked to the outline you relocated, so I clearly have affirmed the step you took in creating the new location. I contest the amalgamation of a general outline intended for all, with your individual writing and you must know this, because this is what you reverted. PLEASE, READ AND REPLY TO WHAT I ACTUALLY WROTE, AND NOT TO A THOUGHT THAT COMES INTO YOUR MIND THE SPLIT SECOND YOU BEGIN TO READ. BL: Un-amalgamated outline—where? Le Prof Leprof 7272 ( talk) 05:49, 15 July 2014 (UTC)
- @ Leprof 7272: Why do you make things so complicated? Sometimes outlines become overly abstract and in these situations it helps to flesh out the outline to determine if the outline is workable. Hence my creation of the amalgam was intentional. Why can't we continue to work on the amalgam? Boghog ( talk) 06:00, 15 July 2014 (UTC)
- That an amalgam is simpler than the outline is nonsense; easier for you perhaps, better for you perhaps, yes. Simpler, no. Considerate of others, certainly not. So if you cannot see how ones personal editing would intrude upon an outline document intended for general, overall, unbiased, all-editor friendly discussion, there is nothing to say. I ask again, as I have for weeks now, where is the un-amalgamated outline that all can discuss and work on together (leaving your editing in your sandbox)? No more argument about this, if you cannot yield, you cannot, and I will not waste another moment of either of our time. Le Prof Leprof 7272 ( talk) 06:07, 15 July 2014 (UTC)
- @ Leprof 7272: Your outline was also an amalgam with comments interspersed throughout. The prose essentially is just a more extensive comment. And how is prose any less friendly than comments? I added the prose in direct response to your comment:
Clearly, later full sections of information cannot be subsumed into the brief, early, organizational, bullet-style Classes section
- I wanted to demonstrate one can write this section without bullet points. One last point: There is a completely clean outline minus comments and prose at the top of the Talk:Natural_product/Outline page that is labeled "contents". Boghog ( talk) 06:46, 15 July 2014 (UTC)
- Since you seem incapable of grasping the plain distinction I am making, and of taking the real temperature of this interaction, let me end it here: feel free to delete all explanatory comments in the earlier draft outline that I offered, to make it fulfill your perception of a simple bare, decimal type outline that all users can add to or edit. (The autogenerated TOC is not such.) Please let me know where this simple outline resides, or tell me you will not do so. (As simple as that; "I will not do so.") I will not comment further here, or there, unless the decision is to work together. Choose one place at the Talk page, to report the location of the simple outline you create (or report that you are not willing to work with me). Done here. Le Prof
Leprof 7272 (
talk)
07:05, 15 July 2014 (UTC)
- I am working with my nephew on Latin. It is interesting how very descriptive and revealing old languages can be. For instance, L. subterfugio means literally, to flee or escape under…
Leprof 7272 (
talk)
07:09, 15 July 2014 (UTC)
- @
Leprof 7272: You have not explained why editors cannot work on
Talk:Natural_product/Outline. There is no difference in the mechanics of editing this page as opposed to the talk page. In contrast, interactively working on a large complex outline on a talk page is a really bad idea. It amount to refactoring other edit's comments and complicates the revision history. Talk pages were simply not designed for this. It is much better to place the draft outline in a subdirectory which separates comments from the collaboratively worked on proposal. Furthermore separating the outline from comments and draft prose is inefficient. One gets a birds eye view from the "contents" outline at the top with more details below.
Boghog (
talk)
07:25, 15 July 2014 (UTC)
- quam subterfugium. Change of location long granted, as you well know. Le Prof
Leprof 7272 (
talk)
07:32, 15 July 2014 (UTC)
- @
Leprof 7272: OK, good, we can move the location of the outline. We can also
collapse the prose if you like.
Boghog (
talk)
07:40, 15 July 2014 (UTC)
- Reproduced from article Talk page, please reply only there Le Prof
Leprof 7272 (
talk)
07:25, 15 July 2014 (UTC)
- Replied there. Boghog ( talk) 14:10, 15 July 2014 (UTC)
- Reproduced from article Talk page, please reply only there Le Prof
Leprof 7272 (
talk)
07:25, 15 July 2014 (UTC)
- @
Leprof 7272: OK, good, we can move the location of the outline. We can also
collapse the prose if you like.
Boghog (
talk)
07:40, 15 July 2014 (UTC)
- quam subterfugium. Change of location long granted, as you well know. Le Prof
Leprof 7272 (
talk)
07:32, 15 July 2014 (UTC)
- @
Leprof 7272: You have not explained why editors cannot work on
Talk:Natural_product/Outline. There is no difference in the mechanics of editing this page as opposed to the talk page. In contrast, interactively working on a large complex outline on a talk page is a really bad idea. It amount to refactoring other edit's comments and complicates the revision history. Talk pages were simply not designed for this. It is much better to place the draft outline in a subdirectory which separates comments from the collaboratively worked on proposal. Furthermore separating the outline from comments and draft prose is inefficient. One gets a birds eye view from the "contents" outline at the top with more details below.
Boghog (
talk)
07:25, 15 July 2014 (UTC)
- I am working with my nephew on Latin. It is interesting how very descriptive and revealing old languages can be. For instance, L. subterfugio means literally, to flee or escape under…
Leprof 7272 (
talk)
07:09, 15 July 2014 (UTC)
- Since you seem incapable of grasping the plain distinction I am making, and of taking the real temperature of this interaction, let me end it here: feel free to delete all explanatory comments in the earlier draft outline that I offered, to make it fulfill your perception of a simple bare, decimal type outline that all users can add to or edit. (The autogenerated TOC is not such.) Please let me know where this simple outline resides, or tell me you will not do so. (As simple as that; "I will not do so.") I will not comment further here, or there, unless the decision is to work together. Choose one place at the Talk page, to report the location of the simple outline you create (or report that you are not willing to work with me). Done here. Le Prof
Leprof 7272 (
talk)
07:05, 15 July 2014 (UTC)
August 2014
![]() Hello, I'm
BracketBot. I have automatically detected that
your edit to
TAS1R2 may have broken the
syntax by modifying 2 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
Hello, I'm
BracketBot. I have automatically detected that
your edit to
TAS1R2 may have broken the
syntax by modifying 2 "{}"s. If you have, don't worry: just
edit the page again to fix it. If I misunderstood what happened, or if you have any questions, you can leave a message on
my operator's talk page.
- List of unpaired brackets remaining on the page:
- 3 | pages = 293–301 | year = 2003 | pmid = 12581520 | doi = 10.1016/S0092-8674(03)00071-0 }}</ref>}}
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, BracketBot ( talk) 09:20, 3 August 2014 (UTC)
Reference Errors on 3 August
![]() Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
- On the 5-HT2A receptor page, your edit caused a DOI error ( help) and a cite error ( help). ( Fix | Ask for help)
Please check this page and fix the errors highlighted. If you think this is a false positive, you can report it to my operator. Thanks, ReferenceBot ( talk) 00:25, 4 August 2014 (UTC)
Minor typo in image
Hey Boghog, I noticed a small typo in an image you uploaded: File:Inverse_agonist_2.png (activty - in blue text). It's not really a big deal since the intended word is obvious, but I figured I should point it out. Best, Seppi333 ( Insert 2¢ | Maintained) 18:28, 10 August 2014 (UTC)
-
 Fixed Hi Seppi. You have sharp eyes. I have fixed the typo and created a higher quality svg version here:
File:Inverse agonist 3.svg. Thanks for the heads up.
Boghog (
talk)
19:47, 10 August 2014 (UTC)
Fixed Hi Seppi. You have sharp eyes. I have fixed the typo and created a higher quality svg version here:
File:Inverse agonist 3.svg. Thanks for the heads up.
Boghog (
talk)
19:47, 10 August 2014 (UTC)
- Looks great - noticeably sharper than the png version. Seppi333 ( Insert 2¢ | Maintained) 02:14, 12 August 2014 (UTC)
Image for signaling cascade in addiction
References
- ^
a
b
c Renthal W, Nestler EJ (September 2009).
"Chromatin regulation in drug addiction and depression". Dialogues in Clinical Neuroscience. 11 (3): 257–268.
doi:
10.31887/DCNS.2009.11.3/wrenthal.
PMC
2834246.
PMID
19877494.
[Psychostimulants] increase cAMP levels in striatum, which activates protein kinase A (PKA) and leads to phosphorylation of its targets. This includes the cAMP response element binding protein (CREB), the phosphorylation of which induces its association with the histone acetyltransferase, CREB binding protein (CBP) to acetylate histones and facilitate gene activation. This is known to occur on many genes including fosB and c-fos in response to psychostimulant exposure. ΔFosB is also upregulated by chronic psychostimulant treatments, and is known to activate certain genes (eg, cdk5) and repress others (eg, c-fos) where it recruits HDAC1 as a corepressor. ... Chronic exposure to psychostimulants increases glutamatergic [signaling] from the prefrontal cortex to the NAc. Glutamatergic signaling elevates Ca2+ levels in NAc postsynaptic elements where it activates CaMK (calcium/calmodulin protein kinases) signaling, which, in addition to phosphorylating CREB, also phosphorylates HDAC5.
Figure 2: Psychostimulant-induced signaling events -
^ Broussard JI (January 2012).
"Co-transmission of dopamine and glutamate". The Journal of General Physiology. 139 (1): 93–96.
doi:
10.1085/jgp.201110659.
PMC
3250102.
PMID
22200950.
Coincident and convergent input often induces plasticity on a postsynaptic neuron. The NAc integrates processed information about the environment from basolateral amygdala, hippocampus, and prefrontal cortex (PFC), as well as projections from midbrain dopamine neurons. Previous studies have demonstrated how dopamine modulates this integrative process. For example, high frequency stimulation potentiates hippocampal inputs to the NAc while simultaneously depressing PFC synapses (Goto and Grace, 2005). The converse was also shown to be true; stimulation at PFC potentiates PFC–NAc synapses but depresses hippocampal–NAc synapses. In light of the new functional evidence of midbrain dopamine/glutamate co-transmission (references above), new experiments of NAc function will have to test whether midbrain glutamatergic inputs bias or filter either limbic or cortical inputs to guide goal-directed behavior.
-
^ Kanehisa Laboratories (10 October 2014).
"Amphetamine – Homo sapiens (human)". KEGG Pathway. Retrieved 31 October 2014.
Most addictive drugs increase extracellular concentrations of dopamine (DA) in nucleus accumbens (NAc) and medial prefrontal cortex (mPFC), projection areas of mesocorticolimbic DA neurons and key components of the "brain reward circuit". Amphetamine achieves this elevation in extracellular levels of DA by promoting efflux from synaptic terminals. ... Chronic exposure to amphetamine induces a unique transcription factor delta FosB, which plays an essential role in long-term adaptive changes in the brain.
- ^ Cadet JL, Brannock C, Jayanthi S, Krasnova IN (2015). "Transcriptional and epigenetic substrates of methamphetamine addiction and withdrawal: evidence from a long-access self-administration model in the rat". Molecular Neurobiology. 51 (2): 696–717 ( Figure 1). doi: 10.1007/s12035-014-8776-8. PMC 4359351. PMID 24939695.
- ^
a
b
c Robison AJ, Nestler EJ (November 2011).
"Transcriptional and epigenetic mechanisms of addiction". Nature Reviews Neuroscience. 12 (11): 623–637.
doi:
10.1038/nrn3111.
PMC
3272277.
PMID
21989194.
ΔFosB serves as one of the master control proteins governing this structural plasticity. ... ΔFosB also represses G9a expression, leading to reduced repressive histone methylation at the cdk5 gene. The net result is gene activation and increased CDK5 expression. ... In contrast, ΔFosB binds to the c-fos gene and recruits several co-repressors, including HDAC1 (histone deacetylase 1) and SIRT 1 (sirtuin 1). ... The net result is c-fos gene repression.
Figure 4: Epigenetic basis of drug regulation of gene expression - ^
a
b
c Nestler EJ (December 2012).
"Transcriptional mechanisms of drug addiction". Clinical Psychopharmacology and Neuroscience. 10 (3): 136–143.
doi:
10.9758/cpn.2012.10.3.136.
PMC
3569166.
PMID
23430970.
The 35-37 kD ΔFosB isoforms accumulate with chronic drug exposure due to their extraordinarily long half-lives. ... As a result of its stability, the ΔFosB protein persists in neurons for at least several weeks after cessation of drug exposure. ... ΔFosB overexpression in nucleus accumbens induces NFκB ... In contrast, the ability of ΔFosB to repress the c-Fos gene occurs in concert with the recruitment of a histone deacetylase and presumably several other repressive proteins such as a repressive histone methyltransferase
-
^ Nestler EJ (October 2008).
"Transcriptional mechanisms of addiction: Role of ΔFosB". Philosophical Transactions of the Royal Society B: Biological Sciences. 363 (1507): 3245–3255.
doi:
10.1098/rstb.2008.0067.
PMC
2607320.
PMID
18640924.
Recent evidence has shown that ΔFosB also represses the c-fos gene that helps create the molecular switch—from the induction of several short-lived Fos family proteins after acute drug exposure to the predominant accumulation of ΔFosB after chronic drug exposure
-
^ (Text color) Transcription factors
 |
Hey Boghog. I was wondering if you'd happen to know of any any information resources (e.g., databases) that would have content on on signaling cascades / communication networks between receptors and gene transcription factors.
I'm working on an image ( File:ΔFosB.svg - a DA neuron/synapse in the nucleus accumbens) for the addiction section in amphetamine at the moment to illustrate how the NDMA receptors, calcium channels, and ΔFosB are related, since it's not entirely apparent in the text. This KEGG diagram (the bottom pathway) and my molecular neuropharmacology textbook are the only 2 resources I'm working off of at the moment. Unlike the KEGG pathway, I'm just doing the synaptic cleft and postsynaptic neuron since the schematic for the presynaptic half is exactly what's shown in pharmacodynamics. I know all 5 DA receptors are present in accumbens DA neurons, so I'm planning to add the 4 other DA receptors into the KEGG schematic that I recreate in order to show their influence on the cAMP cascade. Seppi333 ( Insert 2¢ | Maintained) 02:50, 17 June 2014 (UTC)
- Also if you have any suggestions or feedback for me on the diagram itself, feel free to chime in.
 Seppi333 (
Insert 2¢ |
Maintained)
03:42, 17 June 2014 (UTC)
Seppi333 (
Insert 2¢ |
Maintained)
03:42, 17 June 2014 (UTC)
- Update: I just realized that I saved a few papers on this topic on my website in a subpage that I forgot I made around 2 months ago. I unfortunately lost a large number of research papers when my laptop died, but at least I still have these signaling cascade models. In any event, I probably don't need more information resources for the model, but I'm always open to feedback! Seppi333 ( Insert 2¢ | Maintained) 00:42, 18 June 2014 (UTC)
- Hi Seppi. Sorry for not responding sooner. Your graphic looks great! One minor issue. Shouldn't the arrow between ΔFosB and DNA be reversed? According to the KEGG diagram, CREB regulates the expression of ΔFosB (i.e., CREB transcription factor → DNA (ΔFosB gene promoter) → ΔFosB protein). Also the transcription factor NFKB is a heterodimer of Fos and Jun. The KEGG diagram suggests that Fos (through the NFKB heterodimer) up regulates the expression of c-Fos. It appears that part of the diagram needs to be adjusted as well. Boghog ( talk) 17:27, 21 June 2014 (UTC)
- Ahh, alright. I'll make those tweaks when I work on the image within the next couple of days; several of the resources I used as a cross reference had a differing schematic inside the nucleus, so I wasn't really sure how to model that part. I don't expect to be done for another week or 2 since I'd like to look for additional reference models in reviews and on reactome first - assuming I can find anything relevant.
Thanks for the feedback by the way. Seppi333 ( Insert 2¢ | Maintained) 11:44, 22 June 2014 (UTC)
- Ahh, alright. I'll make those tweaks when I work on the image within the next couple of days; several of the resources I used as a cross reference had a differing schematic inside the nucleus, so I wasn't really sure how to model that part. I don't expect to be done for another week or 2 since I'd like to look for additional reference models in reviews and on reactome first - assuming I can find anything relevant.
@ Boghog: I've revised the image; I believe it's now consistent with the two KEGG diagrams and two literature reviews that I've listed in the description on the file page - File:ΔFosB.svg. The image legend is the same as KEGG's, with exception for the "+exp" and "-exp" terms which I use to denote gene expression and gene repression respectively. I'm hoping that I can make it more accessible to the average reader by removing all the SVG text and replacing it with annotated wikilinks in the finished version (similar to the amphetamine metabolites diagram). I'm also going to get some external feedback on the image's accessibility once I've finished drawing and annotating it. Consequently, it'll probably be at least another two weeks before I put it in an article.
In any event, you're more the expert in this area than I, so if there's anything you think I should add, edit, or remove from the current version of the image, please let me know! – Seppi333 ( Insert 2¢ | Maintained) 00:07, 12 July 2014 (UTC)
- It is looking even better! Two minor suggestions: Change GRIN3A → NMDAR and GRIA1 → AMPAR. GRIN3A and GRIA1 are the genes that encode subunits of NMDAR and AMPAR. The individual gene products themselves are not functional in isolation but require another subunit encoded by a distinct gene to form functional heterotetramers. Cheers. Boghog ( talk) 07:48, 12 July 2014 (UTC)
- @ Boghog: Thanks again for the feedback. More or less done now - just need to make cosmetic tweaks (the synaptic cleft and some minor tweaks) and replace the labels with a template after I remove them from the image. I might shrink the legend too, since some of those are redundant. Anyway, is there anything you think should be modified? Seppi333 ( Insert 2¢ | Maintained) 09:05, 18 July 2014 (UTC)
- @
Seppi333: Looks fantastic! As far as I can tell, it is technically correct. One further suggestion is to color code the function of the various proteins/factors. For example (using backgrounds instead of colored text):
- DRD1-5, Gs, Gi/o – GPCRs and G proteins
- ion channels
- AMPAR and NMDAR – ligand gated ion channels
- Cav1.2 – voltage gated ion channel
- cAMP, CaM – second messengers
- enzymes
- CaMKII – kinase
- DARPP-32. PP1, PP2B – phosphatases
- HDAC, SIRT1 – deacetylases
- CREB, Fos, Jun, c-Fos – transcription factors
- The following tool seems like a good way to select a color pallet. Also you might want to submit the diagram as a featured picture candidate. A few examples of diagrams that have been designated feature pictures:
- Cheers.
Boghog (
talk)
07:09, 20 July 2014 (UTC)
- Ah, that's a pretty good idea. I'll work on that sometime tomorrow, There's not much room to work with listing the color descriptions on the image itself, but I can use an annotation transclude a note/reference which contains the list you just wrote with the appropriate colors and has the group parameter "Color code". Then - assuming the transcluded page has a color code references group below the refs section - it'll transclude the named ref group along with the list which will appear (hopefully) appear in a reference tooltip whenever people mouse over it, like below. Seppi333 ( Insert 2¢ | Maintained) 09:20, 20 July 2014 (UTC)
- @
Seppi333: Looks fantastic! As far as I can tell, it is technically correct. One further suggestion is to color code the function of the various proteins/factors. For example (using backgrounds instead of colored text):
I tried a variety of color combinations with this image to see how things looked. I think I'd want to use at most 1 additional background color in the boxes (4 color groups total); it just looks too busy/complicated with more than that. Should I just identify the transcript factors, GPCRs, and ion channels, or is there a better grouping? Seppi333 ( Insert 2¢ | Maintained) 01:11, 21 July 2014 (UTC)
- @
Seppi333: I agree that adding too many colors can be counter productive (both for aesthetics and clarity). Highlighting the GPCRs, G-proteins, ion channels (membrane associated) and transcription factors (nuclear) in one color and everything else (enzymes and second messengers) in another color helps. Membrane associated proteins and the transcription factors are at least on opposite sides of the diagram and therefore already spatially distinguished. Two additional suggestions: (1) Using two different shapes for DRD-1,5 (spirals) and DRD-2,3,4 (ellipses) is confusing. Wouldn't it be better to use single ellipses for DRD-2,3,4 and retain double ellipses for DRD-1,5? That way it is immediately clear that the former are monomers while the latter are dimers. (2) The shapes for the ion channels could be altered so they are a bit closer to their actual low resolution shape (see for example
File:Ion channels.svg). This would make clearer the distinction between the ion conduction pore and the ligand binding sites.
Boghog (
talk)
05:38, 21 July 2014 (UTC)
- Hey Boghog - I updated the dopamine receptors and LGIC's, although I was actually already using the voltage gated ion channel structure in the graphic for the nuclear pore (I more or less duplicated the structure of the nuclear pore and DNA from this svg derivation of your image - File:NF-κB.svg. I improvised and created a new structure that reflected the channel structure of the LGICs for the VGIC. Hope that looks ok. I recolored them for consistency and to differentiate them from the nuclear pore; I think it's completely obvious what part constitutes the ion channel now though. I figure I can just use the multicol template to indicate color relationships.
- How's it look? Seppi333 ( Insert 2¢ | Maintained) 23:37, 21 July 2014 (UTC)
Edit: I'm still fairly certainAs is currently evident on FosB, I can transclude the color code in on an annotated reference to take advantage of the tooltip. Seppi333 ( Insert 2¢ | Maintained) 23:39, 21 July 2014 (UTC)- 2nd edit - I captioned the image with enough generality to hopefully make it reasonably accessible. Feel free to tweak it; the parts of the 3rd and 4th paper I used are in the quote parameter.
Seppi333 (
Insert 2¢ |
Maintained)
02:41, 22 July 2014 (UTC)
- @
Seppi333: Thanks for incorporating the suggested changes above. The figure and the caption are now close to perfect. Great job! Cheers.
Boghog (
talk)
15:06, 22 July 2014 (UTC)
- I may nominate this for EN-wiki FP soon, though I have no clue how people will respond to the annotation template.
File:ΔFosB.svg doesn't exactly reflect
template:psychostimulant addiction very well, even with the Commons annotations.
 In any event, I dropped by to ask you if you knew what the compound in the center of the
synthesis diagram on Bupropion is. I've turned the file into an annotated image in order to add the compound names and fix some formatting issues with that section.
Seppi333 (
Insert 2¢ |
Maintained)
23:49, 20 August 2014 (UTC)
In any event, I dropped by to ask you if you knew what the compound in the center of the
synthesis diagram on Bupropion is. I've turned the file into an annotated image in order to add the compound names and fix some formatting issues with that section.
Seppi333 (
Insert 2¢ |
Maintained)
23:49, 20 August 2014 (UTC)
- A name for the compound is 2-bromo-1-(3-chlorophenyl)propan-1-one. Beyond that, the compound probably has no special significance other than it is a synthetic intermediate. I am not sure how people will respond to the template either. From a technical standpoint, I think it is excellent. But to be featured, it also has to have aesthetic appeal. File:Steroidogenesis.svg demonstrates that it is possible to get a technical diagram designated as a feature picture. It will be interesting to see how others respond to it. Cheers. Boghog ( talk) 19:23, 21 August 2014 (UTC)
- I may nominate this for EN-wiki FP soon, though I have no clue how people will respond to the annotation template.
File:ΔFosB.svg doesn't exactly reflect
template:psychostimulant addiction very well, even with the Commons annotations.
- @
Seppi333: Thanks for incorporating the suggested changes above. The figure and the caption are now close to perfect. Great job! Cheers.
Boghog (
talk)
15:06, 22 July 2014 (UTC)
Disambiguation link notification for August 24
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited REG1B, you added a link pointing to the disambiguation page Regeneration. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 09:17, 24 August 2014 (UTC)
ANKK1 and the mistitlted ANNK1 and addictive behaviors article
Hey Boghog, I figured I should probably run this by you before doing it. Is there any reason the latter article shouldn't be merged into the former? Seppi333 ( Insert 2¢ | Maintained) 00:36, 4 September 2014 (UTC)
- @
Seppi333: Thanks for the heads up. Yes, I agree that
ANNK1 and addictive behaviors should be merged into
ANKK1.
Boghog (
talk) 06:32, 6 September 2014 (UTC)
 DoneI was bold and went ahead and merged the two article.
Boghog (
talk)
07:28, 6 September 2014 (UTC)
DoneI was bold and went ahead and merged the two article.
Boghog (
talk)
07:28, 6 September 2014 (UTC)
Disambiguation link notification for October 4
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited Morphogen, you added links pointing to the disambiguation pages Posterior and Smad. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 09:12, 4 October 2014 (UTC)
Lactase
Greetings, and thank you for your contributions. I am referring to your edit to the Lactase article, which you edited on March 3rd 2010. You changed the optimal temperature to 25 degrees Celcius, and added a reference for this claim. However, I cannot find this in the referenced paper, and a friend who is more used to chemistry publications also could not help me with that. I'm not saying you are wrong, but I kindly ask you to re-check the reference, and if possible hint me at where the temperature optimum of 25 degrees Celcius is mentioned in it. Thank you C-Otto ( talk) 21:57, 9 October 2014 (UTC)
A barnstar for you!

|
The Original Barnstar |
| Appreciate the template work you do :-) Doc James ( talk · contribs · email) (if I write on your page reply on mine) 09:34, 12 October 2014 (UTC) |
There you go, barnstar boy
Hi Boghog. Thanks for the message. Until yesterday, the page regarding PCP4 was totally empty, and I have tryied to add relevant information to it. However, I saw no editions of yours previous to that in this page. Would you possibly be more interested in deleting than adding value? About original research, that would mean research that has never been published. And the immunohistochemistry of Purkinje cell protein 4 in Purkinje cells, as well as the Western blot of PCP4 in H295R cells are well known and have been previously published. If you have access to the actual scientific journals you will be able to check that out yourself. But hey, why not just Google it? Google images id great finding scientific material of that sort sometimes. I know deleting apparently irrelevant material and adding tags to new pages can help adding up a badge of honor as a cure something-something award of Wikipedia to one's user talk page. But I see no reason to hide scientific information that has been published (and therefore does no longer constitute original research) from the public. About the content being too complex for featuring in an encyclopaedia, may I ask what would you mean by that? What I see in Wikipedia right now is some articles containing medical information that is wrong, although those are about topics that are well known since the 50's. Nevertheless, no one with a badge goes there to fix them. And no, I am not going to list these articles. I am sure you can do a good job looking for and fixing all of them. By the way, why don't you add some information about PCP4 to its page? You seem to know the topic well, I mean, you seem to know what you are talking about, right? Please do make sure to add your contribution. I will be looking forward to read it. Cheers. Herizora ( talk) 22:30, 17 October 2014 (UTC)
- Thanks for your response, however you still have not provided a clear answer to the question I asked above. Namely, where did these figures come from? Above you state that the figures are not original research and imply that they were obtained from some published source. If in fact you did obtain these images from a published source, then we have a larger problem. Unless you have obtained permission from the publisher to reuse these images, then it is a
copyright violation and the these images must be removed from Wikipedia. On the other hand, in the image summaries you state that:
-
File:PCP4 immunohistochemistry in human cerebellum.jpg:
I have performed the immunohistochemical analysis of PCP4 in 3 human cerebellum specimens
-
File:PCP4 Western blot with knock-down and control H295R adrenocortical cells.jpg:
I have performed Western blotting of PCP4 and beta-actin
-
File:PCP4 immunohistochemistry in human cerebellum.jpg:
- So which is it? Did you create these figures yourself or did you copy them from somewhere? Please answer the question.
- One of the first things I in fact did was search for the source of these images using Google Image search, and here is what I found:
- Finally what you should be doing is paraphrasing the discussion/conclusion parts of papers and not the results section. Even better would be to paraphrase relevant secondary source (e.g., review articles). If anyone is interested in experimental details, they will read directly the primary literature, not a Wikipedia article. What a Wikipedia article should do is to give a general overview of the subject. Including experimental details (e.g.,
after 6h of angiotensin-II treatment in H295R cells
) is almost never appropriate. Boghog ( talk) 14:26, 18 October 2014 (UTC)
- Hi.
- I have no obligation of answering your questions what-so-ever. But let me tell you. I have published the researched in a peer reviewed journal, and the pictures you deleted are from my PhD thesis, which was subject to peer review for several months from 4 independent peers, which makes it a secondary source.
- I thought of making those files available, both because of the good quality of the content and because my thesis is not be available online and therefore those figures cannot be accessed from the internet.
- So there you have your answer. I HAVE made those figures, and I HAVE copied them from my own peer-reviewed PUBLISHED material.
- At any rate, I will be deleting those files.
- Take your Wikipedia and stick it. I won't make material available to you any more.
- Herizora ( talk) 14:52, 18 October 2014 (UTC)
- BTW, let's keep the discussion at one place, here in your talk page. Hope it won't spoil your shiny badge. Herizora ( talk) 16:20, 18 October 2014 (UTC)
- @
Herizora: I started the discussion on your talk page, so why do you keep moving it back here? Also if someone questions the copyright status of material that you include in Wikipedia, you do have an obligation to answer to the question or agree to the removal of the material. In any case, thank you for answering my question. However your answer raises some additional concerns. While you claim your research has been published, you need to prove it (e.g., by including a filled out {{
cite thesis}} template). Otherwise your figures and associated text must be considered
WP:OR. Completed theses or dissertations may be considered reliable sources but it is preferable to cite journal articles which have undergone external peer review (see
WP:SCHOLARSHIP). Wikipedia is not a good place to publish what are essentially unpublished parts of your thesis. There is also the potential for
WP:COI. While not prohibited, including sources that you have authored in a Wikipedia article is generally frowned upon. Finally there is the appropriateness of the figures. The figures may be appropriate (e.g., to illustrate that PCP4 is expressed in Purkinje cells), but the experimental details of how the figures were obtained are not appropriate for Wikipedia. Including a citation to the published work should be sufficient.
Boghog (
talk)
19:17, 18 October 2014 (UTC)
peer review for several months from 4 independent peers, which makes it a secondary source
– No, it is still primary. Please read secondary.
- Thanks for trimming the PCP4 article. While significantly shorter, IMHO it is now much improved. Boghog ( talk) 21:32, 18 October 2014 (UTC)
- @
Herizora: I started the discussion on your talk page, so why do you keep moving it back here? Also if someone questions the copyright status of material that you include in Wikipedia, you do have an obligation to answer to the question or agree to the removal of the material. In any case, thank you for answering my question. However your answer raises some additional concerns. While you claim your research has been published, you need to prove it (e.g., by including a filled out {{
cite thesis}} template). Otherwise your figures and associated text must be considered
WP:OR. Completed theses or dissertations may be considered reliable sources but it is preferable to cite journal articles which have undergone external peer review (see
WP:SCHOLARSHIP). Wikipedia is not a good place to publish what are essentially unpublished parts of your thesis. There is also the potential for
WP:COI. While not prohibited, including sources that you have authored in a Wikipedia article is generally frowned upon. Finally there is the appropriateness of the figures. The figures may be appropriate (e.g., to illustrate that PCP4 is expressed in Purkinje cells), but the experimental details of how the figures were obtained are not appropriate for Wikipedia. Including a citation to the published work should be sufficient.
Boghog (
talk)
19:17, 18 October 2014 (UTC)
- My pleasure about the PCP4 article. Hope it will stay that way for the next few years.
- Also, thanks for the explanation about the proceedings regarding the {{ cite thesis}} template. I wish I could use the figures from the peer-reviewed article, if that was the case, but then there would be a copyright conflict with the publishing company, as I only have the legal rights to self-archive in one repository previously agreed at the time of publishing.
- While the figures that were uploaded are not unpublished, my thesis won't be available online and I have no intention of filling the {{ cite thesis}} template at present.
- Please flag the two photographs for deletion, if you find appropriate.
- And please make sure to do a more useful job. There are several supposedly serious medical articles in Wikipedia that are full of inappropriate information and pictures (ie. months ago I had to exclude a figure from a major article in medical sciences. Basically, a guy uploaded his own abdominal X-ray with the objective to show his kidneys to the world. And it had been there for months, maybe years). Maybe it's time to start addressing that kind of situation in order for Wikipedia to gain reliability.
-
Herizora (
talk)
22:10, 18 October 2014 (UTC)
- I tend to focus more on articles within the scope of WP:MCB rather than WP:MED. Furthermore there are approximately 19,000 articles in the former and 32,000 articles in the later with significant overlap between the two projects. Given the enormous number of articles and the limited number of active editors, it is not realistic that that all errors will be corrected. The best we can do right now is to concentrate on the most accessed articles. Wikipedia clearly needs to recruit and retain more editors so that we can do a better job of maintaining articles. If you see errors, please continue to correct them. We need all the help we can get! Cheers. Boghog ( talk) 08:05, 19 October 2014 (UTC)
Citation policy
Thanks very much for your comments at the Chemistry project in response to my query. I hope that you will also comment on my follow-up question. In many cases, say an article about a chemical compound or a drug etc, there are hundreds or thousands of papers. Your insights/opinions about how to select from these? -- Smokefoot ( talk) 22:53, 22 October 2014 (UTC)
Class assignment
Good afternoon, I saw you delate my contribution to the article of the pancreatic elastase because it was too much exprimental, I agree with you, but I did it for a university project. The project is due on Wenesday, after that I will delate it. So can you leave it just for now. I will really apreciate it.
Thank you so much. Have a nice week end.
BQUB14 Vmourre — Preceding unsigned comment added by BQUB14 Vmourre ( talk • contribs) 14:47, 24 October 2014 (UTC)
- While I am sympathetic to your situation, Wikipedia does not solely exist for the purpose of class assignments. Your instructor should be aware that added material must conform to Wikipedia standards, particularly with respective to WP:NOTJOURNAL (no experimental details) and WP:Verifiability (any material added must be supported by reliable sources) and communicated this to your class. Your contributions have not been deleted and can still be found in the history of the article. Your instructor should already have been in contact with the people at Wikipedia:Education program. If not, I strongly encourage your instructor to do so. Boghog ( talk) 19:54, 24 October 2014 (UTC)
FERMT3
Dear Boghog, I would really appreciate it if you stopped deleting my work from Wikipedia. The article on FERMT3 is an unfinished university assignment. We are in the process of adding online citations and references but it doesn't help that every time we work on it something has been deleted. Thank you. — Preceding unsigned comment added by Marinabuene ( talk • contribs) 10:20, 26 October 2014 (UTC)
- @ Marinabuene: Thanks for your message. If you carefully take a look at the history of the FERMT3, I have actually restored some of the content your class mates have added with WP:MEDRS compliant sourcing. I and several other editors were becoming very frustrated that our requests to your class mates to provide in-line citations were being ignored (see for example here). The only way seem to get your attention was by deleting unsourced text. I am delighted that you and your class mates are now doing so. Good luck on the completion of your assignment. Cheers. Boghog ( talk) 10:42, 26 October 2014 (UTC)
Thanks a lot for your help. We're new in Wikipedia so we don't know how it really works. Some of our references are repeated and I don't know how to delete one (there are two equal references, the same article). I would be glad if you show me how. Our work is almost done. In your opinion, is it good? — Preceding unsigned comment added by BQUB14-pdelamata ( talk • contribs) 11:55, 26 October 2014 (UTC)
- @ BQUB14-pdelamata:. Actually, it looks very good for a first effort. It needs some additional wikification and copyedit, but all in all, not too bad. You and your classmates are to be congratulated. Concerning duplicate citations, these can be combined by using named ref tags. For example, if your first citation looks like "<ref name="Doe_2014"> {{cite journal | author = Doe J | year = 2014 }}</ref>", then you can cite this source again by inserting the named ref tag "<ref name="Doe_2014"/>" (note the "/" character that is in the reused citation and not in the original). Boghog ( talk) 13:23, 26 October 2014 (UTC)
Wikipedia template filling - ISBN bug
There is no form named "frmconvert" at /data/project/citation-template-filling/perl5/lib/perl5/WWW/Mechanize.pm line 1930. at
[5]
Ihaveacatonmydesk (
talk)
14:10, 26 October 2014 (UTC)
- a lot of other examples don't work either. Ihaveacatonmydesk ( talk) 17:24, 26 October 2014 (UTC)
- I know. I took over the maintenance of the tool from
User:Diberri, and I have never been able to get this to work either. I was just happy to get the PMID and PMC searches to work and these are the most frequent type of searches. I am not a Perl expert, but I tried debugging it, but was not successful. The source code is located
here is you wanted to look into it yourself.
Boghog (
talk)
17:45, 26 October 2014 (UTC)
- Maybe put a banner on it with a notice, or remove the non-working options. Ihaveacatonmydesk ( talk) 17:17, 27 October 2014 (UTC)
- I know. I took over the maintenance of the tool from
User:Diberri, and I have never been able to get this to work either. I was just happy to get the PMID and PMC searches to work and these are the most frequent type of searches. I am not a Perl expert, but I tried debugging it, but was not successful. The source code is located
here is you wanted to look into it yourself.
Boghog (
talk)
17:45, 26 October 2014 (UTC)
Dear Boghog, sorry for any inconvenience. The deadline for the project is today, therefore we won't be making any more modifications. Thanks a lot for your kindness and for helping us with this. — Preceding unsigned comment added by BQUB14-pdelamata ( talk • contribs) 19:05, 26 October 2014 (UTC)
Citation tools
Hi Boghog. Am I correct in thinking the Cite doi and cite pmid templates have been deprecated? They don't seem to work any more and I saw that you added full references to some I'd done as cite doi, and had linked to some discussion about the citation templates. I understand you have resurrected that old template filler tool that works with pmid, but do you know if there is an equivalent for doi? Just as these days I often find myself editing on a mobile device and copy pasting the full citation is much more fiddly than just the doi. Any suggestions would be appreciated. Meodipt ( talk) 08:54, 30 October 2014 (UTC)
- Hi Meodipt. Yes, there was a consensus reached here that transcluded cite templates should no longer be used. For the top 1000 medical articles, I have gone through and substituted transcluded cite templates. I was planning to create a bot to go through all articles within the scope of the WP:MED, WP:PHARMA, and WP:MCB projects and substitute these templates, but I have not gotten around to it yet. Substitution is easy, but respecting WP:CITEVAR can be tricky. In theory, Diberri's citation tool could be extended to create citations from a DOI, but Diberri no longer maintains the tool and it is beyond my limited perl scripting abilities. With a concerted effort, I might be able to figure out how to do this, but I don't have much time to devote to this at the moment. As an alternative, you could use the RefToolbar. I personally don't like that solution because of the number of firstn, lastn parameters it will generate if there is a long author list. But it is still better than creating these by hand. Also you could try searching PubMed using the DOI as a query. This sometimes returns a PMID, especially for more recent citations. Boghog ( talk) 08:40, 1 November 2014 (UTC)
Thanks for your help
Thanks for correcting me over here, I didn't realize that NCBI stuff was public domain. Knowing that will help in the future, so I appreciate it :) Kyle Barbour 00:19, 10 November 2014 (UTC)
nuklear
In my view Nulkear is WP:NOTHERE and WP:DISRUPTIVE - it is not clear to me what your point is with the comments you are making at the SPI discussion. You seem to be trying to lay the ground for unblocking him. Is that your goal? Thanks. Jytdog ( talk) 13:49, 13 November 2014 (UTC)
- The point of my comments at the SPI discussion is that the discussion should be focused on issues relevant to an SPI. Nulkear is a serial sock puppeteer. That reason alone is sufficient for blocking confirmed socks of Nulkear and I do not dispute that for a millisecond. I was responding to Doc James' narrower question of whether the
chemical diagrams are copyright violations. If they are copyright violations, they should be removed from Wikimedia commons. For the reasons I have already stated at the SPI discussion, I think it is pretty clear that these diagrams are not copyright violations. Furthermore in the proper context with proper sources, these diagrams could legitimately be included in a Wikipedia article.
Boghog (
talk)
21:54, 13 November 2014 (UTC)
- thanks, that is very clear! and thanks for all your great work, all the time. 22:22, 13 November 2014 (UTC)
Please reinstate my fixes to Nuclear receptor
This edit reverted my fix of displayauthors errors along with the Citation bot edits to which you are objecting. You should have been able to undo just the Citation bot edits while leaving my fixes in place. Please reinstate my fixes and be more selective next time. Thanks. – Jonesey95 ( talk) 00:17, 15 November 2014 (UTC)
- @ Jonesey95: What on earth is this Help:CS1_errors#.7Cdisplayauthors.3D_suggested non-sense and why should I possibly care? The error documentation is complete gibberish. The documentation needs to clearly explain why there is a problem. Boghog ( talk) 00:33, 15 November 2014 (UTC)
The new Lua-based templates cannot know if editors created citations with exactly nine authors because there were only nine authors or, because old-style citations were limited to nine authors
– huh ??? This seems like a made up problem. In short, there is no problem. Boghog ( talk) 00:37, 15 November 2014 (UTC)
-
"Fixed" in this edit. (
|displayauthors=9) There were exactly 9 authors, not 8. Boghog ( talk) 00:54, 15 November 2014 (UTC)
- In answer to your "What on earth" question, here's a step-by-step explanation, as far as I understand it:
- The old versions of the cite templates had a limit of nine authors.
- Because of those limits, many editors and citation-creation tools created citations with only nine authors, even if the cited source had more than nine authors. Eight authors were displayed, followed by "et al."
- When the cite templates were modernized and converted to Lua, the nine-author limit was eliminated.
- Citations created before the cite templates were modernized, and that had exactly nine authors displayed, were marked with an error message, since there was no way to know whether eight authors and "et al." was correct, or whether exactly nine authors should be displayed.
- I, along with a few other gnome editors, have been comparing these citations with the source articles to determine whether more authors should be added, or if all nine authors should be displayed. In some cases, it is difficult to locate the source article, so
|displayauthors=8is added to match the old citation format. From a total of over 10,000 articles, we are down to 175. Once these 175 are fixed, the error category and error message can be eliminated from the citation template system, and the number of authors displayed will be controlled by the number in the citation, unless|displayauthors=is added intentionally by an editor. – Jonesey95 ( talk) 06:31, 15 November 2014 (UTC)
- In answer to your "What on earth" question, here's a step-by-step explanation, as far as I understand it:
- OK, thanks for the explanation. It seems like a lot of effort is going into fixing a minor problem that most readers would never notice, but at least it will completely disappear once your sweep is over.
Boghog (
talk)
09:26, 15 November 2014 (UTC)
- "a lot of effort is going into fixing a minor problem that most readers would never notice": That's my chosen vocation on WP. To each his own.... – Jonesey95 ( talk) 15:31, 15 November 2014 (UTC)
- OK, thanks for the explanation. It seems like a lot of effort is going into fixing a minor problem that most readers would never notice, but at least it will completely disappear once your sweep is over.
Boghog (
talk)
09:26, 15 November 2014 (UTC)
Reference Errors on 16 November
![]() Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
Hello, I'm
ReferenceBot. I have automatically detected that an edit performed by you may have introduced errors in referencing. It is as follows:
- On the Anabolic steroid page, your edit caused an unsupported parameter error ( help). ( Fix | Ask for help)
Please check this page and fix the errors highlighted. If you think this is a false positive, you can report it to my operator. Thanks, ReferenceBot ( talk) 00:39, 17 November 2014 (UTC)
Copyright checks when performing AfC reviews
Hello Boghog. This message is part of a mass mailing to people who appear active in reviewing articles for creation submissions. First of all, thank you for taking part in this important work! I'm sorry this message is a form letter – it really was the only way I could think of to covey the issue economically. Of course, this also means that I have not looked to see whether the matter is applicable to you in particular.
The issue is in rather large numbers of copyright violations ("copyvios") making their way through AfC reviews without being detected (even when easy to check, and even when hallmarks of copyvios in the text that should have invited a check, were glaring). A second issue is the correct method of dealing with them when discovered.
If you don't do so already, I'd like to ask for your to help with this problem by taking on the practice of performing a copyvio check as the first step in any AfC review. The most basic method is to simply copy a unique but small portion of text from the draft body and run it through a search engine in quotation marks. Trying this from two different paragraphs is recommended. (If you have any question about whether the text was copied from the draft, rather than the other way around (a "backwards copyvio"), the Wayback Machine is very useful for sussing that out.)
If you do find a copyright violation, please do not decline the draft on that basis. Copyright violations need to be dealt with immediately as they may harm those whose content is being used and expose Wikipedia to potential legal liability. If the draft is substantially a copyvio, and there's no non-infringing version to revert to, please mark the page for speedy deletion right away using {{db-g12|url=URL of source}}. If there is an assertion of permission, please replace the draft article's content with {{subst:copyvio|url=URL of source}}.
Some of the more obvious indicia of a copyvio are use of the first person ("we/our/us..."), phrases like "this site", or apparent artifacts of content written for somewhere else ("top", "go to top", "next page", "click here", use of smartquotes, etc.); inappropriate tone of voice, such as an overly informal tone or a very slanted marketing voice with weasel words; including intellectual property symbols (™,®); and blocks of text being added all at once in a finished form with no misspellings or other errors.
I hope this message finds you well and thanks again you for your efforts in this area. Best regards-- Fuhghettaboutit ( talk) 02:20, 18 November 2014 (UTC).
Sent via-- MediaWiki message delivery ( talk) 02:20, 18 November 2014 (UTC)
Disambiguation link notification for November 20
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited HRG (gene), you added a link pointing to the disambiguation page Rosette. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 14:21, 20 November 2014 (UTC)
Boghog -- thanks for your feedback on my Quercetin edits. Respectfully, however, biochemistry is not a natural science but rather is a major field of health science at every medical school. On the WP:SCIRS page is stated "The scope of this page is limited to the natural sciences, including astronomy, biology, chemistry, geoscience, physics, and interdiscliplinary fields. For articles about medicine, see Wikipedia:Reliable sources (medicine-related articles)."
Is it a matter of interpretation or definition? The edits I removed were made on the topic of mechanisms of action of quercetin in vitro. By my experience, I'd say this is biochemistry, so WP:MEDRS applies. All Best. -- Zefr ( talk) 23:41, 4 December 2014 (UTC)
- Respectfully, biochemistry is a natural science, not a medical science. While biochemistry is taught in medical school, biochemistry departments are normally part of colleges of liberal arts and not in medical schools. I double majored in biochemistry as an undergraduate and never attended medical school. More importantly, a biochemical mechanism of action falls within the scope of WP:MCB, not WP:MED. Boghog ( talk) 23:49, 4 December 2014 (UTC)
- "The scope of this page is limited to ... biology, chemistry". biology ∩ chemistry = biochemistry. Boghog ( talk) 23:55, 4 December 2014 (UTC)
Thank you for being one of Wikipedia's top medical contributors!
- please help translate this message into the local language

|
The Cure Award |
| In 2013 you were one of the top 300 medical editors across any language of Wikipedia. Thank you so much for helping bring free, complete, accurate, up-to-date medical information to the public. We really appreciate you and the vital work you do! |
We are wondering about the educational background of our top medical editors. Would you please complete a quick 5-question survey? (please only fill this out if you received the award)
Thanks again :) -- Ocaasi, Doc James and the team at Wiki Project Med Foundation
Deleting the message is fine
Your Talk page is yours to do with as you please. Message was intended just for you, in any case. Sad that you can see only nonsense, and nothing worth your consideration, but that is the state of things. Cheers. Le Prof
In reply to your talk my page also appearing immediately above
I have no clue as to who Flyer22 is in relation to the question of highlighting Talk text to make minority voices clearer, or what "refactoring" or "archiving" are, and what they have to do with highlighting. Here, brevity is your enemy; I am not chasing possible meanings down for you. This you need to clarify, significantly, to simply be understood. As for Padillah's statement, I have asked him to make clear what he wants, and when he does, I will act. As for your "protest", you can assuredly speak to what I do with your comments, but—as far as I see in the WPs, and as far as I have been counseled—not about what I do with other others' comments. I have highlighted none of your comments. I have queried all others, and will respond to their requests. It is, I understand, up to them, individually. (There is no overriding, firm policy, whatever you might think; this is what I am told.) So for now, I will await Padillah et al's replies, and yours if you care to say more. (But no action taken until.) Cheers. Le Prof
Disambiguation link notification for December 7
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited IGFBP3, you added a link pointing to the disambiguation page Esophageal. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:57, 7 December 2014 (UTC)
Wikipedia study- Thank you
Hello Boghog, I hope you remember speaking to me in the summer of 2012 about your motivations for contributing to the health-related pages on Wikipedia. The great news is that the study got published this Wednesday in JMIR (Journal of Medical Internet Research). You can read it here: http://www.jmir.org/2014/12/e260 This would not have been possible without your contributions so once again, I would like to thank you for taking the time and sharing your experiences with me. I also wrote an entry about my own experience with the study, about additional observations and how I plan to further extend my research - published in the WMF blog today: https://blog.wikimedia.org.uk/2014/12/who-writes-wikipedias-health-and-medical-pages-and-why/If you have any comments or questions please get in touch.Perhaps see you at the next Wikimania conference in Mexico! Best Wishes Hydra Rain ( talk) 21:03, 7 December 2014 (UTC)
Need to update template:Amphetamine pharmacodynamics...
Hey Boghog. I noticed in the recently updated KEGG pathway as well as in a recently published paper that presynaptic CaMKIIα signaling plays a role in amphetamine-induced dopamine efflux. Unfortunately, the mechanism of this isn't published because the KEGG and the DAT researchers working on this somehow have never heard of trace amine-associated receptors...
Notes from my sandbox
|
|---|
|
–Amphetamine causes
EAAT3 endocytosis (presynaptic, possibly also postsynaptic) and presynaptic CaMKIIα signaling in DA neurons -
KEGG pathway and
PMID
25033183
References
|
|
After reading all the papers, it seems intuitive to me how this works, but I wanted to run these steps by you before I update the diagram:
- Since TAAR1 is Gq-coupled, its activation causes an influx of calcium via IP3R in the presynaptic neuron (as noted in PMID 16831861 and IUPHAR's TAAR1 page).
- From this point, the intermediate pathway steps between the calcium channel and CaMKIIα are analogous to the pathway in template:Psychostimulant addiction
- CaMKIIα-dependent DAT phosphorylation occurs and results in dopamine efflux/reverse transport
In a nutshell, I'd basically be drawing the following connections: TAAR1 → IP3R → CaM → CaMKIIα → DAT (I'd need to draw IP3R). That said, is there anything missing, or does that sound right?
Seppi333 (
Insert 2¢ |
Maintained)
02:55, 12 November 2014 (UTC)
- I can't even find a review that includes amphetamine and CAMKII, so I'm just going to ignore this until someone decides to write one. The only primary source I found that mentioned both TAAR1-mediated PKC and CAMKII as mechanisms of amphetamine efflux didn't draw the conclusion I jumped to either, so I probably shouldn't do this anyway. I still think that a bit odd how there isn't a paper covering all three kinases yet.
Seppi333 (
Insert 2¢ |
Maintained)
08:15, 8 December 2014 (UTC)
- @
Seppi333: Sorry for not responding sooner. I did take a look at this a couple of weeks ago and could not find any sources that made the connection either. While your reasoning appears to be sound, unfortunately we cannot include the connection without a reliable source to support the connection, otherwise it become OR. Cheers.
Boghog (
talk)
21:23, 8 December 2014 (UTC)
- Yeah... that was my main reason for not wanting to follow up on this. At best, I can point out the associations between the three as I did in the text of
amphetamine, but I don't have any sources to adequately cite that causal chain of events. Besides that, there could even be another intracellular receptor that amph binds to which promotes CAMKII, which was my other reservation. In any event, I moved a lot of the CAMK articles around because
Ca2+/calmodulin-dependent protein kinase was originally going to the article that I moved to
Ca2+/calmodulin-dependent protein kinase II. I turned The former into a redirect to
CAMK, though I'm not sure if "Ca2+/calmodulin-dependent protein kinase" or "CAMK" should be the set index for these kinases. There doesn't appear to be any general article written that includes an overview of all subtypes.
CAMKI, the CAMKII family, and CAMKIV at the moment (I believe that's all the CAMKs, based upon BRENDA at least).Seppi333 ( Insert 2¢ | Maintained) 02:34, 9 December 2014 (UTC)
- Yeah... that was my main reason for not wanting to follow up on this. At best, I can point out the associations between the three as I did in the text of
amphetamine, but I don't have any sources to adequately cite that causal chain of events. Besides that, there could even be another intracellular receptor that amph binds to which promotes CAMKII, which was my other reservation. In any event, I moved a lot of the CAMK articles around because
Ca2+/calmodulin-dependent protein kinase was originally going to the article that I moved to
Ca2+/calmodulin-dependent protein kinase II. I turned The former into a redirect to
CAMK, though I'm not sure if "Ca2+/calmodulin-dependent protein kinase" or "CAMK" should be the set index for these kinases. There doesn't appear to be any general article written that includes an overview of all subtypes.
- @
Seppi333: Sorry for not responding sooner. I did take a look at this a couple of weeks ago and could not find any sources that made the connection either. While your reasoning appears to be sound, unfortunately we cannot include the connection without a reliable source to support the connection, otherwise it become OR. Cheers.
Boghog (
talk)
21:23, 8 December 2014 (UTC)
References
- ^ a b c d e Miller GM (January 2011). "The emerging role of trace amine-associated receptor 1 in the functional regulation of monoamine transporters and dopaminergic activity". J. Neurochem. 116 (2): 164–176. doi: 10.1111/j.1471-4159.2010.07109.x. PMC 3005101. PMID 21073468.
- ^
a
b Eiden LE, Weihe E (January 2011).
"VMAT2: a dynamic regulator of brain monoaminergic neuronal function interacting with drugs of abuse". Ann. N. Y. Acad. Sci. 1216 (1): 86–98.
Bibcode:
2011NYASA1216...86E.
doi:
10.1111/j.1749-6632.2010.05906.x.
PMC
4183197.
PMID
21272013.
VMAT2 is the CNS vesicular transporter for not only the biogenic amines DA, NE, EPI, 5-HT, and HIS, but likely also for the trace amines TYR, PEA, and thyronamine (THYR) ... [Trace aminergic] neurons in mammalian CNS would be identifiable as neurons expressing VMAT2 for storage, and the biosynthetic enzyme aromatic amino acid decarboxylase (AADC). ... AMPH release of DA from synapses requires both an action at VMAT2 to release DA to the cytoplasm and a concerted release of DA from the cytoplasm via "reverse transport" through DAT.
-
^ Sulzer D, Cragg SJ, Rice ME (August 2016).
"Striatal dopamine neurotransmission: regulation of release and uptake". Basal Ganglia. 6 (3): 123–148.
doi:
10.1016/j.baga.2016.02.001.
PMC
4850498.
PMID
27141430.
Despite the challenges in determining synaptic vesicle pH, the proton gradient across the vesicle membrane is of fundamental importance for its function. Exposure of isolated catecholamine vesicles to protonophores collapses the pH gradient and rapidly redistributes transmitter from inside to outside the vesicle. ... Amphetamine and its derivatives like methamphetamine are weak base compounds that are the only widely used class of drugs known to elicit transmitter release by a non-exocytic mechanism. As substrates for both DAT and VMAT, amphetamines can be taken up to the cytosol and then sequestered in vesicles, where they act to collapse the vesicular pH gradient.
-
^ Ledonne A, Berretta N, Davoli A, Rizzo GR, Bernardi G, Mercuri NB (July 2011).
"Electrophysiological effects of trace amines on mesencephalic dopaminergic neurons". Front. Syst. Neurosci. 5: 56.
doi:
10.3389/fnsys.2011.00056.
PMC
3131148.
PMID
21772817.
Three important new aspects of TAs action have recently emerged: (a) inhibition of firing due to increased release of dopamine; (b) reduction of D2 and GABAB receptor-mediated inhibitory responses (excitatory effects due to disinhibition); and (c) a direct TA1 receptor-mediated activation of GIRK channels which produce cell membrane hyperpolarization.
-
^
"TAAR1". GenAtlas. University of Paris. 28 January 2012. Retrieved 29 May 2014.
• tonically activates inwardly rectifying K(+) channels, which reduces the basal firing frequency of dopamine (DA) neurons of the ventral tegmental area (VTA)
-
^ Underhill SM, Wheeler DS, Li M, Watts SD, Ingram SL, Amara SG (July 2014).
"Amphetamine modulates excitatory neurotransmission through endocytosis of the glutamate transporter EAAT3 in dopamine neurons". Neuron. 83 (2): 404–416.
doi:
10.1016/j.neuron.2014.05.043.
PMC
4159050.
PMID
25033183.
AMPH also increases intracellular calcium (Gnegy et al., 2004) that is associated with calmodulin/CamKII activation (Wei et al., 2007) and modulation and trafficking of the DAT (Fog et al., 2006; Sakrikar et al., 2012). ... For example, AMPH increases extracellular glutamate in various brain regions including the striatum, VTA and NAc (Del Arco et al., 1999; Kim et al., 1981; Mora and Porras, 1993; Xue et al., 1996), but it has not been established whether this change can be explained by increased synaptic release or by reduced clearance of glutamate. ... DHK-sensitive, EAAT2 uptake was not altered by AMPH (Figure 1A). The remaining glutamate transport in these midbrain cultures is likely mediated by EAAT3 and this component was significantly decreased by AMPH
-
^ Vaughan RA, Foster JD (September 2013).
"Mechanisms of dopamine transporter regulation in normal and disease states". Trends Pharmacol. Sci. 34 (9): 489–496.
doi:
10.1016/j.tips.2013.07.005.
PMC
3831354.
PMID
23968642.
AMPH and METH also stimulate DA efflux, which is thought to be a crucial element in their addictive properties [80], although the mechanisms do not appear to be identical for each drug [81]. These processes are PKCβ– and CaMK–dependent [72, 82], and PKCβ knock-out mice display decreased AMPH-induced efflux that correlates with reduced AMPH-induced locomotion [72].
Merge proposal
You have proposed that Environmental fate of TNT be merged to TNT#Bio-degradation, but you have not opened a discussion regarding said merger. I have an opinion on the matter, but in the interest of maintaining a proper chronolgy of the discussion, I'd like you to open a discussion first. WikiDan61 ChatMe! ReadMe!! 21:48, 11 December 2014 (UTC)
 Done OK, per your request, I have opened a merge discussion
here. Sorry for not doing this sooner.
Boghog (
talk)
05:06, 12 December 2014 (UTC)
Done OK, per your request, I have opened a merge discussion
here. Sorry for not doing this sooner.
Boghog (
talk)
05:06, 12 December 2014 (UTC)
Disambiguation link notification for December 20
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that when you edited BCAR1, you added a link pointing to the disambiguation page Hypoxia. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 08:56, 20 December 2014 (UTC)
Thanks for your input.
As you may have gathered, this article has a new anonymous editor, who posted a question today...
Wikipedia:Help_desk#Help:Cite_errors.2FCite_error_ref_no_input. I didn’t want to frighten her off right at the start by mentioning COI (she is one of the authors of the paper she posted). However, as she first was trying to place it in Further Reading, then later included it in Ref 8, the former is now superfluous. Do you suggest I delete it now or wait till she comes back? I'm thinking that it can be off-putting for a any new editor to have loads of other editors making changes before one has had a chance to get a hang of our wiki-ways. She appears to have in depth knowledge of this subject so I welcomed her
here.--
Aspro (
talk)
00:54, 21 December 2014 (UTC)
- Hi. Thanks for you diligence in following up on this and for encouraging this new editor. Yes, clearly the editor is an expert in the field as she is a senior author on the paper she cited. While that constitutes a conflict of interest, the paper itself is both relevant and notable (it has been highly cited by independent reliable sources, see for example Google Scholar, Cited by 37 and Cited by 16 PubMed Central articles). Hence I am less concerned about the conflict of interest. Of bigger concern however is that the source is primary. Per WP:MEDRS secondary sources are needed to support medical claims. Hence I suggest that this source be replaced by a relevant review article. Boghog ( talk) 10:15, 21 December 2014 (UTC)
Disambiguation link notification for December 27
Hi. Thank you for your recent edits. Wikipedia appreciates your help. We noticed though that you've added some links pointing to disambiguation pages. Such links are almost always unintended, since a disambiguation page is merely a list of "Did you mean..." article titles. Read the FAQ • Join us at the DPL WikiProject.
-
ARHGEF7
- added a link pointing to Rho GTPase
-
Ecological impact of explosives
- added a link pointing to Environment
It's OK to remove this message. Also, to stop receiving these messages, follow these opt-out instructions. Thanks, DPL bot ( talk) 09:32, 27 December 2014 (UTC)
science the endless frontier
did you ever read this? Jytdog ( talk) 15:24, 29 December 2014 (UTC)
- I was aware of the Vannevar Bush report, but never read it in detail. It was enormously influential and lead to a major expansion of Federal support for research after World War II. I am not exactly sure why you brought this up, but the following quote from the report may be relevant to this recent MEDMOS discussion:
Further [medical] progress requires that the entire front of medicine and the underlying sciences of chemistry, physics, anatomy, biochemistry, physiology, pharmacology, bacteriology, pathology, parasitology, etc., be broadly developed.
Boghog (
talk)
02:31, 30 December 2014 (UTC)