| THAP3 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | THAP3, THAP domain containing 3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 612532; MGI: 1917126; HomoloGene: 18413; GeneCards: THAP3; OMA: THAP3 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||

THAP domain-containing protein 3 (THAP3) is a protein that, in Homo sapiens (humans), is encoded by the THAP3 gene. [7] The THAP3 protein is as known as MGC33488, LOC90326, and THAP domain-containing, apoptosis associated protein 3. This protein contains the Thanatos-associated protein (THAP) domain [8] and a host-cell factor 1C binding motif. [9] These domains allow THAP3 to influence a variety of processes, including transcription and neuronal development. [10] THAP3 is ubiquitously expressed in H. sapiens, though expression is highest in the kidneys. [7]
Gene
The H. sapiens THAP3 gene is a protein-encoding gene that is located on the plus strand of chromosome 1 [7] at cytogenetic location 1p36.31. [11] It is 10,727 base pairs long, spanning from genomic coordinates 6,624,868-6,635,595. [11] It contains 6 exons. [12]

Expression
In H. sapiens, THAP3 gene is expressed ubiquitously throughout different tissues, and expression is greatest in the kidneys. [13] It has also been determined that expression of THAP3 tends to be slightly higher in organs located in the abdomen and male and female sexual organs, such as the ovaries, testes, prostate, adrenal gland, spleen, liver, and colon, though expression in the kidneys is 1.4-1.5x higher than those organs. [13] THAP3 mRNA is 1.3x. more abundant in H. sapiens fetal brain tissue than in H. sapiens adult kidney tissue. [14]
mRNA
Transcription of the THAP3 gene can result in 11 different mRNA variants, of which 8 are alternatively spliced and 3 are unspliced. [7] Variant 1 is the predominant variant and encodes THAP3 protein isoform 1. [7]
| Variant | Sequence length (nucleotides) | Accession number [7] |
|---|---|---|
| 1 | 1358 | NM_001195752.2 [16] |
| 2 | 2071 | NM_138350.4 [17] |
| 3 | 1361 | NM_001195753.2 [18] |
| 4 | 1262 | NM_001394496.1 [19] |
| 5 | 2050 | NM_001394497.1 [20] |
| 6 | 2047 | NM_001394498.1 [21] |
| 7 | 1123 | NM_001394499.1 [22] |
| 8 | 1120 | NM_001394500.1 [23] |
Protein
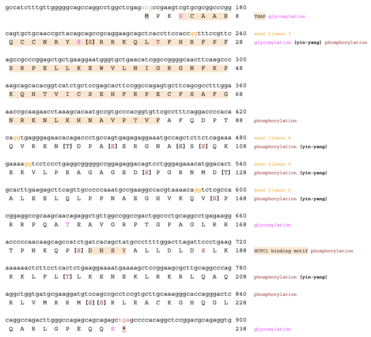
The H. sapiens THAP3 protein is predicted to have a molecular weight of 26.9 kilodaltons [24] and a pI of 10.26. [25] The amino acid sequence is isoleucine and tyrosine rich and arginine poor. [24] Characteristics domains of H. sapiens are the THAP domain (THAP) and the hell-cell factor 1C binding motif (HCM). [7]
Isoforms
Due to having 8 alternatively spliced variants, there are 8 THAP3 isoforms. [7]
| Isoform | Sequence length (amino acids) | Accession number | Encoded by |
|---|---|---|---|
| 1 | 238 | NP_001182681.1 [26] | Variant 1 |
| 2 | 175 | NP_612359.2 [27] | Variant 2 |
| 3 | 239 | NP_001182682.1 [28] | Variant 3 |
| 4 | 236 | NP_001381425.1 [29] | Variant 4 |
| 5 | 168 | NP_001381426.1 [30] | Variant 5 |
| 6 | 167 | NP_001381427.1 [31] | Variant 6 |
| 7 | 148 | NP_001381428.1 [32] | Variant 7 |
| 8 | 147 | NP_001381429.1 [33] | Variant 8 |
Structure
The predicted H. sapiens THAP3 tertiary structure contains a globular region and an alpha helix. [5] [6] The globular region is located near the N-terminus of the sequence and is the structure of the THAP domain. It spans amino acids 4-82. [37] The alpha helix is located from amino acids 186-230 and contains the host-cell factor 1C binding motif. [37]
Regulation
Localization
THAP3 can be localized in the nucleus or mitochondria of H. sapiens cells. [38]
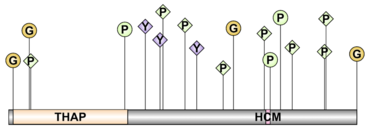
Post-translation modifications
The H. sapiens the THAP3 protein has 30 predicted phosphorylation sites, 28 predicted O-β-glycosylation sites, and 11 predicted Yin-Yang sites. [35] [36] Many proteins involved in transcription regulation are influenced by phosphorylation and glycosylation sites, which corroborates THAP3's function. [39]
Homology and evolution
Paralogs
The H. sapiens THAP3 protein, along with several other proteins, is part of the THAP family of proteins. [40] All of these proteins contain the THAP domain and are, thus, paralogs of H. sapiens THAP3. [15]
| Protein Name | E-Value! [15] | Percent Identity to THAP3 [15] |
|---|---|---|
| THAP1 [41] | 8×10−23 | 48.00 |
| THAP2 [42] | 6×10−17 | 45.24 |
| THAP5 [43] | 4×10−13 | 31.96 |
| THAP6 [44] | 6×10−6 | 34.44 |
| THAP7 [45] | 1×10−7 | 33.33 |
| THAP8 [46] | 8×10−11 | 31.96 |
| THAP9 [47] | 2×10−8 | 32.99 |
Orthologs

There are approximately 206 orthlologs of H. sapiens THAP3. [7] Orthologs can be found in a variety of taxomonic classes, including mammals, reptiles, amphibians, bony fishes, and cartilaginous fishes. [15] However, there are no orthologs in bacteria, fungi, protists, archaea, plants, invertebrates, or birds. [15] Additionally, not all orders are represented with in a class. For example, in reptiles, orthologs to H. sapiens THAP3 are found in testudines (turtles or tortoises) and not found in crocodilia (crocodiles and alligators) or squamata (lizards and snakes). [15] Similarly, there are only orthologs in apoda within amphibians. [15] There are no orthologs in anura (frogs) or urodela (salamanders). [15]
In closely related organisms, those diverged 0-160 million years ago (MYA), percent similarity of orthologs ranges from 36-82.9%. THAP3 sequences in rodents are the least conserved compared to H. sapiens. Sequences that diverged 319-353 MYA, those moderately related, have 47.2-68.9% similarity to H. sapiens THAP3, and 41.3-54.1% similarity in organisms that are distantly related, diverged 431-464 MYA.
| Taxonomic Class | Scientific Name | Common Name | Taxonomic Order | Date of Divergence [50] | Accession Number [15] | Percent Identity to THAP3 | Percent Similarity to THAP3 [51] |
|---|---|---|---|---|---|---|---|
| Mammals | Marmota flaviventris | Yellow-bellied marmot | Rodentia | 87 | XP_027803226.1 [52] | 29.9 | 36 |
| Lontra canadensis | North American river otter | Carnivora | 94 | XP_032719186.1 [53] | 59.5 | 65.8 | |
| Eptesicus fuscus | Big brown bat | Chiroptera | 94 | XP_028016747.1 [54] | 65.0 | 69.6 | |
| Balaenoptera musculus | Blue whale | Cetacea | 94 | XP_036686252.1 [55] | 77.5 | 82.9 | |
| Dromiciops gliroides | Colocolo opossum | Microbiotheria | 160 | XP_043850206.1 [56] | 64.6 | 74.5 | |
| Phascolarctos cinereus | Koala | Diprotodontia | 160 | XP_020830574.1 [57] | 65.7 | 76.4 | |
| Reptiles | Caretta caretta | Loggerhead turtle | Testudines | 319 | XP_048680971.1 [58] | 36.9 | 47.2 |
| Gopherus evgoodei | Goode's thornscrub tortoise | Testudines | 319 | XP_030393185.1 [59] | 48.9 | 58.1 | |
| Chelonoidis abingdonii | Abingdon Island giant tortoise | Testudines | 319 | XP_032619750.1 [60] | 48.9 | 61.4 | |
| Mauremys mutica | Yellow pond turtle | Testudines | 319 | XP_044852367.1 [61] | 49.0 | 60.9 | |
| Amphibians | Microcaecilia unicolor | Microcaecilia unicolor | Gymnophiona | 353 | XP_030041702.1 [62] | 41.2 | 56.8 |
| Geotrypetes seraphini | Gaboon caecilian | Gymnophiona | 353 | XP_033777236.1 [63] | 44.2 | 57.8 | |
| Bony Fishes | Electrophorus electricus | Electric eel | Gymnotiformes | 431 | XP_026873261.2 [64] | 31.9 | 41.3 |
| Coregonus clupeaformis | Lake whitefish | Salmoniformes | 431 | XP_041712304.2 [65] | 32.9 | 47.7 | |
| Brienomyrus brachyistius | Baby whale | Osteoglossiformes | 431 | XP_048872538.1 [66] | 33.5 | 46.3 | |
| Puntigrus tetrazona | Sumatra barb | Cypriniformes | 431 | XP_043081346.1 [67] | 34.0 | 48.1 | |
| Cartilaginous Fishes | Rhincodon typus | Whale shark | Orectolobiformes | 464 | XP_020386430.1 [68] | 39.0 | 53.4 |
| Chiloscyllium plagiosum | White-spotted bamboo shark | Orectolobiformes | 464 | XP_043531920.1 [69] | 39.0 | 53.0 | |
| Amblyraja radiata | Thorny skate | Rajiformes | 464 | XP_032904038.1 [70] | 40.2 | 54.1 |
Evolution
H. sapiens THAP3 has evolved at a rate similar to H. sapiens fibrinogen alpha, which is involved in the immune system. [15]
Protein interactions
H. sapiens THAP3 interacts with proteins involved in various cellular processes, like transcription regulation and neuronal development. [10] It is also interacts with molecular chaperones during its translation.
| Process | Protein Name | Identified By! [72] | Interaction Type |
|---|---|---|---|
| Transcription Regulation | CHAT | two hybrid assay | Functional |
| FGFR3 | two hybrid assay | Functional | |
| HCF1C [73] | affinity capture - mass spectrometry | Functional | |
| OGT [73] | affinity capture - mass spectrometry | Functional | |
| PKN1 | two hybrid assay | Functional | |
| POLR2A | two hybrid assay | Functional | |
| TARDBP | two hybrid assay | Functional | |
| Neuronal Development | LSAMP | two hybrid assay | Functional |
| DNAJB6 | two hybrid assay | Functional | |
| Protein Folding | BAG6 | two hybrid assay | Developmental |
Clinical significance
THAP3 contributes to the presentation of X-linked Dystonia-Parkinsonism, also known as Lubag Syndrome. [74] This disease is a neurodegenerative movement disorder that predominantly affects males of Filipino descent. [75] Symptoms include tremors, bradykinesia, rigidity, postural instability, shuffling gait and dystonia, which typically develops later in life. [75]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000041988 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000039759 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. (August 2021). "Highly accurate protein structure prediction with AlphaFold". Nature. 596 (7873): 583–589. Bibcode: 2021Natur.596..583J. doi: 10.1038/s41586-021-03819-2. PMC 8371605. PMID 34265844.
- ^ a b Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, et al. (January 2022). "AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models". Nucleic Acids Research. 50 (D1): D439–D444. doi: 10.1093/nar/gkab1061. PMC 8728224. PMID 34791371.
- ^ a b c d e f g h i j k l "THAP3 THAP domain containing 3 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2022-12-08.
- ^ Roussigne M, Kossida S, Lavigne AC, Clouaire T, Ecochard V, Glories A, et al. (February 2003). "The THAP domain: a novel protein motif with similarity to the DNA-binding domain of P element transposase". Trends in Biochemical Sciences. 28 (2): 66–69. doi: 10.1016/S0968-0004(02)00013-0. PMID 12575992.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 1, mRNA". 2022-04-22.
- ^ a b Sabogal A, Lyubimov AY, Corn JE, Berger JM, Rio DC (January 2010). "THAP proteins target specific DNA sites through bipartite recognition of adjacent major and minor grooves". Nature Structural & Molecular Biology. 17 (1): 117–123. doi: 10.1038/nsmb.1742. PMC 2933787. PMID 20010837.
- ^ a b "Entry - *612532 - THAP Doman-Containing Protein 3; THAP3 - OMIM". www.omim.org. Retrieved 2022-12-15.
- ^ a b "THAP domain-containing protein 3 isoform 1 [Homo sapiens] - Protein - NCBI". National Center of Biotechnology Information. Retrieved 2022-12-16.
- ^ a b Fagerberg L, Hallström BM, Oksvold P, Kampf C, Djureinovic D, Odeberg J, et al. (February 2014). "Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics". Molecular & Cellular Proteomics. 13 (2): 397–406. doi: 10.1074/mcp.M113.035600. PMC 3916642. PMID 24309898.
- ^ Duff MO, Olson S, Wei X, Garrett SC, Osman A, Bolisetty M, et al. (May 2015). "Genome-wide identification of zero nucleotide recursive splicing in Drosophila". Nature. 521 (7552): 376–379. Bibcode: 2015Natur.521..376D. doi: 10.1038/nature14475. PMC 4529404. PMID 25970244.
- ^ a b c d e f g h i j k "Protein BLAST: search protein databases using a protein query". National Center of Biotechnology Information. Retrieved 2022-12-08.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 1, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 2, m - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. 22 April 2022.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 3, m - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. 10 June 2022.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 4, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 5, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 6, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 7, m - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. 22 April 2022.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 8, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ a b Brendel V, Bucher P, Nourbakhsh IR, Blaisdell BE, Karlin S (March 1992). "Methods and algorithms for statistical analysis of protein sequences". Proceedings of the National Academy of Sciences of the United States of America. 89 (6): 2002–2006. Bibcode: 1992PNAS...89.2002B. doi: 10.1073/pnas.89.6.2002. PMC 48584. PMID 1549558.
- ^ Gasteiger E., Hoogland C., Gattiker A., Duvaud S., Wilkins M.R., Appel R.D., Bairoch A.; Protein Identification and Analysis Tools on the Expasy Server; (In) John M. Walker (ed): The Proteomics Protocols Handbook, Humana Press (2005).
- ^ "THAP domain-containing protein 3 isoform 1 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 2 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 3 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 4 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 5 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 6 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 7 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 8 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ Liu W, Xie Y, Ma J, Luo X, Nie P, Zuo Z, et al. (October 2015). "IBS: an illustrator for the presentation and visualization of biological sequences". Bioinformatics. 31 (20): 3359–3361. doi: 10.1093/bioinformatics/btv362. PMC 4595897. PMID 26069263.
- ^ a b c Gupta, R. (2001). Prediction of glycosylation sites in proteomes: from post-translational modifications to protein function. Technical University of Denmark.
- ^ a b Blom N, Gammeltoft S, Brunak S (December 1999). "Sequence and structure-based prediction of eukaryotic protein phosphorylation sites". Journal of Molecular Biology. 294 (5): 1351–1362. doi: 10.1006/jmbi.1999.3310. PMID 10600390.
- ^ a b Wang J, Youkharibache P, Marchler-Bauer A, Lanczycki C, Zhang D, Lu S, et al. (2022). "iCn3D: From Web-Based 3D Viewer to Structural Analysis Tool in Batch Mode". Frontiers in Molecular Biosciences. 9: 831740. doi: 10.3389/fmolb.2022.831740. PMC 8892267. PMID 35252351.
- ^ "PSORT II Prediction". psort.hgc.jp. Retrieved 2022-12-16.
- ^ Filtz TM, Vogel WK, Leid M (February 2014). "Regulation of transcription factor activity by interconnected post-translational modifications". Trends in Pharmacological Sciences. 35 (2): 76–85. doi: 10.1016/j.tips.2013.11.005. PMC 3954851. PMID 24388790.
- ^ Sanghavi HM, Mallajosyula SS, Majumdar S (March 2019). "Classification of the human THAP protein family identifies an evolutionarily conserved coiled coil region". BMC Structural Biology. 19 (1): 4. doi: 10.1186/s12900-019-0102-2. PMC 6402169. PMID 30836974.
- ^ "THAP1 THAP domain containing 1 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP2 THAP domain containing 2 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP5 THAP domain containing 5 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP6 THAP domain containing 6 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP7 THAP domain containing 7 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP8 THAP domain containing 8 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP9 THAP domain containing 9 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, et al. (October 2011). "Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega". Molecular Systems Biology. 7 (1): 539. doi: 10.1038/msb.2011.75. PMC 3261699. PMID 21988835.
- ^ "THAP domain-containing protein 3 isoform 1 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2022-12-08.
- ^ Kumar S, Suleski M, Craig JM, Kasprowicz AE, Sanderford M, Li M, et al. (August 2022). "TimeTree 5: An Expanded Resource for Species Divergence Times". Molecular Biology and Evolution. 39 (8): msac174. doi: 10.1093/molbev/msac174. PMC 9400175. PMID 35932227.
- ^ "EMBOSS Needle < Pairwise Sequence Alignment < EMBL-EBI". www.ebi.ac.uk. Retrieved 2022-12-08.
- ^ "THAP domain-containing protein 3 isoform X1 [Marmota flaviventris] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Lontra canadensis] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Eptesicus fuscus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Balaenoptera musculus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Dromiciops gliroides] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Phascolarctos cinereus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Caretta caretta] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Gopherus evgoodei] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Chelonoidis abingdonii] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Mauremys mutica] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Microcaecilia unicolor] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Geotrypetes seraphini] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Electrophorus electricus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Coregonus clupeaformis] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Brienomyrus brachyistius] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Puntigrus tetrazona] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Rhincodon typus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Chiloscyllium plagiosum] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Amblyraja radiata] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "UniProt". www.uniprot.org. Retrieved 2022-12-16.
- ^ "IntAct Portal". www.ebi.ac.uk. Retrieved 2022-12-16.
- ^ a b "THAP3 Result Summary | BioGRID". thebiogrid.org. Retrieved 2022-12-16.
- ^ "THAP3 Gene - GeneCards | THAP3 Protein | THAP3 Antibody". www.genecards.org. Retrieved 2022-12-08.
- ^ a b Rosales RL (October 2010). "X-linked dystonia parkinsonism: clinical phenotype, genetics and therapeutics". Journal of Movement Disorders. 3 (2): 32–38. doi: 10.14802/jmd.10009. PMC 4027667. PMID 24868378.
| THAP3 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | THAP3, THAP domain containing 3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 612532; MGI: 1917126; HomoloGene: 18413; GeneCards: THAP3; OMA: THAP3 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||

THAP domain-containing protein 3 (THAP3) is a protein that, in Homo sapiens (humans), is encoded by the THAP3 gene. [7] The THAP3 protein is as known as MGC33488, LOC90326, and THAP domain-containing, apoptosis associated protein 3. This protein contains the Thanatos-associated protein (THAP) domain [8] and a host-cell factor 1C binding motif. [9] These domains allow THAP3 to influence a variety of processes, including transcription and neuronal development. [10] THAP3 is ubiquitously expressed in H. sapiens, though expression is highest in the kidneys. [7]
Gene
The H. sapiens THAP3 gene is a protein-encoding gene that is located on the plus strand of chromosome 1 [7] at cytogenetic location 1p36.31. [11] It is 10,727 base pairs long, spanning from genomic coordinates 6,624,868-6,635,595. [11] It contains 6 exons. [12]

Expression
In H. sapiens, THAP3 gene is expressed ubiquitously throughout different tissues, and expression is greatest in the kidneys. [13] It has also been determined that expression of THAP3 tends to be slightly higher in organs located in the abdomen and male and female sexual organs, such as the ovaries, testes, prostate, adrenal gland, spleen, liver, and colon, though expression in the kidneys is 1.4-1.5x higher than those organs. [13] THAP3 mRNA is 1.3x. more abundant in H. sapiens fetal brain tissue than in H. sapiens adult kidney tissue. [14]
mRNA
Transcription of the THAP3 gene can result in 11 different mRNA variants, of which 8 are alternatively spliced and 3 are unspliced. [7] Variant 1 is the predominant variant and encodes THAP3 protein isoform 1. [7]
| Variant | Sequence length (nucleotides) | Accession number [7] |
|---|---|---|
| 1 | 1358 | NM_001195752.2 [16] |
| 2 | 2071 | NM_138350.4 [17] |
| 3 | 1361 | NM_001195753.2 [18] |
| 4 | 1262 | NM_001394496.1 [19] |
| 5 | 2050 | NM_001394497.1 [20] |
| 6 | 2047 | NM_001394498.1 [21] |
| 7 | 1123 | NM_001394499.1 [22] |
| 8 | 1120 | NM_001394500.1 [23] |
Protein

The H. sapiens THAP3 protein is predicted to have a molecular weight of 26.9 kilodaltons [24] and a pI of 10.26. [25] The amino acid sequence is isoleucine and tyrosine rich and arginine poor. [24] Characteristics domains of H. sapiens are the THAP domain (THAP) and the hell-cell factor 1C binding motif (HCM). [7]
Isoforms
Due to having 8 alternatively spliced variants, there are 8 THAP3 isoforms. [7]
| Isoform | Sequence length (amino acids) | Accession number | Encoded by |
|---|---|---|---|
| 1 | 238 | NP_001182681.1 [26] | Variant 1 |
| 2 | 175 | NP_612359.2 [27] | Variant 2 |
| 3 | 239 | NP_001182682.1 [28] | Variant 3 |
| 4 | 236 | NP_001381425.1 [29] | Variant 4 |
| 5 | 168 | NP_001381426.1 [30] | Variant 5 |
| 6 | 167 | NP_001381427.1 [31] | Variant 6 |
| 7 | 148 | NP_001381428.1 [32] | Variant 7 |
| 8 | 147 | NP_001381429.1 [33] | Variant 8 |
Structure

The predicted H. sapiens THAP3 tertiary structure contains a globular region and an alpha helix. [5] [6] The globular region is located near the N-terminus of the sequence and is the structure of the THAP domain. It spans amino acids 4-82. [37] The alpha helix is located from amino acids 186-230 and contains the host-cell factor 1C binding motif. [37]
Regulation
Localization
THAP3 can be localized in the nucleus or mitochondria of H. sapiens cells. [38]
Post-translation modifications
The H. sapiens the THAP3 protein has 30 predicted phosphorylation sites, 28 predicted O-β-glycosylation sites, and 11 predicted Yin-Yang sites. [35] [36] Many proteins involved in transcription regulation are influenced by phosphorylation and glycosylation sites, which corroborates THAP3's function. [39]
Homology and evolution
Paralogs
The H. sapiens THAP3 protein, along with several other proteins, is part of the THAP family of proteins. [40] All of these proteins contain the THAP domain and are, thus, paralogs of H. sapiens THAP3. [15]
| Protein Name | E-Value! [15] | Percent Identity to THAP3 [15] |
|---|---|---|
| THAP1 [41] | 8×10−23 | 48.00 |
| THAP2 [42] | 6×10−17 | 45.24 |
| THAP5 [43] | 4×10−13 | 31.96 |
| THAP6 [44] | 6×10−6 | 34.44 |
| THAP7 [45] | 1×10−7 | 33.33 |
| THAP8 [46] | 8×10−11 | 31.96 |
| THAP9 [47] | 2×10−8 | 32.99 |
Orthologs

There are approximately 206 orthlologs of H. sapiens THAP3. [7] Orthologs can be found in a variety of taxomonic classes, including mammals, reptiles, amphibians, bony fishes, and cartilaginous fishes. [15] However, there are no orthologs in bacteria, fungi, protists, archaea, plants, invertebrates, or birds. [15] Additionally, not all orders are represented with in a class. For example, in reptiles, orthologs to H. sapiens THAP3 are found in testudines (turtles or tortoises) and not found in crocodilia (crocodiles and alligators) or squamata (lizards and snakes). [15] Similarly, there are only orthologs in apoda within amphibians. [15] There are no orthologs in anura (frogs) or urodela (salamanders). [15]
In closely related organisms, those diverged 0-160 million years ago (MYA), percent similarity of orthologs ranges from 36-82.9%. THAP3 sequences in rodents are the least conserved compared to H. sapiens. Sequences that diverged 319-353 MYA, those moderately related, have 47.2-68.9% similarity to H. sapiens THAP3, and 41.3-54.1% similarity in organisms that are distantly related, diverged 431-464 MYA.
| Taxonomic Class | Scientific Name | Common Name | Taxonomic Order | Date of Divergence [50] | Accession Number [15] | Percent Identity to THAP3 | Percent Similarity to THAP3 [51] |
|---|---|---|---|---|---|---|---|
| Mammals | Marmota flaviventris | Yellow-bellied marmot | Rodentia | 87 | XP_027803226.1 [52] | 29.9 | 36 |
| Lontra canadensis | North American river otter | Carnivora | 94 | XP_032719186.1 [53] | 59.5 | 65.8 | |
| Eptesicus fuscus | Big brown bat | Chiroptera | 94 | XP_028016747.1 [54] | 65.0 | 69.6 | |
| Balaenoptera musculus | Blue whale | Cetacea | 94 | XP_036686252.1 [55] | 77.5 | 82.9 | |
| Dromiciops gliroides | Colocolo opossum | Microbiotheria | 160 | XP_043850206.1 [56] | 64.6 | 74.5 | |
| Phascolarctos cinereus | Koala | Diprotodontia | 160 | XP_020830574.1 [57] | 65.7 | 76.4 | |
| Reptiles | Caretta caretta | Loggerhead turtle | Testudines | 319 | XP_048680971.1 [58] | 36.9 | 47.2 |
| Gopherus evgoodei | Goode's thornscrub tortoise | Testudines | 319 | XP_030393185.1 [59] | 48.9 | 58.1 | |
| Chelonoidis abingdonii | Abingdon Island giant tortoise | Testudines | 319 | XP_032619750.1 [60] | 48.9 | 61.4 | |
| Mauremys mutica | Yellow pond turtle | Testudines | 319 | XP_044852367.1 [61] | 49.0 | 60.9 | |
| Amphibians | Microcaecilia unicolor | Microcaecilia unicolor | Gymnophiona | 353 | XP_030041702.1 [62] | 41.2 | 56.8 |
| Geotrypetes seraphini | Gaboon caecilian | Gymnophiona | 353 | XP_033777236.1 [63] | 44.2 | 57.8 | |
| Bony Fishes | Electrophorus electricus | Electric eel | Gymnotiformes | 431 | XP_026873261.2 [64] | 31.9 | 41.3 |
| Coregonus clupeaformis | Lake whitefish | Salmoniformes | 431 | XP_041712304.2 [65] | 32.9 | 47.7 | |
| Brienomyrus brachyistius | Baby whale | Osteoglossiformes | 431 | XP_048872538.1 [66] | 33.5 | 46.3 | |
| Puntigrus tetrazona | Sumatra barb | Cypriniformes | 431 | XP_043081346.1 [67] | 34.0 | 48.1 | |
| Cartilaginous Fishes | Rhincodon typus | Whale shark | Orectolobiformes | 464 | XP_020386430.1 [68] | 39.0 | 53.4 |
| Chiloscyllium plagiosum | White-spotted bamboo shark | Orectolobiformes | 464 | XP_043531920.1 [69] | 39.0 | 53.0 | |
| Amblyraja radiata | Thorny skate | Rajiformes | 464 | XP_032904038.1 [70] | 40.2 | 54.1 |
Evolution
H. sapiens THAP3 has evolved at a rate similar to H. sapiens fibrinogen alpha, which is involved in the immune system. [15]
Protein interactions
H. sapiens THAP3 interacts with proteins involved in various cellular processes, like transcription regulation and neuronal development. [10] It is also interacts with molecular chaperones during its translation.
| Process | Protein Name | Identified By! [72] | Interaction Type |
|---|---|---|---|
| Transcription Regulation | CHAT | two hybrid assay | Functional |
| FGFR3 | two hybrid assay | Functional | |
| HCF1C [73] | affinity capture - mass spectrometry | Functional | |
| OGT [73] | affinity capture - mass spectrometry | Functional | |
| PKN1 | two hybrid assay | Functional | |
| POLR2A | two hybrid assay | Functional | |
| TARDBP | two hybrid assay | Functional | |
| Neuronal Development | LSAMP | two hybrid assay | Functional |
| DNAJB6 | two hybrid assay | Functional | |
| Protein Folding | BAG6 | two hybrid assay | Developmental |
Clinical significance
THAP3 contributes to the presentation of X-linked Dystonia-Parkinsonism, also known as Lubag Syndrome. [74] This disease is a neurodegenerative movement disorder that predominantly affects males of Filipino descent. [75] Symptoms include tremors, bradykinesia, rigidity, postural instability, shuffling gait and dystonia, which typically develops later in life. [75]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000041988 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000039759 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. (August 2021). "Highly accurate protein structure prediction with AlphaFold". Nature. 596 (7873): 583–589. Bibcode: 2021Natur.596..583J. doi: 10.1038/s41586-021-03819-2. PMC 8371605. PMID 34265844.
- ^ a b Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, et al. (January 2022). "AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models". Nucleic Acids Research. 50 (D1): D439–D444. doi: 10.1093/nar/gkab1061. PMC 8728224. PMID 34791371.
- ^ a b c d e f g h i j k l "THAP3 THAP domain containing 3 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2022-12-08.
- ^ Roussigne M, Kossida S, Lavigne AC, Clouaire T, Ecochard V, Glories A, et al. (February 2003). "The THAP domain: a novel protein motif with similarity to the DNA-binding domain of P element transposase". Trends in Biochemical Sciences. 28 (2): 66–69. doi: 10.1016/S0968-0004(02)00013-0. PMID 12575992.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 1, mRNA". 2022-04-22.
- ^ a b Sabogal A, Lyubimov AY, Corn JE, Berger JM, Rio DC (January 2010). "THAP proteins target specific DNA sites through bipartite recognition of adjacent major and minor grooves". Nature Structural & Molecular Biology. 17 (1): 117–123. doi: 10.1038/nsmb.1742. PMC 2933787. PMID 20010837.
- ^ a b "Entry - *612532 - THAP Doman-Containing Protein 3; THAP3 - OMIM". www.omim.org. Retrieved 2022-12-15.
- ^ a b "THAP domain-containing protein 3 isoform 1 [Homo sapiens] - Protein - NCBI". National Center of Biotechnology Information. Retrieved 2022-12-16.
- ^ a b Fagerberg L, Hallström BM, Oksvold P, Kampf C, Djureinovic D, Odeberg J, et al. (February 2014). "Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics". Molecular & Cellular Proteomics. 13 (2): 397–406. doi: 10.1074/mcp.M113.035600. PMC 3916642. PMID 24309898.
- ^ Duff MO, Olson S, Wei X, Garrett SC, Osman A, Bolisetty M, et al. (May 2015). "Genome-wide identification of zero nucleotide recursive splicing in Drosophila". Nature. 521 (7552): 376–379. Bibcode: 2015Natur.521..376D. doi: 10.1038/nature14475. PMC 4529404. PMID 25970244.
- ^ a b c d e f g h i j k "Protein BLAST: search protein databases using a protein query". National Center of Biotechnology Information. Retrieved 2022-12-08.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 1, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 2, m - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. 22 April 2022.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 3, m - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. 10 June 2022.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 4, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 5, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 6, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 7, m - Nucleotide - NCBI". www.ncbi.nlm.nih.gov. 22 April 2022.
- ^ "Homo sapiens THAP domain containing 3 (THAP3), transcript variant 8, mRNA". April 22, 2022 – via NCBI Nucleotide.
- ^ a b Brendel V, Bucher P, Nourbakhsh IR, Blaisdell BE, Karlin S (March 1992). "Methods and algorithms for statistical analysis of protein sequences". Proceedings of the National Academy of Sciences of the United States of America. 89 (6): 2002–2006. Bibcode: 1992PNAS...89.2002B. doi: 10.1073/pnas.89.6.2002. PMC 48584. PMID 1549558.
- ^ Gasteiger E., Hoogland C., Gattiker A., Duvaud S., Wilkins M.R., Appel R.D., Bairoch A.; Protein Identification and Analysis Tools on the Expasy Server; (In) John M. Walker (ed): The Proteomics Protocols Handbook, Humana Press (2005).
- ^ "THAP domain-containing protein 3 isoform 1 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 2 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 3 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 4 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 5 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 6 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 7 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform 8 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ Liu W, Xie Y, Ma J, Luo X, Nie P, Zuo Z, et al. (October 2015). "IBS: an illustrator for the presentation and visualization of biological sequences". Bioinformatics. 31 (20): 3359–3361. doi: 10.1093/bioinformatics/btv362. PMC 4595897. PMID 26069263.
- ^ a b c Gupta, R. (2001). Prediction of glycosylation sites in proteomes: from post-translational modifications to protein function. Technical University of Denmark.
- ^ a b Blom N, Gammeltoft S, Brunak S (December 1999). "Sequence and structure-based prediction of eukaryotic protein phosphorylation sites". Journal of Molecular Biology. 294 (5): 1351–1362. doi: 10.1006/jmbi.1999.3310. PMID 10600390.
- ^ a b Wang J, Youkharibache P, Marchler-Bauer A, Lanczycki C, Zhang D, Lu S, et al. (2022). "iCn3D: From Web-Based 3D Viewer to Structural Analysis Tool in Batch Mode". Frontiers in Molecular Biosciences. 9: 831740. doi: 10.3389/fmolb.2022.831740. PMC 8892267. PMID 35252351.
- ^ "PSORT II Prediction". psort.hgc.jp. Retrieved 2022-12-16.
- ^ Filtz TM, Vogel WK, Leid M (February 2014). "Regulation of transcription factor activity by interconnected post-translational modifications". Trends in Pharmacological Sciences. 35 (2): 76–85. doi: 10.1016/j.tips.2013.11.005. PMC 3954851. PMID 24388790.
- ^ Sanghavi HM, Mallajosyula SS, Majumdar S (March 2019). "Classification of the human THAP protein family identifies an evolutionarily conserved coiled coil region". BMC Structural Biology. 19 (1): 4. doi: 10.1186/s12900-019-0102-2. PMC 6402169. PMID 30836974.
- ^ "THAP1 THAP domain containing 1 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP2 THAP domain containing 2 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP5 THAP domain containing 5 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP6 THAP domain containing 6 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP7 THAP domain containing 7 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP8 THAP domain containing 8 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ "THAP9 THAP domain containing 9 [Homo sapiens (human)] - Gene - NCBI". National Center of Biotechnology Information.
- ^ Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, et al. (October 2011). "Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega". Molecular Systems Biology. 7 (1): 539. doi: 10.1038/msb.2011.75. PMC 3261699. PMID 21988835.
- ^ "THAP domain-containing protein 3 isoform 1 [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2022-12-08.
- ^ Kumar S, Suleski M, Craig JM, Kasprowicz AE, Sanderford M, Li M, et al. (August 2022). "TimeTree 5: An Expanded Resource for Species Divergence Times". Molecular Biology and Evolution. 39 (8): msac174. doi: 10.1093/molbev/msac174. PMC 9400175. PMID 35932227.
- ^ "EMBOSS Needle < Pairwise Sequence Alignment < EMBL-EBI". www.ebi.ac.uk. Retrieved 2022-12-08.
- ^ "THAP domain-containing protein 3 isoform X1 [Marmota flaviventris] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Lontra canadensis] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Eptesicus fuscus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Balaenoptera musculus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Dromiciops gliroides] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Phascolarctos cinereus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Caretta caretta] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Gopherus evgoodei] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Chelonoidis abingdonii] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Mauremys mutica] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Microcaecilia unicolor] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Geotrypetes seraphini] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Electrophorus electricus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Coregonus clupeaformis] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Brienomyrus brachyistius] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Puntigrus tetrazona] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Rhincodon typus] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Chiloscyllium plagiosum] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "THAP domain-containing protein 3 isoform X1 [Amblyraja radiata] - Protein - NCBI". www.ncbi.nlm.nih.gov.
- ^ "UniProt". www.uniprot.org. Retrieved 2022-12-16.
- ^ "IntAct Portal". www.ebi.ac.uk. Retrieved 2022-12-16.
- ^ a b "THAP3 Result Summary | BioGRID". thebiogrid.org. Retrieved 2022-12-16.
- ^ "THAP3 Gene - GeneCards | THAP3 Protein | THAP3 Antibody". www.genecards.org. Retrieved 2022-12-08.
- ^ a b Rosales RL (October 2010). "X-linked dystonia parkinsonism: clinical phenotype, genetics and therapeutics". Journal of Movement Disorders. 3 (2): 32–38. doi: 10.14802/jmd.10009. PMC 4027667. PMID 24868378.



