| Envelope protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
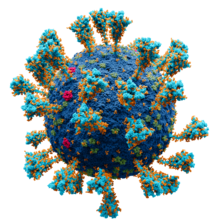 Model of the external structure of the SARS-CoV-2
virion
[1]
● Blue: envelope ● Turquoise: spike glycoprotein (S) ● Bright Pink: envelope proteins (E) ● Green: membrane proteins (M) ● Orange: glycans | |||||||||
| Identifiers | |||||||||
| Symbol | CoV_E | ||||||||
| Pfam | PF02723 | ||||||||
| InterPro | IPR003873 | ||||||||
| PROSITE | PS51926 | ||||||||
| |||||||||
The envelope (E) protein is the smallest and least well-characterized of the four major structural proteins found in coronavirus virions. [2] [3] [4] It is an integral membrane protein less than 110 amino acid residues long; [2] in SARS-CoV-2, the causative agent of Covid-19, the E protein is 75 residues long. [5] Although it is not necessarily essential for viral replication, absence of the E protein may produce abnormally assembled viral capsids or reduced replication. [2] [3] E is a multifunctional protein [6] and, in addition to its role as a structural protein in the viral capsid, it is thought to be involved in viral assembly, likely functions as a viroporin, and is involved in viral pathogenesis. [2] [5]
Structure

The E protein consists of a short hydrophilic N-terminal region, a hydrophobic helical transmembrane domain, and a somewhat hydrophilic C-terminal region. In SARS-CoV and SARS-CoV-2, the C-terminal region contains a PDZ-binding motif (PBM). [2] [5] This feature appears to be conserved only in the alpha and beta coronavirus groups, but not gamma. [2] In the beta and gamma groups, a conserved proline residue is found in the C-terminal region likely involved in targeting the protein to the Golgi. [2]
The transmembrane helices of the E proteins of SARS-CoV and SARS-CoV-2 can oligomerize and have been shown in vitro to form pentameric structures with central pores that serve as cation-selective ion channels. [5] Both viruses' E protein pentamers have been structurally characterized by nuclear magnetic resonance spectroscopy. [5] [7]
The membrane topology of the E protein has been studied in a number of coronaviruses with inconsistent results; the protein's orientation in the membrane may be variable. [3] The balance of evidence suggests the most common orientation has the C-terminus oriented toward the cytoplasm. [8] Studies of SARS-CoV-2 E protein are consistent with this orientation. [5] [9]
Post-translational modifications
In some, but not all, coronaviruses, the E protein is post-translationally modified by palmitoylation on conserved cysteine residues. [2] [8] In the SARS-CoV E protein, one glycosylation site has been observed, which may influence membrane topology; [8] however, the functional significance of E glycosylation is unclear. [2] Ubiquitination of SARS-CoV E has also been described, though its functional significance is also not known. [2]
Expression and localization
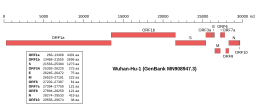
Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2, indicating the location of the E gene | |
| NCBI genome ID | 86693 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
| Genome browser ( UCSC) | |
The E protein is expressed at high abundance in infected cells. However, only a small amount of the total E protein produced is found in assembled virions. [2] [4] E protein is localized to the endoplasmic reticulum, Golgi apparatus, and endoplasmic-reticulum–Golgi intermediate compartment (ERGIC), the intracellular compartment that gives rise to the coronavirus viral envelope. [2] [5]
Function
Essentiality
Studies in different coronaviruses have reached different conclusions about whether E is essential to viral replication. In some coronaviruses, including MERS-CoV, E has been reported to be essential. [10] In others, including mouse coronavirus [11] and SARS-CoV, E is not essential, though its absence reduces viral titer, [12] in some cases by introducing propagation defects or causing abnormal capsid morphology. [2]
Virions and viral assembly

The E protein is found in assembled virions where it forms protein-protein interactions with the coronavirus membrane protein (M), the most abundant of the four structural proteins contained in the viral capsid. [2] [4] The interaction between E and M occurs through their respective C-termini on the cytoplasmic side of the membrane. [2] In most coronaviruses, E and M are sufficient to form virus-like particles, [2] [4] though SARS-CoV has been reported to depend on N as well. [14] There is good evidence that E is involved in inducing membrane curvature to create the typical spherical coronavirus virion. [2] [15] It is likely that E is involved in viral budding or scission, although its role in this process has not been well characterized. [2] [4] [15]
Viroporin
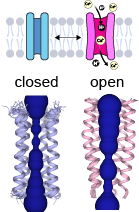
In its pentameric state, E forms cation-selective ion channels and likely functions as a viroporin. [5] NMR studies show that viroporin presents an open conformation at low pH or in the presence of calcium ions, while the closed conformation is favored at basic pH. [16] The NMR structure shows a hydrophobic gate at leucine 28 in the middle of the pore. The passage of ions through the gate is thought to be facilitated by the polar residues at the C-terminus. [17]
The cation leakage may disrupt ion homeostasis, alter membrane permeability, and modulate pH in the host cell, which may facilitate viral release. [2] [4]
The E protein's role as a viroporin appears to be involved in
pathogenesis and may be related to activation of the
inflammasome.
[3]
[18] In SARS-CoV, mutations that disrupt E's ion channel function result in attenuated pathogenesis in
animal models despite little effect on viral growth.
[10]
Interactions with host proteins

Protein-protein interactions between E and proteins in the host cell are best described in SARS-CoV and occur via the C-terminal PDZ domain binding motif. The SARS-CoV E protein has been reported to interact with five host cell proteins: Bcl-xL, PALS1, syntenin, sodium/potassium (Na+/K+) ATPase α-1 subunit, and stomatin. [2] The interaction with PALS1 may be related to pathogenesis via the resulting disruption in tight junctions. [3] [10] This interaction has also been identified in SARS-CoV-2. [19]
Evolution and conservation
The sequence of the E protein is not well conserved across coronavirus genera, with sequence identities reaching under 30%. [12] In laboratory experiments on mouse hepatitis virus, substitution of E proteins from different coronaviruses, even from different groups, could produce viable viruses, suggesting that significant sequence diversity can be tolerated in functional E proteins. [20] The SARS-CoV-2 E protein is very similar to that of SARS-CoV, with three substitutions and one deletion. [4] A study of SARS-CoV-2 sequences suggests that the E protein is evolving relatively slowly compared to other structural proteins. [21] The conserved nature of the envelope protein among SARS-CoV and SARS-CoV-2 variants has led it to be researched as a potential target for universal coronavirus vaccine development. [22] [23]
References
- ^ Solodovnikov, Alexey; Arkhipova, Valeria (2021-07-29). "Достоверно красиво: как мы сделали 3D-модель SARS-CoV-2" [Truly beautiful: how we made the SARS-CoV-2 3D model] (in Russian). N+1. Archived from the original on 2021-07-30. Retrieved 30 July 2021.
- ^ a b c d e f g h i j k l m n o p q r s t Schoeman D, Fielding BC (May 2019). "Coronavirus envelope protein: current knowledge". Virology Journal. 16 (1): 69. doi: 10.1186/s12985-019-1182-0. PMC 6537279. PMID 31133031.
- ^ a b c d e Schoeman D, Fielding BC (2020-09-03). "Is There a Link Between the Pathogenic Human Coronavirus Envelope Protein and Immunopathology? A Review of the Literature". Frontiers in Microbiology. 11: 2086. doi: 10.3389/fmicb.2020.02086. PMC 7496634. PMID 33013759.
- ^ a b c d e f g h Cao Y, Yang R, Lee I, Zhang W, Sun J, Wang W, Meng X (June 2021). "Characterization of the SARS-CoV-2 E Protein: Sequence, Structure, Viroporin, and Inhibitors". Protein Science. 30 (6): 1114–1130. doi: 10.1002/pro.4075. PMC 8138525. PMID 33813796.
- ^ a b c d e f g h i Mandala VS, McKay MJ, Shcherbakov AA, Dregni AJ, Kolocouris A, Hong M (December 2020). "Structure and drug binding of the SARS-CoV-2 envelope protein transmembrane domain in lipid bilayers". Nature Structural & Molecular Biology. 27 (12): 1202–1208. doi: 10.1038/s41594-020-00536-8. PMC 7718435. PMID 33177698.
- ^ Liu DX, Yuan Q, Liao Y (August 2007). "Coronavirus envelope protein: a small membrane protein with multiple functions". Cellular and Molecular Life Sciences. 64 (16): 2043–2048. doi: 10.1007/s00018-007-7103-1. PMC 7079843. PMID 17530462.
- ^ Surya W, Li Y, Torres J (June 2018). "Structural model of the SARS coronavirus E channel in LMPG micelles". Biochimica et Biophysica Acta (BBA) - Biomembranes. 1860 (6): 1309–1317. doi: 10.1016/j.bbamem.2018.02.017. PMC 7094280. PMID 29474890.
- ^ a b c Fung TS, Liu DX (June 2018). "Post-translational modifications of coronavirus proteins: roles and function". Future Virology. 13 (6): 405–430. doi: 10.2217/fvl-2018-0008. PMC 7080180. PMID 32201497.
- ^ Duart G, García-Murria MJ, Grau B, Acosta-Cáceres JM, Martínez-Gil L, Mingarro I (September 2020). "SARS-CoV-2 envelope protein topology in eukaryotic membranes". Open Biology. 10 (9): 200209. doi: 10.1098/rsob.200209. PMC 7536074. PMID 32898469.
- ^ a b c DeDiego ML, Nieto-Torres JL, Jimenez-Guardeño JM, Regla-Nava JA, Castaño-Rodriguez C, Fernandez-Delgado R, et al. (December 2014). "Coronavirus virulence genes with main focus on SARS-CoV envelope gene". Virus Research. 194: 124–137. doi: 10.1016/j.virusres.2014.07.024. PMC 4261026. PMID 25093995.
- ^ Kuo L, Masters PS (April 2003). "The small envelope protein E is not essential for murine coronavirus replication". Journal of Virology. 77 (8): 4597–4608. doi: 10.1128/JVI.77.8.4597-4608.2003. PMC 152126. PMID 12663766.
- ^ a b Ruch TR, Machamer CE (March 2012). "The coronavirus E protein: assembly and beyond". Viruses. 4 (3): 363–382. doi: 10.3390/v4030363. PMC 3347032. PMID 22590676.
- ^ Goodsell DS, Voigt M, Zardecki C, Burley SK (August 2020). "Integrative illustration for coronavirus outreach". PLOS Biology. 18 (8): e3000815. doi: 10.1371/journal.pbio.3000815. PMC 7433897. PMID 32760062.
- ^ Siu YL, Teoh KT, Lo J, Chan CM, Kien F, Escriou N, et al. (November 2008). "The M, E, and N structural proteins of the severe acute respiratory syndrome coronavirus are required for efficient assembly, trafficking, and release of virus-like particles". Journal of Virology. 82 (22): 11318–11330. doi: 10.1128/JVI.01052-08. PMC 2573274. PMID 18753196.
- ^ a b J Alsaadi EA, Jones IM (April 2019). "Membrane binding proteins of coronaviruses". Future Virology. 14 (4): 275–286. doi: 10.2217/fvl-2018-0144. PMC 7079996. PMID 32201500.
- ^ Medeiros-Silva J, Somberg NH, Wang HK, McKay MJ, Mandala VS, Dregni AJ, Hong M (April 2022). "pH- and Calcium-Dependent Aromatic Network in the SARS-CoV-2 Envelope Protein". Journal of the American Chemical Society. 144 (15): 6839–6850. doi: 10.1021/jacs.2c00973. PMC 9188436. PMID 35380805.
- ^ Medeiros-Silva J, Dregni AJ, Somberg NH, Duan P, Hong M (October 2023). "Atomic structure of the open SARS-CoV-2 E viroporin". Science Advances. 9 (41): eadi9007. doi: 10.1126/sciadv.adi9007. PMC 10575589. PMID 37831764.
- ^ Nieto-Torres JL, DeDiego ML, Verdiá-Báguena C, Jimenez-Guardeño JM, Regla-Nava JA, Fernandez-Delgado R, et al. (May 2014). "Severe acute respiratory syndrome coronavirus envelope protein ion channel activity promotes virus fitness and pathogenesis". PLOS Pathogens. 10 (5): e1004077. doi: 10.1371/journal.ppat.1004077. PMC 4006877. PMID 24788150.
- ^ a b Chai J, Cai Y, Pang C, Wang L, McSweeney S, Shanklin J, Liu Q (June 2021). "Structural basis for SARS-CoV-2 envelope protein recognition of human cell junction protein PALS1". Nature Communications. 12 (1): 3433. Bibcode: 2021NatCo..12.3433C. doi: 10.1038/s41467-021-23533-x. PMC 8187709. PMID 34103506.
- ^ Kuo L, Hurst KR, Masters PS (March 2007). "Exceptional flexibility in the sequence requirements for coronavirus small envelope protein function". Journal of Virology. 81 (5): 2249–2262. doi: 10.1128/JVI.01577-06. PMC 1865940. PMID 17182690.
- ^ Rahman MS, Hoque MN, Islam MR, Islam I, Mishu ID, Rahaman MM, et al. (March 2021). "Mutational insights into the envelope protein of SARS-CoV-2". Gene Reports. 22: 100997. doi: 10.1016/j.genrep.2020.100997. PMC 7723457. PMID 33319124.
- ^ Bhattacharya S, Banerjee A, Ray S (March 2021). "Development of new vaccine target against SARS-CoV2 using envelope (E) protein: An evolutionary, molecular modeling and docking based study". International Journal of Biological Macromolecules. 172: 74–81. doi: 10.1016/j.ijbiomac.2020.12.192. PMC 7833863. PMID 33385461.
- ^ Chen J, Deng Y, Huang B, Han D, Wang W, Huang M, et al. (2022-02-24). "DNA Vaccines Expressing the Envelope and Membrane Proteins Provide Partial Protection Against SARS-CoV-2 in Mice". Frontiers in Immunology. 13: 827605. doi: 10.3389/fimmu.2022.827605. PMC 8907653. PMID 35281016.
| Envelope protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Model of the external structure of the SARS-CoV-2
virion
[1]
● Blue: envelope ● Turquoise: spike glycoprotein (S) ● Bright Pink: envelope proteins (E) ● Green: membrane proteins (M) ● Orange: glycans | |||||||||
| Identifiers | |||||||||
| Symbol | CoV_E | ||||||||
| Pfam | PF02723 | ||||||||
| InterPro | IPR003873 | ||||||||
| PROSITE | PS51926 | ||||||||
| |||||||||
The envelope (E) protein is the smallest and least well-characterized of the four major structural proteins found in coronavirus virions. [2] [3] [4] It is an integral membrane protein less than 110 amino acid residues long; [2] in SARS-CoV-2, the causative agent of Covid-19, the E protein is 75 residues long. [5] Although it is not necessarily essential for viral replication, absence of the E protein may produce abnormally assembled viral capsids or reduced replication. [2] [3] E is a multifunctional protein [6] and, in addition to its role as a structural protein in the viral capsid, it is thought to be involved in viral assembly, likely functions as a viroporin, and is involved in viral pathogenesis. [2] [5]
Structure

The E protein consists of a short hydrophilic N-terminal region, a hydrophobic helical transmembrane domain, and a somewhat hydrophilic C-terminal region. In SARS-CoV and SARS-CoV-2, the C-terminal region contains a PDZ-binding motif (PBM). [2] [5] This feature appears to be conserved only in the alpha and beta coronavirus groups, but not gamma. [2] In the beta and gamma groups, a conserved proline residue is found in the C-terminal region likely involved in targeting the protein to the Golgi. [2]
The transmembrane helices of the E proteins of SARS-CoV and SARS-CoV-2 can oligomerize and have been shown in vitro to form pentameric structures with central pores that serve as cation-selective ion channels. [5] Both viruses' E protein pentamers have been structurally characterized by nuclear magnetic resonance spectroscopy. [5] [7]
The membrane topology of the E protein has been studied in a number of coronaviruses with inconsistent results; the protein's orientation in the membrane may be variable. [3] The balance of evidence suggests the most common orientation has the C-terminus oriented toward the cytoplasm. [8] Studies of SARS-CoV-2 E protein are consistent with this orientation. [5] [9]
Post-translational modifications
In some, but not all, coronaviruses, the E protein is post-translationally modified by palmitoylation on conserved cysteine residues. [2] [8] In the SARS-CoV E protein, one glycosylation site has been observed, which may influence membrane topology; [8] however, the functional significance of E glycosylation is unclear. [2] Ubiquitination of SARS-CoV E has also been described, though its functional significance is also not known. [2]
Expression and localization

Genomic organisation of isolate Wuhan-Hu-1, the earliest sequenced sample of SARS-CoV-2, indicating the location of the E gene | |
| NCBI genome ID | 86693 |
|---|---|
| Genome size | 29,903 bases |
| Year of completion | 2020 |
| Genome browser ( UCSC) | |
The E protein is expressed at high abundance in infected cells. However, only a small amount of the total E protein produced is found in assembled virions. [2] [4] E protein is localized to the endoplasmic reticulum, Golgi apparatus, and endoplasmic-reticulum–Golgi intermediate compartment (ERGIC), the intracellular compartment that gives rise to the coronavirus viral envelope. [2] [5]
Function
Essentiality
Studies in different coronaviruses have reached different conclusions about whether E is essential to viral replication. In some coronaviruses, including MERS-CoV, E has been reported to be essential. [10] In others, including mouse coronavirus [11] and SARS-CoV, E is not essential, though its absence reduces viral titer, [12] in some cases by introducing propagation defects or causing abnormal capsid morphology. [2]
Virions and viral assembly

The E protein is found in assembled virions where it forms protein-protein interactions with the coronavirus membrane protein (M), the most abundant of the four structural proteins contained in the viral capsid. [2] [4] The interaction between E and M occurs through their respective C-termini on the cytoplasmic side of the membrane. [2] In most coronaviruses, E and M are sufficient to form virus-like particles, [2] [4] though SARS-CoV has been reported to depend on N as well. [14] There is good evidence that E is involved in inducing membrane curvature to create the typical spherical coronavirus virion. [2] [15] It is likely that E is involved in viral budding or scission, although its role in this process has not been well characterized. [2] [4] [15]
Viroporin

In its pentameric state, E forms cation-selective ion channels and likely functions as a viroporin. [5] NMR studies show that viroporin presents an open conformation at low pH or in the presence of calcium ions, while the closed conformation is favored at basic pH. [16] The NMR structure shows a hydrophobic gate at leucine 28 in the middle of the pore. The passage of ions through the gate is thought to be facilitated by the polar residues at the C-terminus. [17]
The cation leakage may disrupt ion homeostasis, alter membrane permeability, and modulate pH in the host cell, which may facilitate viral release. [2] [4]
The E protein's role as a viroporin appears to be involved in
pathogenesis and may be related to activation of the
inflammasome.
[3]
[18] In SARS-CoV, mutations that disrupt E's ion channel function result in attenuated pathogenesis in
animal models despite little effect on viral growth.
[10]
Interactions with host proteins

Protein-protein interactions between E and proteins in the host cell are best described in SARS-CoV and occur via the C-terminal PDZ domain binding motif. The SARS-CoV E protein has been reported to interact with five host cell proteins: Bcl-xL, PALS1, syntenin, sodium/potassium (Na+/K+) ATPase α-1 subunit, and stomatin. [2] The interaction with PALS1 may be related to pathogenesis via the resulting disruption in tight junctions. [3] [10] This interaction has also been identified in SARS-CoV-2. [19]
Evolution and conservation
The sequence of the E protein is not well conserved across coronavirus genera, with sequence identities reaching under 30%. [12] In laboratory experiments on mouse hepatitis virus, substitution of E proteins from different coronaviruses, even from different groups, could produce viable viruses, suggesting that significant sequence diversity can be tolerated in functional E proteins. [20] The SARS-CoV-2 E protein is very similar to that of SARS-CoV, with three substitutions and one deletion. [4] A study of SARS-CoV-2 sequences suggests that the E protein is evolving relatively slowly compared to other structural proteins. [21] The conserved nature of the envelope protein among SARS-CoV and SARS-CoV-2 variants has led it to be researched as a potential target for universal coronavirus vaccine development. [22] [23]
References
- ^ Solodovnikov, Alexey; Arkhipova, Valeria (2021-07-29). "Достоверно красиво: как мы сделали 3D-модель SARS-CoV-2" [Truly beautiful: how we made the SARS-CoV-2 3D model] (in Russian). N+1. Archived from the original on 2021-07-30. Retrieved 30 July 2021.
- ^ a b c d e f g h i j k l m n o p q r s t Schoeman D, Fielding BC (May 2019). "Coronavirus envelope protein: current knowledge". Virology Journal. 16 (1): 69. doi: 10.1186/s12985-019-1182-0. PMC 6537279. PMID 31133031.
- ^ a b c d e Schoeman D, Fielding BC (2020-09-03). "Is There a Link Between the Pathogenic Human Coronavirus Envelope Protein and Immunopathology? A Review of the Literature". Frontiers in Microbiology. 11: 2086. doi: 10.3389/fmicb.2020.02086. PMC 7496634. PMID 33013759.
- ^ a b c d e f g h Cao Y, Yang R, Lee I, Zhang W, Sun J, Wang W, Meng X (June 2021). "Characterization of the SARS-CoV-2 E Protein: Sequence, Structure, Viroporin, and Inhibitors". Protein Science. 30 (6): 1114–1130. doi: 10.1002/pro.4075. PMC 8138525. PMID 33813796.
- ^ a b c d e f g h i Mandala VS, McKay MJ, Shcherbakov AA, Dregni AJ, Kolocouris A, Hong M (December 2020). "Structure and drug binding of the SARS-CoV-2 envelope protein transmembrane domain in lipid bilayers". Nature Structural & Molecular Biology. 27 (12): 1202–1208. doi: 10.1038/s41594-020-00536-8. PMC 7718435. PMID 33177698.
- ^ Liu DX, Yuan Q, Liao Y (August 2007). "Coronavirus envelope protein: a small membrane protein with multiple functions". Cellular and Molecular Life Sciences. 64 (16): 2043–2048. doi: 10.1007/s00018-007-7103-1. PMC 7079843. PMID 17530462.
- ^ Surya W, Li Y, Torres J (June 2018). "Structural model of the SARS coronavirus E channel in LMPG micelles". Biochimica et Biophysica Acta (BBA) - Biomembranes. 1860 (6): 1309–1317. doi: 10.1016/j.bbamem.2018.02.017. PMC 7094280. PMID 29474890.
- ^ a b c Fung TS, Liu DX (June 2018). "Post-translational modifications of coronavirus proteins: roles and function". Future Virology. 13 (6): 405–430. doi: 10.2217/fvl-2018-0008. PMC 7080180. PMID 32201497.
- ^ Duart G, García-Murria MJ, Grau B, Acosta-Cáceres JM, Martínez-Gil L, Mingarro I (September 2020). "SARS-CoV-2 envelope protein topology in eukaryotic membranes". Open Biology. 10 (9): 200209. doi: 10.1098/rsob.200209. PMC 7536074. PMID 32898469.
- ^ a b c DeDiego ML, Nieto-Torres JL, Jimenez-Guardeño JM, Regla-Nava JA, Castaño-Rodriguez C, Fernandez-Delgado R, et al. (December 2014). "Coronavirus virulence genes with main focus on SARS-CoV envelope gene". Virus Research. 194: 124–137. doi: 10.1016/j.virusres.2014.07.024. PMC 4261026. PMID 25093995.
- ^ Kuo L, Masters PS (April 2003). "The small envelope protein E is not essential for murine coronavirus replication". Journal of Virology. 77 (8): 4597–4608. doi: 10.1128/JVI.77.8.4597-4608.2003. PMC 152126. PMID 12663766.
- ^ a b Ruch TR, Machamer CE (March 2012). "The coronavirus E protein: assembly and beyond". Viruses. 4 (3): 363–382. doi: 10.3390/v4030363. PMC 3347032. PMID 22590676.
- ^ Goodsell DS, Voigt M, Zardecki C, Burley SK (August 2020). "Integrative illustration for coronavirus outreach". PLOS Biology. 18 (8): e3000815. doi: 10.1371/journal.pbio.3000815. PMC 7433897. PMID 32760062.
- ^ Siu YL, Teoh KT, Lo J, Chan CM, Kien F, Escriou N, et al. (November 2008). "The M, E, and N structural proteins of the severe acute respiratory syndrome coronavirus are required for efficient assembly, trafficking, and release of virus-like particles". Journal of Virology. 82 (22): 11318–11330. doi: 10.1128/JVI.01052-08. PMC 2573274. PMID 18753196.
- ^ a b J Alsaadi EA, Jones IM (April 2019). "Membrane binding proteins of coronaviruses". Future Virology. 14 (4): 275–286. doi: 10.2217/fvl-2018-0144. PMC 7079996. PMID 32201500.
- ^ Medeiros-Silva J, Somberg NH, Wang HK, McKay MJ, Mandala VS, Dregni AJ, Hong M (April 2022). "pH- and Calcium-Dependent Aromatic Network in the SARS-CoV-2 Envelope Protein". Journal of the American Chemical Society. 144 (15): 6839–6850. doi: 10.1021/jacs.2c00973. PMC 9188436. PMID 35380805.
- ^ Medeiros-Silva J, Dregni AJ, Somberg NH, Duan P, Hong M (October 2023). "Atomic structure of the open SARS-CoV-2 E viroporin". Science Advances. 9 (41): eadi9007. doi: 10.1126/sciadv.adi9007. PMC 10575589. PMID 37831764.
- ^ Nieto-Torres JL, DeDiego ML, Verdiá-Báguena C, Jimenez-Guardeño JM, Regla-Nava JA, Fernandez-Delgado R, et al. (May 2014). "Severe acute respiratory syndrome coronavirus envelope protein ion channel activity promotes virus fitness and pathogenesis". PLOS Pathogens. 10 (5): e1004077. doi: 10.1371/journal.ppat.1004077. PMC 4006877. PMID 24788150.
- ^ a b Chai J, Cai Y, Pang C, Wang L, McSweeney S, Shanklin J, Liu Q (June 2021). "Structural basis for SARS-CoV-2 envelope protein recognition of human cell junction protein PALS1". Nature Communications. 12 (1): 3433. Bibcode: 2021NatCo..12.3433C. doi: 10.1038/s41467-021-23533-x. PMC 8187709. PMID 34103506.
- ^ Kuo L, Hurst KR, Masters PS (March 2007). "Exceptional flexibility in the sequence requirements for coronavirus small envelope protein function". Journal of Virology. 81 (5): 2249–2262. doi: 10.1128/JVI.01577-06. PMC 1865940. PMID 17182690.
- ^ Rahman MS, Hoque MN, Islam MR, Islam I, Mishu ID, Rahaman MM, et al. (March 2021). "Mutational insights into the envelope protein of SARS-CoV-2". Gene Reports. 22: 100997. doi: 10.1016/j.genrep.2020.100997. PMC 7723457. PMID 33319124.
- ^ Bhattacharya S, Banerjee A, Ray S (March 2021). "Development of new vaccine target against SARS-CoV2 using envelope (E) protein: An evolutionary, molecular modeling and docking based study". International Journal of Biological Macromolecules. 172: 74–81. doi: 10.1016/j.ijbiomac.2020.12.192. PMC 7833863. PMID 33385461.
- ^ Chen J, Deng Y, Huang B, Han D, Wang W, Huang M, et al. (2022-02-24). "DNA Vaccines Expressing the Envelope and Membrane Proteins Provide Partial Protection Against SARS-CoV-2 in Mice". Frontiers in Immunology. 13: 827605. doi: 10.3389/fimmu.2022.827605. PMC 8907653. PMID 35281016.
