| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 5 | ← | Archive 7 | Archive 8 | Archive 9 | Archive 10 | Archive 11 |
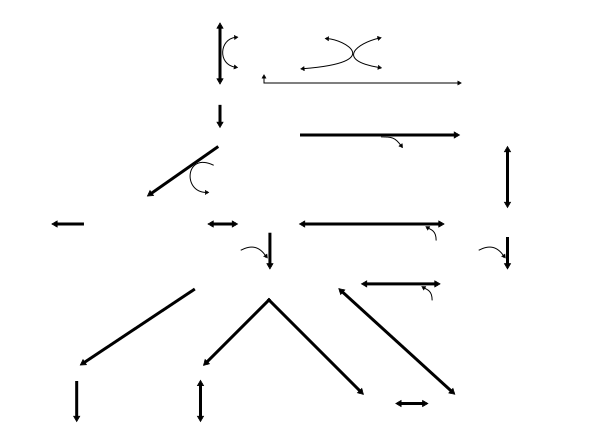
I recently removed all of the image text from File:ISSN HMB statement Fig 1.jpg and replaced it with wikitext. The background image file is likely going to be changed to an svg format soon. Consequently, I'm open to restructuring the background image and repositioning/resizing the annotations based upon feedback from this project.
The original image file that this diagram is based upon was uploaded from a CC-BY-SA 2.0-copyrighted journal article about β-hydroxy β-methylbutyric acid (HMB), hence why there's a lot of emphasis on HMB in this image (note that there are a few minor differences between this image and the original). Unfortunately, the enzymes that catalyze the reactions HMB ↔ HMB-CoA and HMB-CoA ↔ HMG-CoA aren't currently known. Also, some very minor metabolites of leucine aren't shown in this image (L-Leucine → β-Leucine via leucine aminomutase, and β-Leucine → β-ketoisocaproic acid → β-Ketoisocaproyl-CoA → acetyl-CoA via a sequence of uncharacterized enzymes). This textbook diagram illustrates that tertiary pathway. Seppi333 ( Insert 2¢) 19:01, 2 December 2017 (UTC)
- One thing I'm not sure about is if it's correct to write "biotin+CO2" and "CO2+biotin" in this image as an input for MC-CoA carboxylase, or if it should say "CO2" as an input and " cofactor: biotin" next to the line beneath that substrate. Are either of those approaches fine? Seppi333 ( Insert 2¢) 20:07, 2 December 2017 (UTC)
- If anyone can comment on how to illustrate biotin as a cofactor for MC-CoA carboxylase in this image, I'd really appreciate it. Seppi333 ( Insert 2¢) 07:24, 10 December 2017 (UTC)
- Seppi333 Is it necessary to illustrate biotin? As a prosthetic group, it's really just part of the MC-CoA carboxylase holoenzyme. It's different to something like NADH or ATP where the cofactor is different before and after the reaction, and you need some separate process to regenerate it. But if you want to illustrate it, I'd suggest something like the original image, which doesn't give the impression that biotin is consumed like CO2 is. You could also just note the biotin requirement in the image caption.
- Also, is there a reason that reaction and its substrate and product appear in the diagram twice? As in, shouldn't the right-pointing arrow from HMB-CoA point to MC-CoA over on the right edge of the diagram, rather than there being a duplicate near the middle?
Adrian J. Hunter(
talk•
contribs)
04:30, 12 December 2017 (UTC)
- It probably isn't necessary TBH. I think the original author just included it because of the "when biotin is deficient" text in the image. I'll delete biotin from the image since you don't think it's particularly necessary to illustrate. As for why there's two pathways to HMG in the image... I'm not sure why it was illustrated that way. That's actually what I was hoping others would comment on. If you think the redundant MC-CoA and MG-CoA pathway in the bottom-center part of the image should come out, I'm okay with deleting that when the image is redrawn. Seppi333 ( Insert 2¢) 05:53, 12 December 2017 (UTC)
Given that the HMB article has been renominated at FAC for the 4th time after a 12 month wait, further feedback on changes to this diagram for the svg version would be very much appreciated. If you think some part of this diagram should be drawn, depicted, or annotated in a different manner, please say so. I'm looking for both general/non-specific feedback as well as specific feedback on one technical and one cosmetic point:
- The technical issue I'm looking for further feedback on is whether or not cofactors should be indicated and how to illustrate cofactors and/or inputs to each reaction if they're included in the image; i.e., should the inputs be depicted as they are now in the svg version and (re: Adrian's comment above) should biotin be annotated in the same manner as it was in the original image or be omitted? NB: the biotin annotation wasn't included in the current annotated image.
- The cosmetic issue I'm looking for further feedback on is whether the two pathways for HMB↔HMB-CoA↔MC-CoA↔MG-CoA↔HMG-CoA should be illustrated in the svg version, analogous to how it is in the current image, or if these should be merged into a single depiction when the image is redrawn as an svg diagram.
For additional context/information on the pathway being depicted in this diagram, it would probably help to read the article text in Leucine#Metabolism in humans. Seppi333 ( Insert 2¢) 08:22, 21 January 2018 (UTC)
- An essential requirement for Featured Articles is that it should be understood by a wide audience.IMHO the figure is way too complicated (i.e., likely to confuse readers) and should be simplified as much as possible without removing essential information. Hence I think co-factors should left out, unless they are directly referred to in the text (e.g., biotin). Splitting the digram into biosynthesis and metabolism will go a long way in making it easier to digest. Also the lower right hand side (MG-CoA → HMG-CoA → Acetyl-CoA/Acetoacetate) need not be included in either digram since this pathway has to do with metabolism of leucine, not HMB.
Boghog (
talk)
10:05, 21 January 2018 (UTC)
- Hehe, well I guess this speaks to how complicated the image is right now.
 The bottom right portion of this image is showing the metabolism of HMG-CoA, which is depicted as being downstream from isovaleryl-CoA; but, you'll notice that the pathway on the left also shows HMB-CoA being converted to both MC-CoA and HMG-CoA, and both of those are also depicted in the pathway on the right side (NB: if you click on the HMG-CoA, MC-CoA, or MG-CoA in this diagram and then view this image in those articles, you'll see 2 highlighted annotations for each of those pairs). Because HMG-CoA is downstream from HMB, the metabolism of HMG-CoA into acetyl-CoA and acetoacetate is actually downstream from HMB as well; however, this diagram makes it look like it isn't due the depiction of 1-way reactions (← or ↓) on the right side and reversible (↔) reactions on the left. In a nutshell, there's 2 interconnected pathways between HMB and HMG-CoA being shown, where different metabolic fates for HMG-CoA are shown on each side of the diagram (bottom left vs bottom right).
The bottom right portion of this image is showing the metabolism of HMG-CoA, which is depicted as being downstream from isovaleryl-CoA; but, you'll notice that the pathway on the left also shows HMB-CoA being converted to both MC-CoA and HMG-CoA, and both of those are also depicted in the pathway on the right side (NB: if you click on the HMG-CoA, MC-CoA, or MG-CoA in this diagram and then view this image in those articles, you'll see 2 highlighted annotations for each of those pairs). Because HMG-CoA is downstream from HMB, the metabolism of HMG-CoA into acetyl-CoA and acetoacetate is actually downstream from HMB as well; however, this diagram makes it look like it isn't due the depiction of 1-way reactions (← or ↓) on the right side and reversible (↔) reactions on the left. In a nutshell, there's 2 interconnected pathways between HMB and HMG-CoA being shown, where different metabolic fates for HMG-CoA are shown on each side of the diagram (bottom left vs bottom right). - So, I guess the two pathways that connect HMB to HMG-CoA in this image should be merged when creating the svg version.
Seppi333 (
Insert 2¢)
10:44, 21 January 2018 (UTC)
- I think we are on the same page now. There is a duplication in the parent diagram which should be eliminated in the daughter biosynthesis and metabolism diagrams. Boghog ( talk) 11:15, 21 January 2018 (UTC)
- Hehe, well I guess this speaks to how complicated the image is right now.
@Boghog: I'm referring to the reactions illustrated above.
|
@ Boghog: I'm creating the SVG diagram right now, but I just noticed something. Since MC-CoA→HMB-CoA requires H2O as an input and both HMB-CoA→HMG-CoA and MC-CoA→MG-CoA require CO2 as an input, wouldn't the MG-CoA→HMG-CoA reaction catalyzed by MG-CoA hydratase also require H2O as an input? If so, I should illustrate that in the diagram. Seppi333 ( Insert 2¢) 10:40, 28 January 2018 (UTC)
- A
hydratase by definition catalyzes the addition of water to a substrate, so yes, I think it should be illustrated in the diagram.
Boghog (
talk)
10:55, 28 January 2018 (UTC)
- File:HMB biosynthesis and metabolism diagram - no labels.svg - that's the outline. Now I need to annotate it. I'll probably upload a few more versions of that pathway outline with slight changes in the line placement while I'm working on the annotations though. Also, I decided to add a pathway for acetoacetate→ β-hydroxybutyrate to the annotated svg diagram. Seppi333 ( Insert 2¢) 11:34, 28 January 2018 (UTC)
@ Boghog: The current version of the annotated SVG diagram is transcluded from the template talk page into the collapse tab below. I've finished re-annotating most of it, but the annotations with an orange background need to be repositioned. Logging off for now. Let me know if you have any feedback. Seppi333 ( Insert 2¢) 13:00, 28 January 2018 (UTC)
- Looks good, but just to reiterate, I still think the diagram needs to be split in two. Also isovaleryl-CoA should be removed from both daughter diagrams since it is not involved in the biosynthesis of HMB and it appears to be a minor side pathway in HMB metabolism.
Boghog (
talk)
16:50, 28 January 2018 (UTC)
- Based upon the current layout, it would be pretty easy to display just the biosynthesis or the metabolism by template-cropping the top/bottom portions of the diagram and annotating over irrelevant lines using non-breaking spaces with a white background color. The full diagram is used in a number of other articles though. Seppi333 ( Insert 2¢) 18:15, 28 January 2018 (UTC)
@
Boghog: I've finished annotating the new svg diagram below. Unless you have any suggestions for further revision, I'm going to replace the current {{
Leucine metabolism in humans}} template with this one. I can work on splitting this diagram into two tomorrow if
Galobtter thinks it would be best to move the biosynthesis section out of the
beta-Hydroxy beta-methylbutyric acid#Pharmacology section.
Seppi333 (
Insert 2¢)
12:36, 29 January 2018 (UTC)
Annotated SVG version of the diagram
| |
|---|---|
|
|
Section reflist
Reflist for
{{
Leucine metabolism in humans}} |
|---|
|
FYI – 4th FAC nomination
Beta-Hydroxy beta-methylbutyric acid recently became a good article, so I renominated it FAC for the 4th time: Wikipedia:Featured article candidates/Beta-Hydroxy beta-methylbutyric acid/archive4.
I really need more editors to comment at this nomination and review it against the WP:Featured article criteria. For those who haven't reviewed an article at FAC before, this is the "FAQ" page for reviewing articles at FAC. Seppi333 ( Insert 2¢) 03:46, 2 February 2018 (UTC)
Question about the term "mTOR"
Figured that folks may be interested in this discussion. Jo-Jo Eumerus ( talk, contributions) 14:29, 4 February 2018 (UTC)
There is a discussion at the above link. Your input is most welcome. Boghog ( talk) 17:39, 11 February 2018 (UTC)
Requesting help with Cooperative antigen transfer
Hello there! I just came across this article ( Cooperative antigen transfer) and I was wondering if an expert could take a look at it? I know little about the topic. -- TheSandDoctor ( talk) 20:20, 12 February 2018 (UTC)
- Looks like WP:OR to me. I have nominated it for deletion. Boghog ( talk) 21:11, 12 February 2018 (UTC)
Please see the latest discussion at Talk:Sex, which proposes to restrict the scope of the article. -- EncycloPetey ( talk) 16:52, 18 February 2018 (UTC)
Umami and Savoriness
Please share your views here: Talk:Umami#This really should be called savory
Anna Frodesiak ( talk) 01:06, 24 February 2018 (UTC)
Facto Post – Issue 10 – 12 March 2018
Facto Post – Issue 10 – 12 March 2018
 Milestone for mix'n'matchAround the time in February when Wikidata clicked past item Q50000000, another milestone was reached: the mix'n'match tool uploaded its 1000th dataset. Concisely defined by its author, Magnus Manske, it works "to match entries in external catalogs to Wikidata". The total number of entries is now well into eight figures, and more are constantly being added: a couple of new catalogs each day is normal. Since the end of 2013, mix'n'match has gradually come to play a significant part in adding statements to Wikidata. Particularly in areas with the flavour of digital humanities, but datasets can of course be about practically anything. There is a catalog on skyscrapers, and two on spiders. These days mix'n'match can be used in numerous modes, from the relaxed gamified click through a catalog looking for matches, with prompts, to the fantastically useful and often demanding search across all catalogs. I'll type that again: you can search 1000+ datasets from the simple box at the top right. The drop-down menu top left offers "creation candidates", Magnus's personal favourite. m:Mix'n'match/Manual for more. For the Wikidatan, a key point is that these matches, however carried out, add statements to Wikidata if, and naturally only if, there is a Wikidata property associated with the catalog. For everyone, however, the hands-on experience of deciding of what is a good match is an education, in a scholarly area, biographical catalogs being particularly fraught. Underpinning recent rapid progress is an open infrastructure for scraping and uploading. Congratulations to Magnus, our data Stakhanovite! Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. If you wish to receive no further issues of Facto Post, please remove your name from
our mailing list. Alternatively, to opt out of all
massmessage mailings, you may add
Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery ( talk) 12:26, 12 March 2018 (UTC)
Hi. I have been thinking over an issue and would appreciate a second opinion.
We have an article about cobalamin, which is another name for vitamin B12. The problem is, as such things sometimes are with vitamins (see vitamin B3 complex/ niacin), that it is a group name for a number of compounds that have a certain structure in common, but depending on the ligand attached they have different names, and slightly different properties, as well as some differences in biological functions. And I was wondering maybe it should be a disambig page?
On the one hand, something indeed can be said about cobalamin as a group, on the other hand it rapidly boils down to "if ligand is this, then it is such and such name with such and such properties". So basically, it has a disambig function.
What do you think, what could be put in that article, if it should remain an article? PubChem has a lot of them for reference (as well as vitamin B12) -- Helixitta ( t.) 14:04, 10 March 2018 (UTC)
- Maybe "Vitamin B12" should discuss the medical/biological/nutritional aspects and "cobalamin" the chemical ones?
Jo-Jo Eumerus (
talk,
contributions)
15:20, 10 March 2018 (UTC)
- I'm not sure. Chemical properties actually lead to biological ones, and usually the article on a substance would have everything in it. Say,
glucose entry has biological function, chemical and physical properties. But I suppose some chemical properties that are common for the group could be added to the article about
cobalamin... --
Helixitta (
t.)
16:23, 10 March 2018 (UTC)
- Similar issues occurred with
vitamin D and
cholecalciferol. The feeling here was that all the drug and therapeutic issues associated with vitamin D deficiency had to be rehashed at the colecalciferol page, and take priority, rather than the chemical and physiological details.
Jrfw51 (
talk)
19:13, 10 March 2018 (UTC)
- Thanks for that example. -- Helixitta ( t.) 19:57, 12 March 2018 (UTC)
- Similar issues occurred with
vitamin D and
cholecalciferol. The feeling here was that all the drug and therapeutic issues associated with vitamin D deficiency had to be rehashed at the colecalciferol page, and take priority, rather than the chemical and physiological details.
Jrfw51 (
talk)
19:13, 10 March 2018 (UTC)
- I'm not sure. Chemical properties actually lead to biological ones, and usually the article on a substance would have everything in it. Say,
glucose entry has biological function, chemical and physical properties. But I suppose some chemical properties that are common for the group could be added to the article about
cobalamin... --
Helixitta (
t.)
16:23, 10 March 2018 (UTC)
Adenosine triphosphate
Adenosine triphosphate, an article that you or your project may be interested in, has been nominated for an individual good article reassessment. If you are interested in the discussion, please participate by adding your comments to the reassessment page. If concerns are not addressed during the review period, the good article status may be removed from the article. AIRcorn (talk) 22:42, 26 March 2018 (UTC)
A thorough review of the gene article
- Transcluded from Talk:Gene/Review
Article review
|
|---|
|
To WP:MCB, WP:GEN, WP:BIOL and WP:EB The gene article gets 50,000 views per month but has been de-listed as a featured article since 2006. Given the success of the recent blitz on the enzyme article, I thought I'd suggest spending a couple of weeks seeing if we can get it up to a higher standard. I'm going to start with updating some of the images. If you'd like to help out on the article, it'd be great to see you there. T.Shafee(Evo﹠Evo) talk 09:49, 31 March 2015 (UTC)
GlossarySnooping around I encountered Template:Genetics glossary, I don't know it's backstory, but it is a rather cleaver idea for a template in my opinion. I partially reckon it might go well under the first image in place or the second image depicting DNA, which conceptually is a tangent. I am not sure, hence my asking. -- Squidonius ( talk) 21:47, 1 April 2015 (UTC)
ReferencesI'm planning on adding some more Molecular Biology of the Cell references to the article using
I've not done much non-standard reference citation so I'll wait until you've done a couple so that I can see the format in context before doing any more. The ones I added yesterday shouldn't be too difficult to reformat.
T.Shafee(Evo﹠Evo)
talk
12:24, 22 April 2015 (UTC)
MBOC referencesArticle Genes [1]: 2 are numerous [1]: 4 and useful [1]: 4.1 References
So {{ rp}} labels the chapter number but does not provide any easy link to the actual information. Therefore it's combined with a list of chapter links. the benefit is that the {{ rp}} template is relatively easy to maintain and the list of chapter links doesn't require maintainance and places all the MBOC links together. As stated above, there's basically no way to avoid linking individually to chapters if we want to cite MBOC. I'll finish building the chapter list over the next couple of days. T.Shafee(Evo﹠Evo) talk 01:29, 27 April 2015 (UTC)
Collapsed the transclusion. Seppi333 ( Insert 2¢) 04:48, 27 March 2018 (UTC) |
Any interest in rescuing this abandoned draft? Espresso Addict ( talk) 01:27, 9 April 2018 (UTC)
Facto Post – Issue 11 – 9 April 2018
Facto Post – Issue 11 – 9 April 2018
 The 100 Skins of the OnionOpen Citations Month, with its eminently guessable hashtag, is upon us. We should be utterly grateful that in the past 12 months, so much data on which papers cite which other papers has been made open, and that Wikidata is playing its part in hosting it as "cites" statements. At the time of writing, there are 15.3M Wikidata items that can do that. Pulling back to look at open access papers in the large, though, there is is less reason for celebration. Access in theory does not yet equate to practical access. A recent LSE IMPACT blogpost puts that issue down to "heterogeneity". A useful euphemism to save us from thinking that the whole concept doesn't fall into the realm of the oxymoron. Some home truths: aggregation is not content management, if it falls short on reusability. The PDF file format is wedded to how humans read documents, not how machines ingest them. The salami-slicer is our friend in the current downloading of open access papers, but for a better metaphor, think about skinning an onion, laboriously, 100 times with diminishing returns. There are of the order of 100 major publisher sites hosting open access papers, and the predominant offer there is still a PDF.  From the discoverability angle, Wikidata's bibliographic resources combined with the SPARQL query are superior in principle, by far, to existing keyword searches run over papers. Open access content should be managed into consistent HTML, something that is currently strenuous. The good news, such as it is, would be that much of it is already in XML. The organisational problem of removing further skins from the onion, with sensible prioritisation, is certainly not insuperable. The CORE group (the bloggers in the LSE posting) has some answers, but actually not all that is needed for the text and data mining purposes they highlight. The long tail, or in other words the onion heart when it has become fiddly beyond patience to skin, does call for a pis aller. But the real knack is to do more between the XML and the heart. Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. If you wish to receive no further issues of Facto Post, please remove your name from
our mailing list. Alternatively, to opt out of all
massmessage mailings, you may add
Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery ( talk) 16:25, 9 April 2018 (UTC)
Move request
Talk:GBA_(disambiguation)#Requested_move_13_April_2018 Jytdog ( talk) 02:59, 15 April 2018 (UTC)
Remove tags for Wikipedia:WikiProject Cell Signaling?
Wikipedia:WikiProject Cell Signaling has been inactive for several years, yet its tag is on the talk pages of 295 pages. Should we remove the Cell signalling banner from those pages and replace it with the MCB banner where appropriate? I figure the purpose of the project tags on article talk pages is (1) to help projects keep track of pages their editors are interested in, and (2) to point readers/editors towards a Project that could help address concerns about a particular article. It seems like having the banner from a defunct WikiProject just serves to confuse. Thoughts? I assume a bot could be found to do this task if folks think it's a good idea. Cheers Ajpolino ( talk) 19:28, 24 April 2018 (UTC)
- At the top of the Wikipedia:WikiProject Cell Signaling page, there is in big bold letters with links: This MCB project subpage is no longer in use and is kept as a historical archive. Please go to the MCB project homepage or talk page for currently active sections. That seems like a sufficient pointer to our active WP. I don't think we need to completely bypass the cell signaling sub-WP. On the other hand, I don't know what is usually done with defunct WP--are they typically bypassed? -- Mark viking ( talk) 20:06, 24 April 2018 (UTC)
Merger discussion for Vitamin B3

An article that you have been involved in editing— Vitamin B3—has been proposed for merging with another article. If you are interested, please participate in the merger discussion. Thank you. SusanLesch ( talk) 13:59, 30 April 2018 (UTC)
Wikiproject Anatomy and the scope of your project
Hi, I am not a member of your project but do most of my work at Wikiproject Anatomy. At the anatomy project we recently finished classifying our articles and we have just under 1500 articles and redirects related to cells and microanatomy. What I would to know is what is under the scope of your project? Do you include individual cell types such as alveolar cells, osteocytes or melanocytes? If such articles are within your scope please let me know and I will gladly start tagging/adding your project banner to their talk page (in the hope that your project members can find and will work on some of the articles of interest to both wikiprojects). Kind regards JakobSteenberg ( talk) 20:47, 29 April 2018 (UTC)
- In my opinion such cell types would also fall within the scope WP:MCB. Some of them are already tagged (e.g. most of the immune cell types). I'll be interested to hear the opinions of others.
T.Shafee(Evo&Evo)
talk
23:56, 29 April 2018 (UTC)
- Thanks for your input. I will wait for others to comment before doing anything. Just be sure;
histology-articles such as
histology of the vocal folds,
Loop of Henle or
Portal triad are not within the scope of MCB correct?
JakobSteenberg (
talk)
17:55, 30 April 2018 (UTC)
- I agree with Evo. With respect to morphology and cell type, the two projects clearly overlap. Boghog ( talk) 18:38, 30 April 2018 (UTC)
- Thanks for your input. I will wait for others to comment before doing anything. Just be sure;
histology-articles such as
histology of the vocal folds,
Loop of Henle or
Portal triad are not within the scope of MCB correct?
JakobSteenberg (
talk)
17:55, 30 April 2018 (UTC)
LCHN: An Error has occurred retrieving Wikidata item for infobox
I just moved Draft:LCHN into mainspace. @ M.Gray97: The infobox_gene template is generating An Error has occurred retrieving Wikidata item for infobox. My template-fu is insufficient to fix this. Could a SME take a look? Thanks. -- RoySmith (talk) 21:17, 6 May 2018 (UTC)
Draft:Comparison_of_DNA_melting_prediction_software
Draft:Comparison_of_DNA_melting_prediction_software I'm a Afc reviewer with cell biology background but still struggling whether this bioinformatics page can be accepted . do give me an opinion. thanks much Quek157 ( talk) 16:22, 8 May 2018 (UTC)
3D protein/nucleic acid structures could be rotateable 3D models: NGL
Hey everyone. I'm no wiki expert and there might be internal reasons not to do this, but I thought I'd float it.
There is an excellent+beautiful+open source molecular graphics thing called NGL. It is being developed at UCSD as part of the protein data bank, and I know the person doing it. Some examples: Ribosome https://www.rcsb.org/3d-view/4V4J RNA pseudoknot http://www.rcsb.org/3d-view/2LC8 Dorothy Hodgkin's Insulin https://www.rcsb.org/3d-view/4INS
You can see a (super cool+interactive) presentation on it here http://nglviewer.org/talks/ngl-3dsig/
Adopting ngl would be great for wiki because: -Everything would look super beautiful and make the molecular biology pages more readable -It would allow you to have images of any molecule for which we have a model (= any molecule for which you would want to see an "image") on any page, because you wouldn't need to find a public domain picture. In principle you could just have some markup like <ngl window: "pdbID:4ins"> or whatever and suddenly your webpage has an excellent up-to-date structure
NGL is very cross platform and works well on smartphones. However, it is obviously dependent on webGL.
Using NGL would take more bandwidth. For each structure you'd need to download a pdb file, plus the ngl javascript (which you'd have to do for every page?). That said, I'd like to point out that a 500KB pdb file is smaller than a 1MB animated gif of the same molecule performing a rotation, so a small number of pages would be compressed by this adoption.
NGL is better than the visualizer that proteopedia use http://proteopedia.org/wiki/index.php/Main_Page as you can verify for yourself. — Preceding unsigned comment added by Hamishtodd1 ( talk • contribs) 12:06, 10 May 2018 (UTC)
- The proteopedia visualizer I use is FirstGlance ... see http://proteopedia.org/wiki/index.php/FirstGlance_in_Jmol and https://bioinformatics.org/firstglance/fgij/. - Dank ( push to talk) 13:56, 10 May 2018 (UTC)
Discussion at Wikipedia talk:WikiProject Medicine#MicroDNA
![]() You are invited to join the discussion at
Wikipedia talk:WikiProject Medicine#MicroDNA. –
Finnusertop (
talk ⋅
contribs)
22:53, 10 May 2018 (UTC)
You are invited to join the discussion at
Wikipedia talk:WikiProject Medicine#MicroDNA. –
Finnusertop (
talk ⋅
contribs)
22:53, 10 May 2018 (UTC)
Facto Post – Issue 12 – 28 May 2018
Facto Post – Issue 12 – 28 May 2018
 ScienceSource fundedThe Wikimedia Foundation announced full funding of the ScienceSource grant proposal from ContentMine on May 18. See the ScienceSource Twitter announcement and 60 second video.
The proposal includes downloading 30,000 open access papers, aiming (roughly speaking) to create a baseline for medical referencing on Wikipedia. It leaves open the question of how these are to be chosen. The basic criteria of WP:MEDRS include a concentration on secondary literature. Attention has to be given to the long tail of diseases that receive less current research. The MEDRS guideline supposes that edge cases will have to be handled, and the premature exclusion of publications that would be in those marginal positions would reduce the value of the collection. Prophylaxis misses the point that gate-keeping will be done by an algorithm. Two well-known but rather different areas where such considerations apply are tropical diseases and alternative medicine. There are also a number of potential downloading troubles, and these were mentioned in Issue 11. There is likely to be a gap, even with the guideline, between conditions taken to be necessary but not sufficient, and conditions sufficient but not necessary, for candidate papers to be included. With around 10,000 recognised medical conditions in standard lists, being comprehensive is demanding. With all of these aspects of the task, ScienceSource will seek community help. Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. ScienceSource pages will be announced there, and in this mass message. If you wish to receive no further issues of Facto Post, please remove your name from
our mailing list. Alternatively, to opt out of all
massmessage mailings, you may add
Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery ( talk) 10:16, 28 May 2018 (UTC)
WikiProject collaboration notice from the Portals WikiProject
The reason I am contacting you is because there are one or more portals that fall under this subject, and the Portals WikiProject is currently undertaking a major drive to automate portals that may affect them.
Portals are being redesigned.
The new design features are being applied to existing portals.
At present, we are gearing up for a maintenance pass of portals in which the introduction section will be upgraded to no longer need a subpage. In place of static copied and pasted excerpts will be self-updating excerpts displayed through selective transclusion, using the template {{ Transclude lead excerpt}}.
The discussion about this can be found here.
Maintainers of specific portals are encouraged to sign up as project members here, noting the portals they maintain, so that those portals are skipped by the maintenance pass. Currently, we are interested in upgrading neglected and abandoned portals. There will be opportunity for maintained portals to opt-in later, or the portal maintainers can handle upgrading (the portals they maintain) personally at any time.
Background
On April 8th, 2018, an RfC ("Request for comment") proposal was made to eliminate all portals and the portal namespace. On April 17th, the Portals WikiProject was rebooted to handle the revitalization of the portal system. On May 12th, the RfC was closed with the result to keep portals, by a margin of about 2 to 1 in favor of keeping portals.
Since the reboot, the Portals WikiProject has been busy building tools and components to upgrade portals.
So far, 84 editors have joined.
If you would like to keep abreast of what is happening with portals, see the newsletter archive.
If you have any questions about what is happening with portals or the Portals WikiProject, please post them on the WikiProject's talk page.
Thank you. — The Transhumanist 10:59, 31 May 2018 (UTC)
Is this worth salvaging? (Delete vs. move to mainspace vs. something else?) Calliopejen1 ( talk) 02:22, 4 June 2018 (UTC)
- It probably should be merged into Insulin#Regulation. The sources need to be moved in-line and it needs some copyediting first. I will work on this when I find time. Boghog ( talk) 03:48, 4 June 2018 (UTC)
- Not needed as a standalone article when it fits nicely within Insulin. Natureium ( talk) 14:14, 4 June 2018 (UTC)
Large copy-paste at Lysine
A large paste of text from WikiJournal of Science/Lysine: biosynthesis, catabolism and roles has been dropped into the article on Lysine. In addition to a number of references breaking in the process, there are issues of overly technical content and copy-paste, even if copyright issues do not apply. I'm too busy to dig into the issues more deeply for at least a few days, so I'm going to punt it over here for now. Sorry and thanks, BiologicalMe ( talk) 03:39, 10 June 2018 (UTC)
- Fixed the link. Adrian J. Hunter( talk• contribs) 05:01, 10 June 2018 (UTC)
-
BiologicalMe it looks like
Evolution and evolvability was in the middle of working on this when you posted here, and has edited extensively since. Not sure there's any remaining issues?
Adrian J. Hunter(
talk•
contribs)
05:01, 10 June 2018 (UTC)
- Hello, thanks for flagging this here
BiologicalMe. I've been in the process of integrating an updated version of the article that has been written at the location you mentioned. It's under a CC-BY license, so the copyright isn't a specific issue (links to the relevant sections of the journal's
editorial and
ethics guidelines). You're correct that the ref errors were introduced during the process which I was fixing as i went on (also thanks to boghog for their ever-present assistance!). The main thing of importance is
WP:TECHNICAL. I believe that many of the updates have largely made the article less technical (especially the lead;
prev version), however I fully support improving readability and accessibility where possible. I also removed mentions of lysine deficiency causing blindness for now, since it was unreferenced. As well as this specific case, feedback from Wikipedians on the journals' processes is very welcome (see also
list of Wikipedia articles updated based on academic journal content). We try to operate WikiJournals in a transparent, open and accountable format, so there is a general discussion board for the WikiJournal User Group
here, as well as discussion boards or each journal.
T.Shafee(Evo&Evo)
talk
05:32, 10 June 2018 (UTC)
- Thanks
User:Evolution and evolvability. I knew I was too tired to address it coherently, and I knew I would have no free time today to figure out copyleft licence compatibility or scramble through the references. As an indication of how tired I was, I forgot to provide a diff or to tag you. I'm looking forward to reading through what you've done, fully alert. Hopefully tomorrow.
BiologicalMe (
talk)
15:27, 10 June 2018 (UTC)
- No worries! You were absolutely right to post here. T.Shafee(Evo&Evo) talk 00:39, 11 June 2018 (UTC)
- Thanks
User:Evolution and evolvability. I knew I was too tired to address it coherently, and I knew I would have no free time today to figure out copyleft licence compatibility or scramble through the references. As an indication of how tired I was, I forgot to provide a diff or to tag you. I'm looking forward to reading through what you've done, fully alert. Hopefully tomorrow.
BiologicalMe (
talk)
15:27, 10 June 2018 (UTC)
- Hello, thanks for flagging this here
BiologicalMe. I've been in the process of integrating an updated version of the article that has been written at the location you mentioned. It's under a CC-BY license, so the copyright isn't a specific issue (links to the relevant sections of the journal's
editorial and
ethics guidelines). You're correct that the ref errors were introduced during the process which I was fixing as i went on (also thanks to boghog for their ever-present assistance!). The main thing of importance is
WP:TECHNICAL. I believe that many of the updates have largely made the article less technical (especially the lead;
prev version), however I fully support improving readability and accessibility where possible. I also removed mentions of lysine deficiency causing blindness for now, since it was unreferenced. As well as this specific case, feedback from Wikipedians on the journals' processes is very welcome (see also
list of Wikipedia articles updated based on academic journal content). We try to operate WikiJournals in a transparent, open and accountable format, so there is a general discussion board for the WikiJournal User Group
here, as well as discussion boards or each journal.
T.Shafee(Evo&Evo)
talk
05:32, 10 June 2018 (UTC)
Similar Pages?
Note from OTRS ticket 2018062010005412 - the following pages appear to duplicate the same subject
Ronhjones (Talk) 22:27, 24 June 2018 (UTC)
- I think they're sufficiently distinct to warrant separate pages.
C/EBP is a
family of proteins, which includes
CEBPA,
CEBPB,
CEBPG etc.
T.Shafee(Evo&Evo)
talk
22:52, 24 June 2018 (UTC)
- Yeah, not duplicates. There is more than one C/EBP protein.
Jo-Jo Eumerus (
talk,
contributions)
12:46, 25 June 2018 (UTC)
- Thanks, not my field - mine is Chemistry. I'll let the OP know Ronhjones (Talk) 16:18, 25 June 2018 (UTC)
- Yeah, not duplicates. There is more than one C/EBP protein.
Jo-Jo Eumerus (
talk,
contributions)
12:46, 25 June 2018 (UTC)
We have separate articles on Galactoside O-acetyltransferase and Beta-galactoside transacetylase, but are they really different enzymes? The first two sources used at Beta-galactoside transacetylase seem to classify them as synonymous: galactoside acetyltransferase (thiogalactoside transacetylase) (thiogalactoside transacetylase, LacA, GAT). Vaselineeeeeeee ★★★ 22:31, 22 June 2018 (UTC)
- @ Vaselineeeeeeee: I'm pretty sure you're correct. I think the two names are synonyms fort teh same anzyem (just slightly different description s of its catalytic activity). "Galactoside acetyltransferase" seems to be the most common variant of the name in the literature. T.Shafee(Evo&Evo) talk 00:39, 26 June 2018 (UTC)
- Additional consideration:
Galactoside O-acetyltransferase is the formal
Enzyme Commission number naming but the least-commonly used in publications!
T.Shafee(Evo&Evo)
talk
00:51, 26 June 2018 (UTC)
- @ Evolution and evolvability: Yeah I noticed that too. You think we should merge to the Galactoside O-acetyltransferase page and possibly get rid of the O- for common name? Vaselineeeeeeee ★★★ 02:22, 26 June 2018 (UTC)
Human Genome had a whole section of creationist claptrap for over three years
[1]
[2]
![]() Adrian J. Hunter(
talk•
contribs)
12:40, 25 June 2018 (UTC)
Adrian J. Hunter(
talk•
contribs)
12:40, 25 June 2018 (UTC)
- Ouch! Thanks for spotting and fixing that. Boghog ( talk) 16:34, 25 June 2018 (UTC)
- Seconded. Well done.
T.Shafee(Evo&Evo)
talk
00:01, 26 June 2018 (UTC)
- Ah, sorry my post was misleading. The problem was spotted and fixed in the first ever edit from an IP, in the second diff linked above. Adrian J. Hunter( talk• contribs) 08:53, 26 June 2018 (UTC)
Facto Post – Issue 13 – 29 May 2018
Issue 13 – 29 May 2018
| |
|---|---|
MediaWiki message delivery ( talk) 18:19, 29 June 2018 (UTC) |
Discussion on readability of some medical / biology articles and their infoboxes at WT:PHARM
See Wikipedia_talk:WikiProject_Pharmacology#Readability_of_medical_physiology_articles. -- Tom (LT) ( talk) 18:27, 3 July 2018 (UTC)
Readability of medical physiology articles
There is a discussion concerning readability of {{ Infobox gene}} at the above link that is well within the scope this project. In fact, the discussion probably been started here, rather than there. In any case, your input is welcome. Boghog ( talk) 20:06, 5 July 2018 (UTC)
Cellosaurus
There is a notability/ COI discussion over at Talk:Cellosaurus. Both this discussion and the article itself would benefit from input from WP:MCB members. -- Daniel Mietchen ( talk) 15:49, 10 July 2018 (UTC)
See Wikipedia:Featured article review/Antioxidant/archive1. Seppi333 ( Insert 2¢) 22:34, 10 July 2018 (UTC)
Facto Post – Issue 14 – 21 July 2018
Issue 14 – 21 July 2018
| |
|---|---|
MediaWiki message delivery ( talk) 06:10, 21 July 2018 (UTC) |
Improvements to infobox gene
I can see a great deal of effort has gone to producing infobox gene and it clearly has some amazing wikidata integration. I think it could be improved in a number of ways and propose some specific suggestions here for discussion. My aim is to make the infobox shorter, easier to read, and increase the prominence of basic information. (I would normally make a sandbox of my proposed edits but am unable to because the infobox is written in lua.) For my inspiration, I am viewing BRCA1 and MLH1.
I'm starting this part of the discussion back here (rather than WT:PHARM), as boghog points out, this being a more appropriate venue. Ping to editors that have participated: Boghog, Seppi333, Jytdog, Looie496, Ajpolino, Tryptofish. Apologies if I've left anyone out. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- + Andrew Su, Daniel Mietchen and Opabinia regalis who may have useful ideas. T.Shafee(Evo&Evo) talk 12:05, 16 July 2018 (UTC)
1. Move some data to an 'authority control' template
We're moving in that direction globally on WP. The list of 'external IDs' in the infobox could easily be moved to an authority control template for articles. This would also help shorten the infobox, reduce the length of its code, and more importantly, simplify the infobox, which is my goal here.-- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support - I find many of the links useful, but I'm fairly confident the average reader does not. The minority of our readers who follow these links (probably biologists) can find them just as well at the bottom of the page. This has been done with taxonomic identifiers for many organism articles as well. Ajpolino ( talk) 20:09, 14 July 2018 (UTC)
- Strong oppose – As long as these links are toward the bottom of the infobox, I do not see the need for this. Many of the articles that transcribe this infobox are somewhat obscure and not often read by the average reader. It is more convenient for the specialist to have all the relevant links in the same infobox. Also most of the links in question are not authority control. They are links to specialist databases that provide more detailed information about the gene/protein. Boghog ( talk) 22:16, 14 July 2018 (UTC)
- Oppose I think collapsing this to avoid cluttering the box with things that won't interest many people without moving it to the bottom of the page is the best solution. Natureium ( talk) 20:09, 16 July 2018 (UTC)
2. Remove pictures of chromosomes

I refer here to the full chromosome panel, not the horizontal individual chromosome with the gene's location boxed. These pictures take up a lot of space, add to loading times, and don't contribute anything factual. They are also duplicated in text (twice) in the infobox.-- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Collapse would be a better option. I agree that this figure is mainly eye candy. Boghog ( talk) 22:17, 14 July 2018 (UTC)
- Collapse, with removal as a secondary option. The diagram would be able to justify itself more if the exact chromosomal location was pointed to.
T.Shafee(Evo&Evo)
talk
11:48, 16 July 2018 (UTC)
- Agree. If the exact location were here that'd be great. As it is, our description of "chromosome 9" is backed up by a numbered group of chromosomes and the ninth selected. I just don't see how this is useful? -- Tom (LT) ( talk) 10:06, 19 July 2018 (UTC)
- Oppose I think this is a nice visual for people who aren't very familiar with genetics but may be looking for information for personal health purposes.
Natureium (
talk)
20:09, 16 July 2018 (UTC)
- But what person not familiar with genetics but using this for their personal health will have access to a karyotype, and thereby find comparing this with their karyotype useful...?
3. Collapse orthologs section by default
The section takes up a large amount of vertical space and is limited to links. I propose collapsing it by default. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Oppose – I find those links very useful. Furthermore they are at the bottom of the infobox so they should not obscure more general information at the top of the box. What could be done to reduce it size is to display only one Refseq link like the previous {{ GNF Protein box}} did. Boghog ( talk) 22:23, 14 July 2018 (UTC)
- I'd support either collapsing the section by default (still leaves it accessible to those interested) or reducing the display to one Refseq link (cuts down on the external link overload in the infobox). I'd also support doing both of those together. Ajpolino ( talk) 14:45, 15 July 2018 (UTC)
- Weakly Support This would save space, but I don't think it's much of an improvement. Natureium ( talk) 20:09, 16 July 2018 (UTC)
4. Rename 'available structures' section
There is a title that is called 'available structures' but it seems to be just some links. I don't know what it does or how it relates to genes, and therefore presume most lay readers won't either. I propose renaming it to something that explains what it does, or merging it to identifiers section if it is related. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment – The available structures section refers to 3D structures of the protein stored in the
Protein Data Bank and are quite distinct from identifiers. Function derives from structure hence those links are essential. I use them all the time.
Boghog (
talk)
22:29, 14 July 2018 (UTC)
- Ah. @
Boghog then these would be suitable to move to an external links section of the infobox? --
Tom (LT) (
talk)
22:47, 14 July 2018 (UTC)
- The left-hand side (green shaded) are explanatory internal links. Almost everything in the right-hand side (grey-shaded) is externally linked. It makes no sense to move only the Protein Databank External links to an external link section when almost everything else in the infobox is also an external link. Boghog ( talk) 06:49, 15 July 2018 (UTC)
- The rationale for placing the PBD links right below the infobox graphic is that the graphics are most commonly of the protein structure derived from one the PDB links. There are often more than one available protein structure (for example structures of different domains of the protein, complexes with different ligands or with other proteins, etc.). Having all the PDB links in one location right below the graphic of the structure gives a more complete view of the structure. Boghog ( talk) 07:54, 15 July 2018 (UTC)
- Ah. @
Boghog then these would be suitable to move to an external links section of the infobox? --
Tom (LT) (
talk)
22:47, 14 July 2018 (UTC)
- Support - Rename to something like "Protein structure" or equivalent (i.e. not gene structure, or chromosomal structure). The "Available" is unnecessary (if there were none available, the section wouldn't be shown. Perhaps
{{ Infobox protein}}could be merged in? T.Shafee(Evo&Evo) talk 11:53, 16 July 2018 (UTC)- This option makes clear the point of the section and, as Evolution and evolvability points out, the 'available' is superfluous - so has my full support. -- Tom (LT) ( talk) 10:03, 19 July 2018 (UTC)
- Weakly oppose This is an explanatory enough title and no one has proposed a better alternative. Natureium ( talk) 20:09, 16 July 2018 (UTC)
5. Create a 'basic information' section and put it at the top of the infobox.
A 'basic information' section should be created and placed beneath the picture of the protein ribbon and include basic information about the gene written in a human-readable format. Fields can be discussed once we have some support for this section. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support - There was substantial discussion at Wikipedia_talk:WikiProject_Pharmacology#Readability_of_medical_physiology_articles regarding what fields might constitute basic information and where database entries to populate these fields could be found. I agree with points raised by Boghog and Seppi333 at that earlier discussion that to be both reliable and human-readable, this may have to be curated (or completely written) manually. I'm happy to help with this. Ajpolino ( talk) 20:09, 14 July 2018 (UTC)
- Support – Giving a birds eye view of the function, mechanism, and location of the protein is a very worthwhile goal. This can be done by manually selecting the most important GO terms for display while leaving the complete list in a collapsed state. Boghog ( talk) 22:32, 14 July 2018 (UTC)
- support per Boghog rationale-- Ozzie10aaaa ( talk) 10:11, 16 July 2018 (UTC)
- oppose and strongly this is trying to solve a content problem with an infobox which is wrong headed -- as wrong headed as the WMF trying to solve our cluttered leads by grabbing the "description" field from Wikidata and forcing it onto WP articles on mobile and in apps. (which in turn has generated a ton of more work for us, since the "solution" was creating a new
{{ short description}}field for every article in WP.) Now a second field of "summary information" is being proposed.. and this is just crazy to me. But I guess people will volunteer doing what they want.
- What bugs me, is that both of these things have nothing to do with fixing the bad leads that are the source of the problems. If the leads of these articles were doing their job this would not be needed. Most importantly the time it would take to do this should really be put into actually fixing the leads. But again, people will spend their time here doing as they will....
Jytdog (
talk)
16:39, 16 July 2018 (UTC)
- I don't really see what the relevance of this infobox discussion is w.r.t. article leads. The infobox summarizes body content (and even content not covered in the body) in a succinct tabular format while also providing relevant external links. A lead and an infobox serve complementary roles in the sense that one conveys information in prose and the other in a data table. Each has its own advantages over the other; hence, the use of both is ideal. Fixing article leads is another issue altogether. Seppi333 ( Insert 2¢) 03:18, 17 July 2018 (UTC)
- The goal of the
Gene Wiki is to create a Wikipedia article for every notable human gene starting with an infobox and public domain Entrez description of the gene/protein. The intention was this seed material would encourage editors in expanding these articles. Unfortunately majority of these articles are still stubs. In addition, many of the Entrez descriptions are either incomplete or missing all together. Adding a "key properties" section to the infobox would go a long way to provide a minimum level of information for each stub. For more complete articles, the "key properties" section may highlight basic information that is lacking in the lead and encourage editors to add that information. Finally it should be pointed out that there are currently over 11,000
Gene Wiki articles. Expanding and cleaning up the leads for all of these articles would be an enormous task and there are simply not enough editors to do this in a reasonable time frame. Adding a manually curated "key properties" section is also a lot of work, but could be accomplished much more quickly. After the "key properties" sections have been added, we will of course continue to work on the leads, but that work will take much longer.
Boghog (
talk)
13:24, 17 July 2018 (UTC)
- The OP
started by saying
I often use WP as revision and for my work as I'm sure many people do. I find many of our articles (e.g. Beta-2 adrenergic receptor, which I'm looking at now) very difficult to read and lacking key information which (I feel at least) could be included in the infobox - e.g. where is that thing, what does that thing do, what makes that thing).
. The problem they point out is that articles don't have basic information, and their solution was "infobox". This is the wrong solution in my view. But again people will put time in where they want. This just seems to be a diversion of our most valuable resource to me. - Additionally,
User:Boghog I am curious why you say it would take less time and effort to add this content manually to an infobox compared to an article. Would you please explain?
Jytdog (
talk)
19:16, 17 July 2018 (UTC)
- As I have stated before, the required GO data is already stored in wikidata. We just need to manually select a subset to be highlighted. In addition, because of within family similarities of the data, processing the data one family at a time will further speed up the process. The entire process can be automatic with a manually assisted script. It is trivial to display this type of data in an infobox. It is much more difficult to automate the integration of this data into existing prose. And regardless of whether we have a understandable lead or not, having the key properties listed in the infobox will be useful in its own right.
Boghog (
talk)
21:34, 17 July 2018 (UTC)
- Oh, you didn't mean manual at all. This is much more worrisome. I would be curious to see a wikidata entry with the kind of data that you believe would be appropriate for this "basic information" field. Can you point me to one?
Jytdog (
talk)
20:07, 18 July 2018 (UTC)
- An example of the kind of manually selected data that I propose to include in the gene infobox is found in the "Key Properties" table
below. The manual part is an editor making a decision about which of the existing GO terms should be promoted and this will take a considerable amount of work. The proposed work flow is as follows:
- down load the GO data from wikidata into a spread sheet (automated)
- organize by family (automated)
- select preferred data (manual)
- adjust the wikidata ranking of the selected data (automated)
- Please note that each of the GO terms provides one or more sources and none of the GO terms is a medical claim. Many of the claims are uncontroversial and can quickly be processed (e.g., a GPCR is membrane associated and the list of endogenous ligands for a particular GPCR). Identifying the primary function may take a little more work. Boghog ( talk) 20:44, 18 July 2018 (UTC)
- Also please note that the data already is in the infobox, but in a collapsed state. The proposal is to manually select a subset for uncollapsed display.
Boghog (
talk)
04:51, 19 July 2018 (UTC)
- Hey Boghog sorry taking time to respond.
Here is the Wikidata entry on BRCA1. Where in there is the "basic information" about "where is that thing, what does that thing do, what makes that thing" in that wikidata entry? (I acknowledge tissue expression data is there in the png image, as it is the infobox)
Jytdog (
talk)
14:28, 21 July 2018 (UTC)
- Fair enough question. The relevant wikidata is here: BRCA1 DNA repair associated (Q17487737) BRCA1 is an "edge case" in that it is a multifunctional protein that has intensively studied and hence many GO terms have been assigned to it. But hey, that is biology. The "where is that thing" question has two parts. In which tissues is it expressed and which part of the cell is it located. The short answer is that it is expressed everywhere (except plasma). That is reflected in the "cellular component" GO terms: plasma membrane, cytoplasm, and cell nucleus. The GO terms generally don't distinguish between tissues. The RNA expression pattern suggests wide tissue distribution and that is also seen in BRAC1 protein expression ( human protein atlas). "What does that thing do" is function. "Biological process" GO lists "DNA repair" which I think qualifies as it primary function. As I have mentioned elsewhere "what makes that thing" is somewhat of a generic question. The short answer is where all proteins are made, ribosomes. Digging deeper, what differentiates proteins synthesis is post-translational modification and GO terms do not cover this. So in summary, the promoted GO terms would look like:
- Hey Boghog sorry taking time to respond.
Here is the Wikidata entry on BRCA1. Where in there is the "basic information" about "where is that thing, what does that thing do, what makes that thing" in that wikidata entry? (I acknowledge tissue expression data is there in the png image, as it is the infobox)
Jytdog (
talk)
14:28, 21 July 2018 (UTC)
- An example of the kind of manually selected data that I propose to include in the gene infobox is found in the "Key Properties" table
below. The manual part is an editor making a decision about which of the existing GO terms should be promoted and this will take a considerable amount of work. The proposed work flow is as follows:
- Oh, you didn't mean manual at all. This is much more worrisome. I would be curious to see a wikidata entry with the kind of data that you believe would be appropriate for this "basic information" field. Can you point me to one?
Jytdog (
talk)
20:07, 18 July 2018 (UTC)
- As I have stated before, the required GO data is already stored in wikidata. We just need to manually select a subset to be highlighted. In addition, because of within family similarities of the data, processing the data one family at a time will further speed up the process. The entire process can be automatic with a manually assisted script. It is trivial to display this type of data in an infobox. It is much more difficult to automate the integration of this data into existing prose. And regardless of whether we have a understandable lead or not, having the key properties listed in the infobox will be useful in its own right.
Boghog (
talk)
21:34, 17 July 2018 (UTC)
- The OP
started by saying
Key Properties Property Description Molecular mechanism damaged DNA binding Biological function DNA repair Subcellular location plasma membrane, cytoplasm, cell nucleus
- Support with caveats that this be very limited and written manually. Natureium ( talk) 20:09, 16 July 2018 (UTC)
6. Rename 'orthologs' to 'external links'
The orthologs part of this section is in fact just the titles 'human' and 'mouse'. The rest are just external links. The proposed change makes the purpose of this section much clearer to readers and reflects what it actually does.
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment– almost everything in the right-hand side of the infobox (grey shaded areas) is an external link so it makes no sense creating a separate external links section. The orthologs listed (human and mouse) are just example organisms, although very important ones. The present table layout makes sense (the left-hand side contains internal wikilinks that provide and explanation for the right-hand side external links). This layout is also very efficient as the headings need only be displayed only once.
Boghog (
talk)
08:53, 15 July 2018 (UTC)
- If as you say these are just external links, why does the section need to be a title that 99% of readers will not be able to understand? --
Tom (LT) (
talk)
11:29, 15 July 2018 (UTC)
- The whole infobox, not just the orthologs section, is filled with external links. Perhaps "orthologs" could be replaced with "representative species" (or "critters" ;-).
Boghog (
talk)
12:11, 15 July 2018 (UTC)
- @ Boghog Whilst I prefer "critters" as it is both highly technically accurate and also understandable by laypeople, I would also support "representative species" with a piped wikilink (like representative species).-- Tom (LT) ( talk) 09:59, 19 July 2018 (UTC)
- The whole infobox, not just the orthologs section, is filled with external links. Perhaps "orthologs" could be replaced with "representative species" (or "critters" ;-).
Boghog (
talk)
12:11, 15 July 2018 (UTC)
- If as you say these are just external links, why does the section need to be a title that 99% of readers will not be able to understand? --
Tom (LT) (
talk)
11:29, 15 July 2018 (UTC)
- Not to 'external links' Another name might be more readily understood, but there are external links throughout most of the infobox. Orthologs seems to be the most technically correct, and "critters" is just ridiculous.
Natureium (
talk)
20:09, 16 July 2018 (UTC)
- Of course "critters" is ridiculous. Turn on your humor detector. Boghog ( talk) 04:55, 19 July 2018 (UTC)
7. Merge gene location human/mouse together
Merging these two parts together reduces the amount of headings in the infobox (which are many) and hopefully also simplifies its display. The section can include a subheading for human and mouse. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Oppose – Displaying the human and mouse links side-by-side saves space since the headings (Entrez, Ensembl, UniProt, etc.) only need to be displayed once. Putting one on top of the other would require duplicating the headings.
Boghog (
talk)
22:51, 14 July 2018 (UTC)
- @ Boghog this proposal relates to merging the two sections of the infobox titled "gene location (human)" and "gene location (mouse)" into a single section titled "gene location". This proposal does not relate to the orthologs section. -- Tom (LT) ( talk) 11:28, 15 July 2018 (UTC)
- OK, sorry I misunderstood your proposal. For this merge to work, I think the graphics would need to be removed and the merged section would consist of a three column data similar to the orthlog section. We then might as well merge the gene location data to the orthlog section which already contains a partially redundant "Location (UCSC)" row. Boghog ( talk) 11:59, 15 July 2018 (UTC)
- Oppose The human chromosomal information will be relevant to many more people, and it makes sense to me to have the human section uncollapsed and the mouse section collapsed by default. Natureium ( talk) 20:09, 16 July 2018 (UTC)
8. Display only a few things in 'gene ontology', or go back to plain text
It seems like we're on the way to doing this based on WT:PHARM, but I will propose it here as a plan B. The 'gene ontology' section is not useful as is. It contains 20 - 30 text entries describing processes. None of these reflect the article's lead description of the gene, nor the commonly understood function of a gene, e.g. when it is described as encoding an enzyme, membrane receptor, tumour suppressor gene etc. I propose that this section, if it can't be made easier to read and have a more narrow focus, be removed and we just handwrite everything ourselves. Yes that takes time but as an encyclopedia we are never finished and we should be aiming for the best - and spaghetti lists are not helpful to anyone. These lists are not useful at the moment - experts will already know these facts, and lay readers will not be able to make head nor tail of it, nor know how to extract the important items in the list from the chaff. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment – Actually I think leaving this list in place in a collapsed state servers a useful propose. This way editors can easily compare the full list against the promoted list and adjust the promoted list if they don't agree with our selections. In addition, there may be "minor functions" that a individual reader might find interesting. Boghog ( talk) 22:41, 14 July 2018 (UTC)
9. Rename 'gene ontology' to 'gene function'
Renaming ontology to function makes the purpose of the section much clearer to end readers, which should be our goal here. I am not sure that readers will even look at the content of the section because they won't know what the section title means. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
Support–More understandable to the average reader. Boghog ( talk) 22:43, 14 July 2018 (UTC) On second thought, this would be misleading. The gene ontology contains more than function data, it also contain mechanism and location. In any case, if we introduced a "keys fact section" above the "gene ontology" section, the need to rename "gene ontology" becomes less urgent. Boghog ( talk) 07:15, 15 July 2018 (UTC) Labeling the section "protein properties" might work. Boghog ( talk) 10:14, 15 July 2018 (UTC)- I'd support "protein properties", which is satisfactorily descriptive, or almost any title that doesn't contain ontology, which readers will not understand. -- Tom (LT) ( talk) 11:30, 15 July 2018 (UTC)
- Oppose gene function. Ambivalent on protein properties. Natureium ( talk) 20:09, 16 July 2018 (UTC)
10. Merge the two gene location fields
Gene location is provided in three sections: gene location (human), and (mouse), and also as a field in orthologs. I propose there that the field 'Location (UCSC)' present in orthologs be moved to the appropriate gene location (human) and (mouse) sections, thus centralising this information and also reducing some duplication. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment – Unless I am missing something, this proposal is essentially a duplicate of #7 above.
Boghog (
talk)
07:46, 15 July 2018 (UTC)
- Not the same proposal. Within the section called orthologs there is a field called "location (USBC)". IN addition there are two sections called "gene location". The proposal here is to remove the "location (UCSC)" field from orthologs, and move the contents to the sections titled "gene location". --
Tom (LT) (
talk)
11:27, 15 July 2018 (UTC)
- OK, now I undertand. As mentioned above, the merge could be done in the reverse direction (i.e., merge the "gene location" sections into the orthologs section).
Boghog (
talk)
12:04, 15 July 2018 (UTC)
- Boghog I'd support either way, and your proposed solution also has the benefit of removing some top level infobox headings :). -- Tom (LT) ( talk) 10:01, 19 July 2018 (UTC)
- OK, now I undertand. As mentioned above, the merge could be done in the reverse direction (i.e., merge the "gene location" sections into the orthologs section).
Boghog (
talk)
12:04, 15 July 2018 (UTC)
- Not the same proposal. Within the section called orthologs there is a field called "location (USBC)". IN addition there are two sections called "gene location". The proposal here is to remove the "location (UCSC)" field from orthologs, and move the contents to the sections titled "gene location". --
Tom (LT) (
talk)
11:27, 15 July 2018 (UTC)
11. Conservative alternative
Rather than making a large number of changes to the infobox all at once, I think a more limited number of changes could make the infobox much more readable for a general audience without sacrificing the usefulness for specialists. Hence I propose that we implement the following subset of what has been proposed above:
- Collapse Gene location (proposal 2)
- Collapse RNA expression pattern (new proposal)
- Introduce a new "key properties" section near the top (proposal 5). Using the Gene Ontology section in Beta-2 adrenergic receptor as an example, if we were to rank a subset of GO data as "preferred" for the wikidata Adrenoceptor beta 2 (Q287961) data set (see diff), the following could be displayed in the infobox:
| Property | Description |
|---|---|
| Molecular mechanism | G-protein coupled receptor activity, epinephrine binding, norepinephrine binding |
| Biological function | regulation of smooth muscle contraction |
| Subcellular location | membrane |
Of course to implement any of this, we must first get buy-in from the those that maintain the infobox and associated wikidata. For example, we have to make sure that changes to the rank of data do not get overwritten by bots. Boghog ( talk) 10:20, 15 July 2018 (UTC)
- Support as proposer. Boghog ( talk) 07:10, 15 July 2018 (UTC)
- Support would support the batch of proposals above in addition to any others suggested. I've tried to propose some piecemeal changes rather than batch them so that they may be discussed clearly and separately. -- Tom (LT) ( talk) 11:26, 15 July 2018 (UTC)
- Support, although I've long felt the RNA expression data is so small as to be useless even to most specialists (though maybe this is a quirk of how my screen is set up?). Perhaps it would be more useful to the reader to replace the bar graphs with a
|most expressed in=field that auto-filled with text of either the top few hits or all cell types that had expression over some mark (the graphs currently show 3x the median, so perhaps that's a mark of interest?). Bonus points if we could make those top hits wikilink to the relevant wikipedia articles. Of course, the section would also have an external link to the microarray dataset this comes from. As an example, I'm looking at the BRCA1 infobox which shows this file and another. Thoughts? Ajpolino ( talk) 14:45, 15 July 2018 (UTC) - As a general point, collapsing content causes accessibility problems. Some fraction of users will have problems (e.g., RSI pain) with collapsed sections. You should think about whether that's an acceptable compromise. If the material is going to be used a moderate amount, then it's not reasonable to hide it. OTOH, if the material is so unwanted that almost nobody will un-collapse it, then does it need to be in the article at all? WhatamIdoing ( talk) 01:15, 16 July 2018 (UTC)
Regulatory RNA concern
I am a new Wikipedia Fellow in the midst of training. I entered " regulatory RNA" and RNAi came up. Of course that's the best worked out regulatory RNA system today, but there are other regulatory RNAs that should be accessed by the general term, such as Xist and others involved in X inactivation and enhancer RNAs, both of which are treated elsewhere on Wikipedia. Does this seem like a problem to others? Is it worth my while to work on putting together a more general treatment of regulatory RNA while I'm in this course? Thank you for considering this. LLMHoopes ( talk) 17:34, 30 July 2018 (UTC)
- Good catch. If we have additional existing articles that fit within the area of regulatory RNAs (I am not an expert in this field), a good place to start might be to turn regulatory RNA into a disambiguation page with links to RNAi and other regulatory RNAarticles (e.g., Long noncoding RNAs, ...). The disambiguation page might later be turned into a general article about regulatory RNAs. Boghog ( talk) 18:00, 30 July 2018 (UTC)
Estriadol metabolism
Hi everyone.
An editor asked me to reproduce File:Estradiol metabolism.png as a wikitext-annotated image. Would anyone here be willing to upload an svg image file with just the 11 compounds in that figure (without any image text except for the letters for the functional groups on the compounds, e.g., the OH for the hydroxyl group) under the file name File:Estradiol metabolitsm.svg? I can't really draw chemical structure diagrams but I can easily draw the pathways, so I intend to edit the uploaded image and add the lines and compound/enzyme annotations + adjust their placement as needed when annotating the image with a template.
@
Boghog: I'm pinging you in the event you're interested in helping with this again and aren't terribly busy.
![]() Seppi333 (
Insert 2¢)
22:10, 2 July 2018 (UTC)
Seppi333 (
Insert 2¢)
22:10, 2 July 2018 (UTC)
- Hi Seppi33. No problem. I will do this, but I need a day or two. Cheers.
Boghog (
talk)
23:59, 2 July 2018 (UTC)
- Thanks! There's no rush, so take whatever time you need.
Seppi333 (
Insert 2¢)
01:49, 3 July 2018 (UTC)
- @
Seppi333: Is this
File:Estradiol metabolism structures.svg OK?
Boghog (
talk)
07:58, 3 July 2018 (UTC)
- Yep, that's exactly what I was looking for, thanks! Could I trouble you to take a minute or two to annotate
the image file on Commons (via the "Add a note" button) with the name of each compound? It'd make it easier for me to determine where to place each structure diagram when I'm cross-referencing the svg/png files while drawing the pathway. Edit: I annotated 2 that I thought were obvious, but some of the others are throwing me off due to the stylistic/representational differences. E.g., stereochemistry, rotation, etc.
Seppi333 (
Insert 2¢)
08:41, 3 July 2018 (UTC); 08:54, 3 July 2018 (UTC)
 Done
Added names of structures.
Boghog (
talk)
06:07, 4 July 2018 (UTC)
Done
Added names of structures.
Boghog (
talk)
06:07, 4 July 2018 (UTC)
- Yep, that's exactly what I was looking for, thanks! Could I trouble you to take a minute or two to annotate
the image file on Commons (via the "Add a note" button) with the name of each compound? It'd make it easier for me to determine where to place each structure diagram when I'm cross-referencing the svg/png files while drawing the pathway. Edit: I annotated 2 that I thought were obvious, but some of the others are throwing me off due to the stylistic/representational differences. E.g., stereochemistry, rotation, etc.
Seppi333 (
Insert 2¢)
08:41, 3 July 2018 (UTC); 08:54, 3 July 2018 (UTC)
- @
Seppi333: Is this
File:Estradiol metabolism structures.svg OK?
Boghog (
talk)
07:58, 3 July 2018 (UTC)
- Thanks! There's no rush, so take whatever time you need.
Seppi333 (
Insert 2¢)
01:49, 3 July 2018 (UTC)
@ Boghog: The last thing I need before beginning to annotate and uniformly align the objects in this diagram is an svg image of estrone glucuronide's structure diagram. Could you email me that or add it to File:Estradiol metabolism structures.svg? If you'd prefer the former method, my email is in the source immediately following this sentence. Seppi333 ( Insert 2¢) 11:40, 7 August 2018 (UTC)
- Thanks again!
 Seppi333 (
Insert 2¢)
12:06, 7 August 2018 (UTC)
Seppi333 (
Insert 2¢)
12:06, 7 August 2018 (UTC)
Metabolic pathways of
estradiol in humans
|
References
- Hi Seppi. Per your request, I have added estrone glucuronide to the upper righthand corner of
File:Estradiol metabolism structures.svg.
Boghog (
talk)
12:20, 7 August 2018 (UTC)
- Much appreciated. Seppi333 ( Insert 2¢) 12:21, 7 August 2018 (UTC)
- Hi Seppi. Per your request, I have added estrone glucuronide to the upper righthand corner of
File:Estradiol metabolism structures.svg.
Boghog (
talk)
12:20, 7 August 2018 (UTC)
I think I'm more-or-less done. Anyone have any feedback/suggestions? Seppi333 ( Insert 2¢) 22:06, 10 August 2018 (UTC)
| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 5 | ← | Archive 7 | Archive 8 | Archive 9 | Archive 10 | Archive 11 |
I recently removed all of the image text from File:ISSN HMB statement Fig 1.jpg and replaced it with wikitext. The background image file is likely going to be changed to an svg format soon. Consequently, I'm open to restructuring the background image and repositioning/resizing the annotations based upon feedback from this project.
The original image file that this diagram is based upon was uploaded from a CC-BY-SA 2.0-copyrighted journal article about β-hydroxy β-methylbutyric acid (HMB), hence why there's a lot of emphasis on HMB in this image (note that there are a few minor differences between this image and the original). Unfortunately, the enzymes that catalyze the reactions HMB ↔ HMB-CoA and HMB-CoA ↔ HMG-CoA aren't currently known. Also, some very minor metabolites of leucine aren't shown in this image (L-Leucine → β-Leucine via leucine aminomutase, and β-Leucine → β-ketoisocaproic acid → β-Ketoisocaproyl-CoA → acetyl-CoA via a sequence of uncharacterized enzymes). This textbook diagram illustrates that tertiary pathway. Seppi333 ( Insert 2¢) 19:01, 2 December 2017 (UTC)
- One thing I'm not sure about is if it's correct to write "biotin+CO2" and "CO2+biotin" in this image as an input for MC-CoA carboxylase, or if it should say "CO2" as an input and " cofactor: biotin" next to the line beneath that substrate. Are either of those approaches fine? Seppi333 ( Insert 2¢) 20:07, 2 December 2017 (UTC)
- If anyone can comment on how to illustrate biotin as a cofactor for MC-CoA carboxylase in this image, I'd really appreciate it. Seppi333 ( Insert 2¢) 07:24, 10 December 2017 (UTC)
- Seppi333 Is it necessary to illustrate biotin? As a prosthetic group, it's really just part of the MC-CoA carboxylase holoenzyme. It's different to something like NADH or ATP where the cofactor is different before and after the reaction, and you need some separate process to regenerate it. But if you want to illustrate it, I'd suggest something like the original image, which doesn't give the impression that biotin is consumed like CO2 is. You could also just note the biotin requirement in the image caption.
- Also, is there a reason that reaction and its substrate and product appear in the diagram twice? As in, shouldn't the right-pointing arrow from HMB-CoA point to MC-CoA over on the right edge of the diagram, rather than there being a duplicate near the middle?
Adrian J. Hunter(
talk•
contribs)
04:30, 12 December 2017 (UTC)
- It probably isn't necessary TBH. I think the original author just included it because of the "when biotin is deficient" text in the image. I'll delete biotin from the image since you don't think it's particularly necessary to illustrate. As for why there's two pathways to HMG in the image... I'm not sure why it was illustrated that way. That's actually what I was hoping others would comment on. If you think the redundant MC-CoA and MG-CoA pathway in the bottom-center part of the image should come out, I'm okay with deleting that when the image is redrawn. Seppi333 ( Insert 2¢) 05:53, 12 December 2017 (UTC)
Given that the HMB article has been renominated at FAC for the 4th time after a 12 month wait, further feedback on changes to this diagram for the svg version would be very much appreciated. If you think some part of this diagram should be drawn, depicted, or annotated in a different manner, please say so. I'm looking for both general/non-specific feedback as well as specific feedback on one technical and one cosmetic point:
- The technical issue I'm looking for further feedback on is whether or not cofactors should be indicated and how to illustrate cofactors and/or inputs to each reaction if they're included in the image; i.e., should the inputs be depicted as they are now in the svg version and (re: Adrian's comment above) should biotin be annotated in the same manner as it was in the original image or be omitted? NB: the biotin annotation wasn't included in the current annotated image.
- The cosmetic issue I'm looking for further feedback on is whether the two pathways for HMB↔HMB-CoA↔MC-CoA↔MG-CoA↔HMG-CoA should be illustrated in the svg version, analogous to how it is in the current image, or if these should be merged into a single depiction when the image is redrawn as an svg diagram.
For additional context/information on the pathway being depicted in this diagram, it would probably help to read the article text in Leucine#Metabolism in humans. Seppi333 ( Insert 2¢) 08:22, 21 January 2018 (UTC)
- An essential requirement for Featured Articles is that it should be understood by a wide audience.IMHO the figure is way too complicated (i.e., likely to confuse readers) and should be simplified as much as possible without removing essential information. Hence I think co-factors should left out, unless they are directly referred to in the text (e.g., biotin). Splitting the digram into biosynthesis and metabolism will go a long way in making it easier to digest. Also the lower right hand side (MG-CoA → HMG-CoA → Acetyl-CoA/Acetoacetate) need not be included in either digram since this pathway has to do with metabolism of leucine, not HMB.
Boghog (
talk)
10:05, 21 January 2018 (UTC)
- Hehe, well I guess this speaks to how complicated the image is right now.
 The bottom right portion of this image is showing the metabolism of HMG-CoA, which is depicted as being downstream from isovaleryl-CoA; but, you'll notice that the pathway on the left also shows HMB-CoA being converted to both MC-CoA and HMG-CoA, and both of those are also depicted in the pathway on the right side (NB: if you click on the HMG-CoA, MC-CoA, or MG-CoA in this diagram and then view this image in those articles, you'll see 2 highlighted annotations for each of those pairs). Because HMG-CoA is downstream from HMB, the metabolism of HMG-CoA into acetyl-CoA and acetoacetate is actually downstream from HMB as well; however, this diagram makes it look like it isn't due the depiction of 1-way reactions (← or ↓) on the right side and reversible (↔) reactions on the left. In a nutshell, there's 2 interconnected pathways between HMB and HMG-CoA being shown, where different metabolic fates for HMG-CoA are shown on each side of the diagram (bottom left vs bottom right).
The bottom right portion of this image is showing the metabolism of HMG-CoA, which is depicted as being downstream from isovaleryl-CoA; but, you'll notice that the pathway on the left also shows HMB-CoA being converted to both MC-CoA and HMG-CoA, and both of those are also depicted in the pathway on the right side (NB: if you click on the HMG-CoA, MC-CoA, or MG-CoA in this diagram and then view this image in those articles, you'll see 2 highlighted annotations for each of those pairs). Because HMG-CoA is downstream from HMB, the metabolism of HMG-CoA into acetyl-CoA and acetoacetate is actually downstream from HMB as well; however, this diagram makes it look like it isn't due the depiction of 1-way reactions (← or ↓) on the right side and reversible (↔) reactions on the left. In a nutshell, there's 2 interconnected pathways between HMB and HMG-CoA being shown, where different metabolic fates for HMG-CoA are shown on each side of the diagram (bottom left vs bottom right). - So, I guess the two pathways that connect HMB to HMG-CoA in this image should be merged when creating the svg version.
Seppi333 (
Insert 2¢)
10:44, 21 January 2018 (UTC)
- I think we are on the same page now. There is a duplication in the parent diagram which should be eliminated in the daughter biosynthesis and metabolism diagrams. Boghog ( talk) 11:15, 21 January 2018 (UTC)
- Hehe, well I guess this speaks to how complicated the image is right now.
@Boghog: I'm referring to the reactions illustrated above.
|
@ Boghog: I'm creating the SVG diagram right now, but I just noticed something. Since MC-CoA→HMB-CoA requires H2O as an input and both HMB-CoA→HMG-CoA and MC-CoA→MG-CoA require CO2 as an input, wouldn't the MG-CoA→HMG-CoA reaction catalyzed by MG-CoA hydratase also require H2O as an input? If so, I should illustrate that in the diagram. Seppi333 ( Insert 2¢) 10:40, 28 January 2018 (UTC)
- A
hydratase by definition catalyzes the addition of water to a substrate, so yes, I think it should be illustrated in the diagram.
Boghog (
talk)
10:55, 28 January 2018 (UTC)
- File:HMB biosynthesis and metabolism diagram - no labels.svg - that's the outline. Now I need to annotate it. I'll probably upload a few more versions of that pathway outline with slight changes in the line placement while I'm working on the annotations though. Also, I decided to add a pathway for acetoacetate→ β-hydroxybutyrate to the annotated svg diagram. Seppi333 ( Insert 2¢) 11:34, 28 January 2018 (UTC)
@ Boghog: The current version of the annotated SVG diagram is transcluded from the template talk page into the collapse tab below. I've finished re-annotating most of it, but the annotations with an orange background need to be repositioned. Logging off for now. Let me know if you have any feedback. Seppi333 ( Insert 2¢) 13:00, 28 January 2018 (UTC)
- Looks good, but just to reiterate, I still think the diagram needs to be split in two. Also isovaleryl-CoA should be removed from both daughter diagrams since it is not involved in the biosynthesis of HMB and it appears to be a minor side pathway in HMB metabolism.
Boghog (
talk)
16:50, 28 January 2018 (UTC)
- Based upon the current layout, it would be pretty easy to display just the biosynthesis or the metabolism by template-cropping the top/bottom portions of the diagram and annotating over irrelevant lines using non-breaking spaces with a white background color. The full diagram is used in a number of other articles though. Seppi333 ( Insert 2¢) 18:15, 28 January 2018 (UTC)
@
Boghog: I've finished annotating the new svg diagram below. Unless you have any suggestions for further revision, I'm going to replace the current {{
Leucine metabolism in humans}} template with this one. I can work on splitting this diagram into two tomorrow if
Galobtter thinks it would be best to move the biosynthesis section out of the
beta-Hydroxy beta-methylbutyric acid#Pharmacology section.
Seppi333 (
Insert 2¢)
12:36, 29 January 2018 (UTC)
Annotated SVG version of the diagram
| |
|---|---|
|
|
Section reflist
Reflist for
{{
Leucine metabolism in humans}} |
|---|
|
FYI – 4th FAC nomination
Beta-Hydroxy beta-methylbutyric acid recently became a good article, so I renominated it FAC for the 4th time: Wikipedia:Featured article candidates/Beta-Hydroxy beta-methylbutyric acid/archive4.
I really need more editors to comment at this nomination and review it against the WP:Featured article criteria. For those who haven't reviewed an article at FAC before, this is the "FAQ" page for reviewing articles at FAC. Seppi333 ( Insert 2¢) 03:46, 2 February 2018 (UTC)
Question about the term "mTOR"
Figured that folks may be interested in this discussion. Jo-Jo Eumerus ( talk, contributions) 14:29, 4 February 2018 (UTC)
There is a discussion at the above link. Your input is most welcome. Boghog ( talk) 17:39, 11 February 2018 (UTC)
Requesting help with Cooperative antigen transfer
Hello there! I just came across this article ( Cooperative antigen transfer) and I was wondering if an expert could take a look at it? I know little about the topic. -- TheSandDoctor ( talk) 20:20, 12 February 2018 (UTC)
- Looks like WP:OR to me. I have nominated it for deletion. Boghog ( talk) 21:11, 12 February 2018 (UTC)
Please see the latest discussion at Talk:Sex, which proposes to restrict the scope of the article. -- EncycloPetey ( talk) 16:52, 18 February 2018 (UTC)
Umami and Savoriness
Please share your views here: Talk:Umami#This really should be called savory
Anna Frodesiak ( talk) 01:06, 24 February 2018 (UTC)
Facto Post – Issue 10 – 12 March 2018
Facto Post – Issue 10 – 12 March 2018
 Milestone for mix'n'matchAround the time in February when Wikidata clicked past item Q50000000, another milestone was reached: the mix'n'match tool uploaded its 1000th dataset. Concisely defined by its author, Magnus Manske, it works "to match entries in external catalogs to Wikidata". The total number of entries is now well into eight figures, and more are constantly being added: a couple of new catalogs each day is normal. Since the end of 2013, mix'n'match has gradually come to play a significant part in adding statements to Wikidata. Particularly in areas with the flavour of digital humanities, but datasets can of course be about practically anything. There is a catalog on skyscrapers, and two on spiders. These days mix'n'match can be used in numerous modes, from the relaxed gamified click through a catalog looking for matches, with prompts, to the fantastically useful and often demanding search across all catalogs. I'll type that again: you can search 1000+ datasets from the simple box at the top right. The drop-down menu top left offers "creation candidates", Magnus's personal favourite. m:Mix'n'match/Manual for more. For the Wikidatan, a key point is that these matches, however carried out, add statements to Wikidata if, and naturally only if, there is a Wikidata property associated with the catalog. For everyone, however, the hands-on experience of deciding of what is a good match is an education, in a scholarly area, biographical catalogs being particularly fraught. Underpinning recent rapid progress is an open infrastructure for scraping and uploading. Congratulations to Magnus, our data Stakhanovite! Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. If you wish to receive no further issues of Facto Post, please remove your name from
our mailing list. Alternatively, to opt out of all
massmessage mailings, you may add
Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery ( talk) 12:26, 12 March 2018 (UTC)
Hi. I have been thinking over an issue and would appreciate a second opinion.
We have an article about cobalamin, which is another name for vitamin B12. The problem is, as such things sometimes are with vitamins (see vitamin B3 complex/ niacin), that it is a group name for a number of compounds that have a certain structure in common, but depending on the ligand attached they have different names, and slightly different properties, as well as some differences in biological functions. And I was wondering maybe it should be a disambig page?
On the one hand, something indeed can be said about cobalamin as a group, on the other hand it rapidly boils down to "if ligand is this, then it is such and such name with such and such properties". So basically, it has a disambig function.
What do you think, what could be put in that article, if it should remain an article? PubChem has a lot of them for reference (as well as vitamin B12) -- Helixitta ( t.) 14:04, 10 March 2018 (UTC)
- Maybe "Vitamin B12" should discuss the medical/biological/nutritional aspects and "cobalamin" the chemical ones?
Jo-Jo Eumerus (
talk,
contributions)
15:20, 10 March 2018 (UTC)
- I'm not sure. Chemical properties actually lead to biological ones, and usually the article on a substance would have everything in it. Say,
glucose entry has biological function, chemical and physical properties. But I suppose some chemical properties that are common for the group could be added to the article about
cobalamin... --
Helixitta (
t.)
16:23, 10 March 2018 (UTC)
- Similar issues occurred with
vitamin D and
cholecalciferol. The feeling here was that all the drug and therapeutic issues associated with vitamin D deficiency had to be rehashed at the colecalciferol page, and take priority, rather than the chemical and physiological details.
Jrfw51 (
talk)
19:13, 10 March 2018 (UTC)
- Thanks for that example. -- Helixitta ( t.) 19:57, 12 March 2018 (UTC)
- Similar issues occurred with
vitamin D and
cholecalciferol. The feeling here was that all the drug and therapeutic issues associated with vitamin D deficiency had to be rehashed at the colecalciferol page, and take priority, rather than the chemical and physiological details.
Jrfw51 (
talk)
19:13, 10 March 2018 (UTC)
- I'm not sure. Chemical properties actually lead to biological ones, and usually the article on a substance would have everything in it. Say,
glucose entry has biological function, chemical and physical properties. But I suppose some chemical properties that are common for the group could be added to the article about
cobalamin... --
Helixitta (
t.)
16:23, 10 March 2018 (UTC)
Adenosine triphosphate
Adenosine triphosphate, an article that you or your project may be interested in, has been nominated for an individual good article reassessment. If you are interested in the discussion, please participate by adding your comments to the reassessment page. If concerns are not addressed during the review period, the good article status may be removed from the article. AIRcorn (talk) 22:42, 26 March 2018 (UTC)
A thorough review of the gene article
- Transcluded from Talk:Gene/Review
Article review
|
|---|
|
To WP:MCB, WP:GEN, WP:BIOL and WP:EB The gene article gets 50,000 views per month but has been de-listed as a featured article since 2006. Given the success of the recent blitz on the enzyme article, I thought I'd suggest spending a couple of weeks seeing if we can get it up to a higher standard. I'm going to start with updating some of the images. If you'd like to help out on the article, it'd be great to see you there. T.Shafee(Evo﹠Evo) talk 09:49, 31 March 2015 (UTC)
GlossarySnooping around I encountered Template:Genetics glossary, I don't know it's backstory, but it is a rather cleaver idea for a template in my opinion. I partially reckon it might go well under the first image in place or the second image depicting DNA, which conceptually is a tangent. I am not sure, hence my asking. -- Squidonius ( talk) 21:47, 1 April 2015 (UTC)
ReferencesI'm planning on adding some more Molecular Biology of the Cell references to the article using
I've not done much non-standard reference citation so I'll wait until you've done a couple so that I can see the format in context before doing any more. The ones I added yesterday shouldn't be too difficult to reformat.
T.Shafee(Evo﹠Evo)
talk
12:24, 22 April 2015 (UTC)
MBOC referencesArticle Genes [1]: 2 are numerous [1]: 4 and useful [1]: 4.1 References
So {{ rp}} labels the chapter number but does not provide any easy link to the actual information. Therefore it's combined with a list of chapter links. the benefit is that the {{ rp}} template is relatively easy to maintain and the list of chapter links doesn't require maintainance and places all the MBOC links together. As stated above, there's basically no way to avoid linking individually to chapters if we want to cite MBOC. I'll finish building the chapter list over the next couple of days. T.Shafee(Evo﹠Evo) talk 01:29, 27 April 2015 (UTC)
Collapsed the transclusion. Seppi333 ( Insert 2¢) 04:48, 27 March 2018 (UTC) |
Any interest in rescuing this abandoned draft? Espresso Addict ( talk) 01:27, 9 April 2018 (UTC)
Facto Post – Issue 11 – 9 April 2018
Facto Post – Issue 11 – 9 April 2018
 The 100 Skins of the OnionOpen Citations Month, with its eminently guessable hashtag, is upon us. We should be utterly grateful that in the past 12 months, so much data on which papers cite which other papers has been made open, and that Wikidata is playing its part in hosting it as "cites" statements. At the time of writing, there are 15.3M Wikidata items that can do that. Pulling back to look at open access papers in the large, though, there is is less reason for celebration. Access in theory does not yet equate to practical access. A recent LSE IMPACT blogpost puts that issue down to "heterogeneity". A useful euphemism to save us from thinking that the whole concept doesn't fall into the realm of the oxymoron. Some home truths: aggregation is not content management, if it falls short on reusability. The PDF file format is wedded to how humans read documents, not how machines ingest them. The salami-slicer is our friend in the current downloading of open access papers, but for a better metaphor, think about skinning an onion, laboriously, 100 times with diminishing returns. There are of the order of 100 major publisher sites hosting open access papers, and the predominant offer there is still a PDF.  From the discoverability angle, Wikidata's bibliographic resources combined with the SPARQL query are superior in principle, by far, to existing keyword searches run over papers. Open access content should be managed into consistent HTML, something that is currently strenuous. The good news, such as it is, would be that much of it is already in XML. The organisational problem of removing further skins from the onion, with sensible prioritisation, is certainly not insuperable. The CORE group (the bloggers in the LSE posting) has some answers, but actually not all that is needed for the text and data mining purposes they highlight. The long tail, or in other words the onion heart when it has become fiddly beyond patience to skin, does call for a pis aller. But the real knack is to do more between the XML and the heart. Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. If you wish to receive no further issues of Facto Post, please remove your name from
our mailing list. Alternatively, to opt out of all
massmessage mailings, you may add
Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery ( talk) 16:25, 9 April 2018 (UTC)
Move request
Talk:GBA_(disambiguation)#Requested_move_13_April_2018 Jytdog ( talk) 02:59, 15 April 2018 (UTC)
Remove tags for Wikipedia:WikiProject Cell Signaling?
Wikipedia:WikiProject Cell Signaling has been inactive for several years, yet its tag is on the talk pages of 295 pages. Should we remove the Cell signalling banner from those pages and replace it with the MCB banner where appropriate? I figure the purpose of the project tags on article talk pages is (1) to help projects keep track of pages their editors are interested in, and (2) to point readers/editors towards a Project that could help address concerns about a particular article. It seems like having the banner from a defunct WikiProject just serves to confuse. Thoughts? I assume a bot could be found to do this task if folks think it's a good idea. Cheers Ajpolino ( talk) 19:28, 24 April 2018 (UTC)
- At the top of the Wikipedia:WikiProject Cell Signaling page, there is in big bold letters with links: This MCB project subpage is no longer in use and is kept as a historical archive. Please go to the MCB project homepage or talk page for currently active sections. That seems like a sufficient pointer to our active WP. I don't think we need to completely bypass the cell signaling sub-WP. On the other hand, I don't know what is usually done with defunct WP--are they typically bypassed? -- Mark viking ( talk) 20:06, 24 April 2018 (UTC)
Merger discussion for Vitamin B3

An article that you have been involved in editing— Vitamin B3—has been proposed for merging with another article. If you are interested, please participate in the merger discussion. Thank you. SusanLesch ( talk) 13:59, 30 April 2018 (UTC)
Wikiproject Anatomy and the scope of your project
Hi, I am not a member of your project but do most of my work at Wikiproject Anatomy. At the anatomy project we recently finished classifying our articles and we have just under 1500 articles and redirects related to cells and microanatomy. What I would to know is what is under the scope of your project? Do you include individual cell types such as alveolar cells, osteocytes or melanocytes? If such articles are within your scope please let me know and I will gladly start tagging/adding your project banner to their talk page (in the hope that your project members can find and will work on some of the articles of interest to both wikiprojects). Kind regards JakobSteenberg ( talk) 20:47, 29 April 2018 (UTC)
- In my opinion such cell types would also fall within the scope WP:MCB. Some of them are already tagged (e.g. most of the immune cell types). I'll be interested to hear the opinions of others.
T.Shafee(Evo&Evo)
talk
23:56, 29 April 2018 (UTC)
- Thanks for your input. I will wait for others to comment before doing anything. Just be sure;
histology-articles such as
histology of the vocal folds,
Loop of Henle or
Portal triad are not within the scope of MCB correct?
JakobSteenberg (
talk)
17:55, 30 April 2018 (UTC)
- I agree with Evo. With respect to morphology and cell type, the two projects clearly overlap. Boghog ( talk) 18:38, 30 April 2018 (UTC)
- Thanks for your input. I will wait for others to comment before doing anything. Just be sure;
histology-articles such as
histology of the vocal folds,
Loop of Henle or
Portal triad are not within the scope of MCB correct?
JakobSteenberg (
talk)
17:55, 30 April 2018 (UTC)
LCHN: An Error has occurred retrieving Wikidata item for infobox
I just moved Draft:LCHN into mainspace. @ M.Gray97: The infobox_gene template is generating An Error has occurred retrieving Wikidata item for infobox. My template-fu is insufficient to fix this. Could a SME take a look? Thanks. -- RoySmith (talk) 21:17, 6 May 2018 (UTC)
Draft:Comparison_of_DNA_melting_prediction_software
Draft:Comparison_of_DNA_melting_prediction_software I'm a Afc reviewer with cell biology background but still struggling whether this bioinformatics page can be accepted . do give me an opinion. thanks much Quek157 ( talk) 16:22, 8 May 2018 (UTC)
3D protein/nucleic acid structures could be rotateable 3D models: NGL
Hey everyone. I'm no wiki expert and there might be internal reasons not to do this, but I thought I'd float it.
There is an excellent+beautiful+open source molecular graphics thing called NGL. It is being developed at UCSD as part of the protein data bank, and I know the person doing it. Some examples: Ribosome https://www.rcsb.org/3d-view/4V4J RNA pseudoknot http://www.rcsb.org/3d-view/2LC8 Dorothy Hodgkin's Insulin https://www.rcsb.org/3d-view/4INS
You can see a (super cool+interactive) presentation on it here http://nglviewer.org/talks/ngl-3dsig/
Adopting ngl would be great for wiki because: -Everything would look super beautiful and make the molecular biology pages more readable -It would allow you to have images of any molecule for which we have a model (= any molecule for which you would want to see an "image") on any page, because you wouldn't need to find a public domain picture. In principle you could just have some markup like <ngl window: "pdbID:4ins"> or whatever and suddenly your webpage has an excellent up-to-date structure
NGL is very cross platform and works well on smartphones. However, it is obviously dependent on webGL.
Using NGL would take more bandwidth. For each structure you'd need to download a pdb file, plus the ngl javascript (which you'd have to do for every page?). That said, I'd like to point out that a 500KB pdb file is smaller than a 1MB animated gif of the same molecule performing a rotation, so a small number of pages would be compressed by this adoption.
NGL is better than the visualizer that proteopedia use http://proteopedia.org/wiki/index.php/Main_Page as you can verify for yourself. — Preceding unsigned comment added by Hamishtodd1 ( talk • contribs) 12:06, 10 May 2018 (UTC)
- The proteopedia visualizer I use is FirstGlance ... see http://proteopedia.org/wiki/index.php/FirstGlance_in_Jmol and https://bioinformatics.org/firstglance/fgij/. - Dank ( push to talk) 13:56, 10 May 2018 (UTC)
Discussion at Wikipedia talk:WikiProject Medicine#MicroDNA
![]() You are invited to join the discussion at
Wikipedia talk:WikiProject Medicine#MicroDNA. –
Finnusertop (
talk ⋅
contribs)
22:53, 10 May 2018 (UTC)
You are invited to join the discussion at
Wikipedia talk:WikiProject Medicine#MicroDNA. –
Finnusertop (
talk ⋅
contribs)
22:53, 10 May 2018 (UTC)
Facto Post – Issue 12 – 28 May 2018
Facto Post – Issue 12 – 28 May 2018
 ScienceSource fundedThe Wikimedia Foundation announced full funding of the ScienceSource grant proposal from ContentMine on May 18. See the ScienceSource Twitter announcement and 60 second video.
The proposal includes downloading 30,000 open access papers, aiming (roughly speaking) to create a baseline for medical referencing on Wikipedia. It leaves open the question of how these are to be chosen. The basic criteria of WP:MEDRS include a concentration on secondary literature. Attention has to be given to the long tail of diseases that receive less current research. The MEDRS guideline supposes that edge cases will have to be handled, and the premature exclusion of publications that would be in those marginal positions would reduce the value of the collection. Prophylaxis misses the point that gate-keeping will be done by an algorithm. Two well-known but rather different areas where such considerations apply are tropical diseases and alternative medicine. There are also a number of potential downloading troubles, and these were mentioned in Issue 11. There is likely to be a gap, even with the guideline, between conditions taken to be necessary but not sufficient, and conditions sufficient but not necessary, for candidate papers to be included. With around 10,000 recognised medical conditions in standard lists, being comprehensive is demanding. With all of these aspects of the task, ScienceSource will seek community help. Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. ScienceSource pages will be announced there, and in this mass message. If you wish to receive no further issues of Facto Post, please remove your name from
our mailing list. Alternatively, to opt out of all
massmessage mailings, you may add
Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery ( talk) 10:16, 28 May 2018 (UTC)
WikiProject collaboration notice from the Portals WikiProject
The reason I am contacting you is because there are one or more portals that fall under this subject, and the Portals WikiProject is currently undertaking a major drive to automate portals that may affect them.
Portals are being redesigned.
The new design features are being applied to existing portals.
At present, we are gearing up for a maintenance pass of portals in which the introduction section will be upgraded to no longer need a subpage. In place of static copied and pasted excerpts will be self-updating excerpts displayed through selective transclusion, using the template {{ Transclude lead excerpt}}.
The discussion about this can be found here.
Maintainers of specific portals are encouraged to sign up as project members here, noting the portals they maintain, so that those portals are skipped by the maintenance pass. Currently, we are interested in upgrading neglected and abandoned portals. There will be opportunity for maintained portals to opt-in later, or the portal maintainers can handle upgrading (the portals they maintain) personally at any time.
Background
On April 8th, 2018, an RfC ("Request for comment") proposal was made to eliminate all portals and the portal namespace. On April 17th, the Portals WikiProject was rebooted to handle the revitalization of the portal system. On May 12th, the RfC was closed with the result to keep portals, by a margin of about 2 to 1 in favor of keeping portals.
Since the reboot, the Portals WikiProject has been busy building tools and components to upgrade portals.
So far, 84 editors have joined.
If you would like to keep abreast of what is happening with portals, see the newsletter archive.
If you have any questions about what is happening with portals or the Portals WikiProject, please post them on the WikiProject's talk page.
Thank you. — The Transhumanist 10:59, 31 May 2018 (UTC)
Is this worth salvaging? (Delete vs. move to mainspace vs. something else?) Calliopejen1 ( talk) 02:22, 4 June 2018 (UTC)
- It probably should be merged into Insulin#Regulation. The sources need to be moved in-line and it needs some copyediting first. I will work on this when I find time. Boghog ( talk) 03:48, 4 June 2018 (UTC)
- Not needed as a standalone article when it fits nicely within Insulin. Natureium ( talk) 14:14, 4 June 2018 (UTC)
Large copy-paste at Lysine
A large paste of text from WikiJournal of Science/Lysine: biosynthesis, catabolism and roles has been dropped into the article on Lysine. In addition to a number of references breaking in the process, there are issues of overly technical content and copy-paste, even if copyright issues do not apply. I'm too busy to dig into the issues more deeply for at least a few days, so I'm going to punt it over here for now. Sorry and thanks, BiologicalMe ( talk) 03:39, 10 June 2018 (UTC)
- Fixed the link. Adrian J. Hunter( talk• contribs) 05:01, 10 June 2018 (UTC)
-
BiologicalMe it looks like
Evolution and evolvability was in the middle of working on this when you posted here, and has edited extensively since. Not sure there's any remaining issues?
Adrian J. Hunter(
talk•
contribs)
05:01, 10 June 2018 (UTC)
- Hello, thanks for flagging this here
BiologicalMe. I've been in the process of integrating an updated version of the article that has been written at the location you mentioned. It's under a CC-BY license, so the copyright isn't a specific issue (links to the relevant sections of the journal's
editorial and
ethics guidelines). You're correct that the ref errors were introduced during the process which I was fixing as i went on (also thanks to boghog for their ever-present assistance!). The main thing of importance is
WP:TECHNICAL. I believe that many of the updates have largely made the article less technical (especially the lead;
prev version), however I fully support improving readability and accessibility where possible. I also removed mentions of lysine deficiency causing blindness for now, since it was unreferenced. As well as this specific case, feedback from Wikipedians on the journals' processes is very welcome (see also
list of Wikipedia articles updated based on academic journal content). We try to operate WikiJournals in a transparent, open and accountable format, so there is a general discussion board for the WikiJournal User Group
here, as well as discussion boards or each journal.
T.Shafee(Evo&Evo)
talk
05:32, 10 June 2018 (UTC)
- Thanks
User:Evolution and evolvability. I knew I was too tired to address it coherently, and I knew I would have no free time today to figure out copyleft licence compatibility or scramble through the references. As an indication of how tired I was, I forgot to provide a diff or to tag you. I'm looking forward to reading through what you've done, fully alert. Hopefully tomorrow.
BiologicalMe (
talk)
15:27, 10 June 2018 (UTC)
- No worries! You were absolutely right to post here. T.Shafee(Evo&Evo) talk 00:39, 11 June 2018 (UTC)
- Thanks
User:Evolution and evolvability. I knew I was too tired to address it coherently, and I knew I would have no free time today to figure out copyleft licence compatibility or scramble through the references. As an indication of how tired I was, I forgot to provide a diff or to tag you. I'm looking forward to reading through what you've done, fully alert. Hopefully tomorrow.
BiologicalMe (
talk)
15:27, 10 June 2018 (UTC)
- Hello, thanks for flagging this here
BiologicalMe. I've been in the process of integrating an updated version of the article that has been written at the location you mentioned. It's under a CC-BY license, so the copyright isn't a specific issue (links to the relevant sections of the journal's
editorial and
ethics guidelines). You're correct that the ref errors were introduced during the process which I was fixing as i went on (also thanks to boghog for their ever-present assistance!). The main thing of importance is
WP:TECHNICAL. I believe that many of the updates have largely made the article less technical (especially the lead;
prev version), however I fully support improving readability and accessibility where possible. I also removed mentions of lysine deficiency causing blindness for now, since it was unreferenced. As well as this specific case, feedback from Wikipedians on the journals' processes is very welcome (see also
list of Wikipedia articles updated based on academic journal content). We try to operate WikiJournals in a transparent, open and accountable format, so there is a general discussion board for the WikiJournal User Group
here, as well as discussion boards or each journal.
T.Shafee(Evo&Evo)
talk
05:32, 10 June 2018 (UTC)
Similar Pages?
Note from OTRS ticket 2018062010005412 - the following pages appear to duplicate the same subject
Ronhjones (Talk) 22:27, 24 June 2018 (UTC)
- I think they're sufficiently distinct to warrant separate pages.
C/EBP is a
family of proteins, which includes
CEBPA,
CEBPB,
CEBPG etc.
T.Shafee(Evo&Evo)
talk
22:52, 24 June 2018 (UTC)
- Yeah, not duplicates. There is more than one C/EBP protein.
Jo-Jo Eumerus (
talk,
contributions)
12:46, 25 June 2018 (UTC)
- Thanks, not my field - mine is Chemistry. I'll let the OP know Ronhjones (Talk) 16:18, 25 June 2018 (UTC)
- Yeah, not duplicates. There is more than one C/EBP protein.
Jo-Jo Eumerus (
talk,
contributions)
12:46, 25 June 2018 (UTC)
We have separate articles on Galactoside O-acetyltransferase and Beta-galactoside transacetylase, but are they really different enzymes? The first two sources used at Beta-galactoside transacetylase seem to classify them as synonymous: galactoside acetyltransferase (thiogalactoside transacetylase) (thiogalactoside transacetylase, LacA, GAT). Vaselineeeeeeee ★★★ 22:31, 22 June 2018 (UTC)
- @ Vaselineeeeeeee: I'm pretty sure you're correct. I think the two names are synonyms fort teh same anzyem (just slightly different description s of its catalytic activity). "Galactoside acetyltransferase" seems to be the most common variant of the name in the literature. T.Shafee(Evo&Evo) talk 00:39, 26 June 2018 (UTC)
- Additional consideration:
Galactoside O-acetyltransferase is the formal
Enzyme Commission number naming but the least-commonly used in publications!
T.Shafee(Evo&Evo)
talk
00:51, 26 June 2018 (UTC)
- @ Evolution and evolvability: Yeah I noticed that too. You think we should merge to the Galactoside O-acetyltransferase page and possibly get rid of the O- for common name? Vaselineeeeeeee ★★★ 02:22, 26 June 2018 (UTC)
Human Genome had a whole section of creationist claptrap for over three years
[1]
[2]
![]() Adrian J. Hunter(
talk•
contribs)
12:40, 25 June 2018 (UTC)
Adrian J. Hunter(
talk•
contribs)
12:40, 25 June 2018 (UTC)
- Ouch! Thanks for spotting and fixing that. Boghog ( talk) 16:34, 25 June 2018 (UTC)
- Seconded. Well done.
T.Shafee(Evo&Evo)
talk
00:01, 26 June 2018 (UTC)
- Ah, sorry my post was misleading. The problem was spotted and fixed in the first ever edit from an IP, in the second diff linked above. Adrian J. Hunter( talk• contribs) 08:53, 26 June 2018 (UTC)
Facto Post – Issue 13 – 29 May 2018
Issue 13 – 29 May 2018
| |
|---|---|
MediaWiki message delivery ( talk) 18:19, 29 June 2018 (UTC) |
Discussion on readability of some medical / biology articles and their infoboxes at WT:PHARM
See Wikipedia_talk:WikiProject_Pharmacology#Readability_of_medical_physiology_articles. -- Tom (LT) ( talk) 18:27, 3 July 2018 (UTC)
Readability of medical physiology articles
There is a discussion concerning readability of {{ Infobox gene}} at the above link that is well within the scope this project. In fact, the discussion probably been started here, rather than there. In any case, your input is welcome. Boghog ( talk) 20:06, 5 July 2018 (UTC)
Cellosaurus
There is a notability/ COI discussion over at Talk:Cellosaurus. Both this discussion and the article itself would benefit from input from WP:MCB members. -- Daniel Mietchen ( talk) 15:49, 10 July 2018 (UTC)
See Wikipedia:Featured article review/Antioxidant/archive1. Seppi333 ( Insert 2¢) 22:34, 10 July 2018 (UTC)
Facto Post – Issue 14 – 21 July 2018
Issue 14 – 21 July 2018
| |
|---|---|
MediaWiki message delivery ( talk) 06:10, 21 July 2018 (UTC) |
Improvements to infobox gene
I can see a great deal of effort has gone to producing infobox gene and it clearly has some amazing wikidata integration. I think it could be improved in a number of ways and propose some specific suggestions here for discussion. My aim is to make the infobox shorter, easier to read, and increase the prominence of basic information. (I would normally make a sandbox of my proposed edits but am unable to because the infobox is written in lua.) For my inspiration, I am viewing BRCA1 and MLH1.
I'm starting this part of the discussion back here (rather than WT:PHARM), as boghog points out, this being a more appropriate venue. Ping to editors that have participated: Boghog, Seppi333, Jytdog, Looie496, Ajpolino, Tryptofish. Apologies if I've left anyone out. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- + Andrew Su, Daniel Mietchen and Opabinia regalis who may have useful ideas. T.Shafee(Evo&Evo) talk 12:05, 16 July 2018 (UTC)
1. Move some data to an 'authority control' template
We're moving in that direction globally on WP. The list of 'external IDs' in the infobox could easily be moved to an authority control template for articles. This would also help shorten the infobox, reduce the length of its code, and more importantly, simplify the infobox, which is my goal here.-- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support - I find many of the links useful, but I'm fairly confident the average reader does not. The minority of our readers who follow these links (probably biologists) can find them just as well at the bottom of the page. This has been done with taxonomic identifiers for many organism articles as well. Ajpolino ( talk) 20:09, 14 July 2018 (UTC)
- Strong oppose – As long as these links are toward the bottom of the infobox, I do not see the need for this. Many of the articles that transcribe this infobox are somewhat obscure and not often read by the average reader. It is more convenient for the specialist to have all the relevant links in the same infobox. Also most of the links in question are not authority control. They are links to specialist databases that provide more detailed information about the gene/protein. Boghog ( talk) 22:16, 14 July 2018 (UTC)
- Oppose I think collapsing this to avoid cluttering the box with things that won't interest many people without moving it to the bottom of the page is the best solution. Natureium ( talk) 20:09, 16 July 2018 (UTC)
2. Remove pictures of chromosomes

I refer here to the full chromosome panel, not the horizontal individual chromosome with the gene's location boxed. These pictures take up a lot of space, add to loading times, and don't contribute anything factual. They are also duplicated in text (twice) in the infobox.-- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Collapse would be a better option. I agree that this figure is mainly eye candy. Boghog ( talk) 22:17, 14 July 2018 (UTC)
- Collapse, with removal as a secondary option. The diagram would be able to justify itself more if the exact chromosomal location was pointed to.
T.Shafee(Evo&Evo)
talk
11:48, 16 July 2018 (UTC)
- Agree. If the exact location were here that'd be great. As it is, our description of "chromosome 9" is backed up by a numbered group of chromosomes and the ninth selected. I just don't see how this is useful? -- Tom (LT) ( talk) 10:06, 19 July 2018 (UTC)
- Oppose I think this is a nice visual for people who aren't very familiar with genetics but may be looking for information for personal health purposes.
Natureium (
talk)
20:09, 16 July 2018 (UTC)
- But what person not familiar with genetics but using this for their personal health will have access to a karyotype, and thereby find comparing this with their karyotype useful...?
3. Collapse orthologs section by default
The section takes up a large amount of vertical space and is limited to links. I propose collapsing it by default. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Oppose – I find those links very useful. Furthermore they are at the bottom of the infobox so they should not obscure more general information at the top of the box. What could be done to reduce it size is to display only one Refseq link like the previous {{ GNF Protein box}} did. Boghog ( talk) 22:23, 14 July 2018 (UTC)
- I'd support either collapsing the section by default (still leaves it accessible to those interested) or reducing the display to one Refseq link (cuts down on the external link overload in the infobox). I'd also support doing both of those together. Ajpolino ( talk) 14:45, 15 July 2018 (UTC)
- Weakly Support This would save space, but I don't think it's much of an improvement. Natureium ( talk) 20:09, 16 July 2018 (UTC)
4. Rename 'available structures' section
There is a title that is called 'available structures' but it seems to be just some links. I don't know what it does or how it relates to genes, and therefore presume most lay readers won't either. I propose renaming it to something that explains what it does, or merging it to identifiers section if it is related. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment – The available structures section refers to 3D structures of the protein stored in the
Protein Data Bank and are quite distinct from identifiers. Function derives from structure hence those links are essential. I use them all the time.
Boghog (
talk)
22:29, 14 July 2018 (UTC)
- Ah. @
Boghog then these would be suitable to move to an external links section of the infobox? --
Tom (LT) (
talk)
22:47, 14 July 2018 (UTC)
- The left-hand side (green shaded) are explanatory internal links. Almost everything in the right-hand side (grey-shaded) is externally linked. It makes no sense to move only the Protein Databank External links to an external link section when almost everything else in the infobox is also an external link. Boghog ( talk) 06:49, 15 July 2018 (UTC)
- The rationale for placing the PBD links right below the infobox graphic is that the graphics are most commonly of the protein structure derived from one the PDB links. There are often more than one available protein structure (for example structures of different domains of the protein, complexes with different ligands or with other proteins, etc.). Having all the PDB links in one location right below the graphic of the structure gives a more complete view of the structure. Boghog ( talk) 07:54, 15 July 2018 (UTC)
- Ah. @
Boghog then these would be suitable to move to an external links section of the infobox? --
Tom (LT) (
talk)
22:47, 14 July 2018 (UTC)
- Support - Rename to something like "Protein structure" or equivalent (i.e. not gene structure, or chromosomal structure). The "Available" is unnecessary (if there were none available, the section wouldn't be shown. Perhaps
{{ Infobox protein}}could be merged in? T.Shafee(Evo&Evo) talk 11:53, 16 July 2018 (UTC)- This option makes clear the point of the section and, as Evolution and evolvability points out, the 'available' is superfluous - so has my full support. -- Tom (LT) ( talk) 10:03, 19 July 2018 (UTC)
- Weakly oppose This is an explanatory enough title and no one has proposed a better alternative. Natureium ( talk) 20:09, 16 July 2018 (UTC)
5. Create a 'basic information' section and put it at the top of the infobox.
A 'basic information' section should be created and placed beneath the picture of the protein ribbon and include basic information about the gene written in a human-readable format. Fields can be discussed once we have some support for this section. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support - There was substantial discussion at Wikipedia_talk:WikiProject_Pharmacology#Readability_of_medical_physiology_articles regarding what fields might constitute basic information and where database entries to populate these fields could be found. I agree with points raised by Boghog and Seppi333 at that earlier discussion that to be both reliable and human-readable, this may have to be curated (or completely written) manually. I'm happy to help with this. Ajpolino ( talk) 20:09, 14 July 2018 (UTC)
- Support – Giving a birds eye view of the function, mechanism, and location of the protein is a very worthwhile goal. This can be done by manually selecting the most important GO terms for display while leaving the complete list in a collapsed state. Boghog ( talk) 22:32, 14 July 2018 (UTC)
- support per Boghog rationale-- Ozzie10aaaa ( talk) 10:11, 16 July 2018 (UTC)
- oppose and strongly this is trying to solve a content problem with an infobox which is wrong headed -- as wrong headed as the WMF trying to solve our cluttered leads by grabbing the "description" field from Wikidata and forcing it onto WP articles on mobile and in apps. (which in turn has generated a ton of more work for us, since the "solution" was creating a new
{{ short description}}field for every article in WP.) Now a second field of "summary information" is being proposed.. and this is just crazy to me. But I guess people will volunteer doing what they want.
- What bugs me, is that both of these things have nothing to do with fixing the bad leads that are the source of the problems. If the leads of these articles were doing their job this would not be needed. Most importantly the time it would take to do this should really be put into actually fixing the leads. But again, people will spend their time here doing as they will....
Jytdog (
talk)
16:39, 16 July 2018 (UTC)
- I don't really see what the relevance of this infobox discussion is w.r.t. article leads. The infobox summarizes body content (and even content not covered in the body) in a succinct tabular format while also providing relevant external links. A lead and an infobox serve complementary roles in the sense that one conveys information in prose and the other in a data table. Each has its own advantages over the other; hence, the use of both is ideal. Fixing article leads is another issue altogether. Seppi333 ( Insert 2¢) 03:18, 17 July 2018 (UTC)
- The goal of the
Gene Wiki is to create a Wikipedia article for every notable human gene starting with an infobox and public domain Entrez description of the gene/protein. The intention was this seed material would encourage editors in expanding these articles. Unfortunately majority of these articles are still stubs. In addition, many of the Entrez descriptions are either incomplete or missing all together. Adding a "key properties" section to the infobox would go a long way to provide a minimum level of information for each stub. For more complete articles, the "key properties" section may highlight basic information that is lacking in the lead and encourage editors to add that information. Finally it should be pointed out that there are currently over 11,000
Gene Wiki articles. Expanding and cleaning up the leads for all of these articles would be an enormous task and there are simply not enough editors to do this in a reasonable time frame. Adding a manually curated "key properties" section is also a lot of work, but could be accomplished much more quickly. After the "key properties" sections have been added, we will of course continue to work on the leads, but that work will take much longer.
Boghog (
talk)
13:24, 17 July 2018 (UTC)
- The OP
started by saying
I often use WP as revision and for my work as I'm sure many people do. I find many of our articles (e.g. Beta-2 adrenergic receptor, which I'm looking at now) very difficult to read and lacking key information which (I feel at least) could be included in the infobox - e.g. where is that thing, what does that thing do, what makes that thing).
. The problem they point out is that articles don't have basic information, and their solution was "infobox". This is the wrong solution in my view. But again people will put time in where they want. This just seems to be a diversion of our most valuable resource to me. - Additionally,
User:Boghog I am curious why you say it would take less time and effort to add this content manually to an infobox compared to an article. Would you please explain?
Jytdog (
talk)
19:16, 17 July 2018 (UTC)
- As I have stated before, the required GO data is already stored in wikidata. We just need to manually select a subset to be highlighted. In addition, because of within family similarities of the data, processing the data one family at a time will further speed up the process. The entire process can be automatic with a manually assisted script. It is trivial to display this type of data in an infobox. It is much more difficult to automate the integration of this data into existing prose. And regardless of whether we have a understandable lead or not, having the key properties listed in the infobox will be useful in its own right.
Boghog (
talk)
21:34, 17 July 2018 (UTC)
- Oh, you didn't mean manual at all. This is much more worrisome. I would be curious to see a wikidata entry with the kind of data that you believe would be appropriate for this "basic information" field. Can you point me to one?
Jytdog (
talk)
20:07, 18 July 2018 (UTC)
- An example of the kind of manually selected data that I propose to include in the gene infobox is found in the "Key Properties" table
below. The manual part is an editor making a decision about which of the existing GO terms should be promoted and this will take a considerable amount of work. The proposed work flow is as follows:
- down load the GO data from wikidata into a spread sheet (automated)
- organize by family (automated)
- select preferred data (manual)
- adjust the wikidata ranking of the selected data (automated)
- Please note that each of the GO terms provides one or more sources and none of the GO terms is a medical claim. Many of the claims are uncontroversial and can quickly be processed (e.g., a GPCR is membrane associated and the list of endogenous ligands for a particular GPCR). Identifying the primary function may take a little more work. Boghog ( talk) 20:44, 18 July 2018 (UTC)
- Also please note that the data already is in the infobox, but in a collapsed state. The proposal is to manually select a subset for uncollapsed display.
Boghog (
talk)
04:51, 19 July 2018 (UTC)
- Hey Boghog sorry taking time to respond.
Here is the Wikidata entry on BRCA1. Where in there is the "basic information" about "where is that thing, what does that thing do, what makes that thing" in that wikidata entry? (I acknowledge tissue expression data is there in the png image, as it is the infobox)
Jytdog (
talk)
14:28, 21 July 2018 (UTC)
- Fair enough question. The relevant wikidata is here: BRCA1 DNA repair associated (Q17487737) BRCA1 is an "edge case" in that it is a multifunctional protein that has intensively studied and hence many GO terms have been assigned to it. But hey, that is biology. The "where is that thing" question has two parts. In which tissues is it expressed and which part of the cell is it located. The short answer is that it is expressed everywhere (except plasma). That is reflected in the "cellular component" GO terms: plasma membrane, cytoplasm, and cell nucleus. The GO terms generally don't distinguish between tissues. The RNA expression pattern suggests wide tissue distribution and that is also seen in BRAC1 protein expression ( human protein atlas). "What does that thing do" is function. "Biological process" GO lists "DNA repair" which I think qualifies as it primary function. As I have mentioned elsewhere "what makes that thing" is somewhat of a generic question. The short answer is where all proteins are made, ribosomes. Digging deeper, what differentiates proteins synthesis is post-translational modification and GO terms do not cover this. So in summary, the promoted GO terms would look like:
- Hey Boghog sorry taking time to respond.
Here is the Wikidata entry on BRCA1. Where in there is the "basic information" about "where is that thing, what does that thing do, what makes that thing" in that wikidata entry? (I acknowledge tissue expression data is there in the png image, as it is the infobox)
Jytdog (
talk)
14:28, 21 July 2018 (UTC)
- An example of the kind of manually selected data that I propose to include in the gene infobox is found in the "Key Properties" table
below. The manual part is an editor making a decision about which of the existing GO terms should be promoted and this will take a considerable amount of work. The proposed work flow is as follows:
- Oh, you didn't mean manual at all. This is much more worrisome. I would be curious to see a wikidata entry with the kind of data that you believe would be appropriate for this "basic information" field. Can you point me to one?
Jytdog (
talk)
20:07, 18 July 2018 (UTC)
- As I have stated before, the required GO data is already stored in wikidata. We just need to manually select a subset to be highlighted. In addition, because of within family similarities of the data, processing the data one family at a time will further speed up the process. The entire process can be automatic with a manually assisted script. It is trivial to display this type of data in an infobox. It is much more difficult to automate the integration of this data into existing prose. And regardless of whether we have a understandable lead or not, having the key properties listed in the infobox will be useful in its own right.
Boghog (
talk)
21:34, 17 July 2018 (UTC)
- The OP
started by saying
Key Properties Property Description Molecular mechanism damaged DNA binding Biological function DNA repair Subcellular location plasma membrane, cytoplasm, cell nucleus
- Support with caveats that this be very limited and written manually. Natureium ( talk) 20:09, 16 July 2018 (UTC)
6. Rename 'orthologs' to 'external links'
The orthologs part of this section is in fact just the titles 'human' and 'mouse'. The rest are just external links. The proposed change makes the purpose of this section much clearer to readers and reflects what it actually does.
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment– almost everything in the right-hand side of the infobox (grey shaded areas) is an external link so it makes no sense creating a separate external links section. The orthologs listed (human and mouse) are just example organisms, although very important ones. The present table layout makes sense (the left-hand side contains internal wikilinks that provide and explanation for the right-hand side external links). This layout is also very efficient as the headings need only be displayed only once.
Boghog (
talk)
08:53, 15 July 2018 (UTC)
- If as you say these are just external links, why does the section need to be a title that 99% of readers will not be able to understand? --
Tom (LT) (
talk)
11:29, 15 July 2018 (UTC)
- The whole infobox, not just the orthologs section, is filled with external links. Perhaps "orthologs" could be replaced with "representative species" (or "critters" ;-).
Boghog (
talk)
12:11, 15 July 2018 (UTC)
- @ Boghog Whilst I prefer "critters" as it is both highly technically accurate and also understandable by laypeople, I would also support "representative species" with a piped wikilink (like representative species).-- Tom (LT) ( talk) 09:59, 19 July 2018 (UTC)
- The whole infobox, not just the orthologs section, is filled with external links. Perhaps "orthologs" could be replaced with "representative species" (or "critters" ;-).
Boghog (
talk)
12:11, 15 July 2018 (UTC)
- If as you say these are just external links, why does the section need to be a title that 99% of readers will not be able to understand? --
Tom (LT) (
talk)
11:29, 15 July 2018 (UTC)
- Not to 'external links' Another name might be more readily understood, but there are external links throughout most of the infobox. Orthologs seems to be the most technically correct, and "critters" is just ridiculous.
Natureium (
talk)
20:09, 16 July 2018 (UTC)
- Of course "critters" is ridiculous. Turn on your humor detector. Boghog ( talk) 04:55, 19 July 2018 (UTC)
7. Merge gene location human/mouse together
Merging these two parts together reduces the amount of headings in the infobox (which are many) and hopefully also simplifies its display. The section can include a subheading for human and mouse. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Oppose – Displaying the human and mouse links side-by-side saves space since the headings (Entrez, Ensembl, UniProt, etc.) only need to be displayed once. Putting one on top of the other would require duplicating the headings.
Boghog (
talk)
22:51, 14 July 2018 (UTC)
- @ Boghog this proposal relates to merging the two sections of the infobox titled "gene location (human)" and "gene location (mouse)" into a single section titled "gene location". This proposal does not relate to the orthologs section. -- Tom (LT) ( talk) 11:28, 15 July 2018 (UTC)
- OK, sorry I misunderstood your proposal. For this merge to work, I think the graphics would need to be removed and the merged section would consist of a three column data similar to the orthlog section. We then might as well merge the gene location data to the orthlog section which already contains a partially redundant "Location (UCSC)" row. Boghog ( talk) 11:59, 15 July 2018 (UTC)
- Oppose The human chromosomal information will be relevant to many more people, and it makes sense to me to have the human section uncollapsed and the mouse section collapsed by default. Natureium ( talk) 20:09, 16 July 2018 (UTC)
8. Display only a few things in 'gene ontology', or go back to plain text
It seems like we're on the way to doing this based on WT:PHARM, but I will propose it here as a plan B. The 'gene ontology' section is not useful as is. It contains 20 - 30 text entries describing processes. None of these reflect the article's lead description of the gene, nor the commonly understood function of a gene, e.g. when it is described as encoding an enzyme, membrane receptor, tumour suppressor gene etc. I propose that this section, if it can't be made easier to read and have a more narrow focus, be removed and we just handwrite everything ourselves. Yes that takes time but as an encyclopedia we are never finished and we should be aiming for the best - and spaghetti lists are not helpful to anyone. These lists are not useful at the moment - experts will already know these facts, and lay readers will not be able to make head nor tail of it, nor know how to extract the important items in the list from the chaff. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment – Actually I think leaving this list in place in a collapsed state servers a useful propose. This way editors can easily compare the full list against the promoted list and adjust the promoted list if they don't agree with our selections. In addition, there may be "minor functions" that a individual reader might find interesting. Boghog ( talk) 22:41, 14 July 2018 (UTC)
9. Rename 'gene ontology' to 'gene function'
Renaming ontology to function makes the purpose of the section much clearer to end readers, which should be our goal here. I am not sure that readers will even look at the content of the section because they won't know what the section title means. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
Support–More understandable to the average reader. Boghog ( talk) 22:43, 14 July 2018 (UTC) On second thought, this would be misleading. The gene ontology contains more than function data, it also contain mechanism and location. In any case, if we introduced a "keys fact section" above the "gene ontology" section, the need to rename "gene ontology" becomes less urgent. Boghog ( talk) 07:15, 15 July 2018 (UTC) Labeling the section "protein properties" might work. Boghog ( talk) 10:14, 15 July 2018 (UTC)- I'd support "protein properties", which is satisfactorily descriptive, or almost any title that doesn't contain ontology, which readers will not understand. -- Tom (LT) ( talk) 11:30, 15 July 2018 (UTC)
- Oppose gene function. Ambivalent on protein properties. Natureium ( talk) 20:09, 16 July 2018 (UTC)
10. Merge the two gene location fields
Gene location is provided in three sections: gene location (human), and (mouse), and also as a field in orthologs. I propose there that the field 'Location (UCSC)' present in orthologs be moved to the appropriate gene location (human) and (mouse) sections, thus centralising this information and also reducing some duplication. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Support. -- Tom (LT) ( talk) 23:32, 13 July 2018 (UTC)
- Comment – Unless I am missing something, this proposal is essentially a duplicate of #7 above.
Boghog (
talk)
07:46, 15 July 2018 (UTC)
- Not the same proposal. Within the section called orthologs there is a field called "location (USBC)". IN addition there are two sections called "gene location". The proposal here is to remove the "location (UCSC)" field from orthologs, and move the contents to the sections titled "gene location". --
Tom (LT) (
talk)
11:27, 15 July 2018 (UTC)
- OK, now I undertand. As mentioned above, the merge could be done in the reverse direction (i.e., merge the "gene location" sections into the orthologs section).
Boghog (
talk)
12:04, 15 July 2018 (UTC)
- Boghog I'd support either way, and your proposed solution also has the benefit of removing some top level infobox headings :). -- Tom (LT) ( talk) 10:01, 19 July 2018 (UTC)
- OK, now I undertand. As mentioned above, the merge could be done in the reverse direction (i.e., merge the "gene location" sections into the orthologs section).
Boghog (
talk)
12:04, 15 July 2018 (UTC)
- Not the same proposal. Within the section called orthologs there is a field called "location (USBC)". IN addition there are two sections called "gene location". The proposal here is to remove the "location (UCSC)" field from orthologs, and move the contents to the sections titled "gene location". --
Tom (LT) (
talk)
11:27, 15 July 2018 (UTC)
11. Conservative alternative
Rather than making a large number of changes to the infobox all at once, I think a more limited number of changes could make the infobox much more readable for a general audience without sacrificing the usefulness for specialists. Hence I propose that we implement the following subset of what has been proposed above:
- Collapse Gene location (proposal 2)
- Collapse RNA expression pattern (new proposal)
- Introduce a new "key properties" section near the top (proposal 5). Using the Gene Ontology section in Beta-2 adrenergic receptor as an example, if we were to rank a subset of GO data as "preferred" for the wikidata Adrenoceptor beta 2 (Q287961) data set (see diff), the following could be displayed in the infobox:
| Property | Description |
|---|---|
| Molecular mechanism | G-protein coupled receptor activity, epinephrine binding, norepinephrine binding |
| Biological function | regulation of smooth muscle contraction |
| Subcellular location | membrane |
Of course to implement any of this, we must first get buy-in from the those that maintain the infobox and associated wikidata. For example, we have to make sure that changes to the rank of data do not get overwritten by bots. Boghog ( talk) 10:20, 15 July 2018 (UTC)
- Support as proposer. Boghog ( talk) 07:10, 15 July 2018 (UTC)
- Support would support the batch of proposals above in addition to any others suggested. I've tried to propose some piecemeal changes rather than batch them so that they may be discussed clearly and separately. -- Tom (LT) ( talk) 11:26, 15 July 2018 (UTC)
- Support, although I've long felt the RNA expression data is so small as to be useless even to most specialists (though maybe this is a quirk of how my screen is set up?). Perhaps it would be more useful to the reader to replace the bar graphs with a
|most expressed in=field that auto-filled with text of either the top few hits or all cell types that had expression over some mark (the graphs currently show 3x the median, so perhaps that's a mark of interest?). Bonus points if we could make those top hits wikilink to the relevant wikipedia articles. Of course, the section would also have an external link to the microarray dataset this comes from. As an example, I'm looking at the BRCA1 infobox which shows this file and another. Thoughts? Ajpolino ( talk) 14:45, 15 July 2018 (UTC) - As a general point, collapsing content causes accessibility problems. Some fraction of users will have problems (e.g., RSI pain) with collapsed sections. You should think about whether that's an acceptable compromise. If the material is going to be used a moderate amount, then it's not reasonable to hide it. OTOH, if the material is so unwanted that almost nobody will un-collapse it, then does it need to be in the article at all? WhatamIdoing ( talk) 01:15, 16 July 2018 (UTC)
Regulatory RNA concern
I am a new Wikipedia Fellow in the midst of training. I entered " regulatory RNA" and RNAi came up. Of course that's the best worked out regulatory RNA system today, but there are other regulatory RNAs that should be accessed by the general term, such as Xist and others involved in X inactivation and enhancer RNAs, both of which are treated elsewhere on Wikipedia. Does this seem like a problem to others? Is it worth my while to work on putting together a more general treatment of regulatory RNA while I'm in this course? Thank you for considering this. LLMHoopes ( talk) 17:34, 30 July 2018 (UTC)
- Good catch. If we have additional existing articles that fit within the area of regulatory RNAs (I am not an expert in this field), a good place to start might be to turn regulatory RNA into a disambiguation page with links to RNAi and other regulatory RNAarticles (e.g., Long noncoding RNAs, ...). The disambiguation page might later be turned into a general article about regulatory RNAs. Boghog ( talk) 18:00, 30 July 2018 (UTC)
Estriadol metabolism
Hi everyone.
An editor asked me to reproduce File:Estradiol metabolism.png as a wikitext-annotated image. Would anyone here be willing to upload an svg image file with just the 11 compounds in that figure (without any image text except for the letters for the functional groups on the compounds, e.g., the OH for the hydroxyl group) under the file name File:Estradiol metabolitsm.svg? I can't really draw chemical structure diagrams but I can easily draw the pathways, so I intend to edit the uploaded image and add the lines and compound/enzyme annotations + adjust their placement as needed when annotating the image with a template.
@
Boghog: I'm pinging you in the event you're interested in helping with this again and aren't terribly busy.
![]() Seppi333 (
Insert 2¢)
22:10, 2 July 2018 (UTC)
Seppi333 (
Insert 2¢)
22:10, 2 July 2018 (UTC)
- Hi Seppi33. No problem. I will do this, but I need a day or two. Cheers.
Boghog (
talk)
23:59, 2 July 2018 (UTC)
- Thanks! There's no rush, so take whatever time you need.
Seppi333 (
Insert 2¢)
01:49, 3 July 2018 (UTC)
- @
Seppi333: Is this
File:Estradiol metabolism structures.svg OK?
Boghog (
talk)
07:58, 3 July 2018 (UTC)
- Yep, that's exactly what I was looking for, thanks! Could I trouble you to take a minute or two to annotate
the image file on Commons (via the "Add a note" button) with the name of each compound? It'd make it easier for me to determine where to place each structure diagram when I'm cross-referencing the svg/png files while drawing the pathway. Edit: I annotated 2 that I thought were obvious, but some of the others are throwing me off due to the stylistic/representational differences. E.g., stereochemistry, rotation, etc.
Seppi333 (
Insert 2¢)
08:41, 3 July 2018 (UTC); 08:54, 3 July 2018 (UTC)
 Done
Added names of structures.
Boghog (
talk)
06:07, 4 July 2018 (UTC)
Done
Added names of structures.
Boghog (
talk)
06:07, 4 July 2018 (UTC)
- Yep, that's exactly what I was looking for, thanks! Could I trouble you to take a minute or two to annotate
the image file on Commons (via the "Add a note" button) with the name of each compound? It'd make it easier for me to determine where to place each structure diagram when I'm cross-referencing the svg/png files while drawing the pathway. Edit: I annotated 2 that I thought were obvious, but some of the others are throwing me off due to the stylistic/representational differences. E.g., stereochemistry, rotation, etc.
Seppi333 (
Insert 2¢)
08:41, 3 July 2018 (UTC); 08:54, 3 July 2018 (UTC)
- @
Seppi333: Is this
File:Estradiol metabolism structures.svg OK?
Boghog (
talk)
07:58, 3 July 2018 (UTC)
- Thanks! There's no rush, so take whatever time you need.
Seppi333 (
Insert 2¢)
01:49, 3 July 2018 (UTC)
@ Boghog: The last thing I need before beginning to annotate and uniformly align the objects in this diagram is an svg image of estrone glucuronide's structure diagram. Could you email me that or add it to File:Estradiol metabolism structures.svg? If you'd prefer the former method, my email is in the source immediately following this sentence. Seppi333 ( Insert 2¢) 11:40, 7 August 2018 (UTC)
- Thanks again!
 Seppi333 (
Insert 2¢)
12:06, 7 August 2018 (UTC)
Seppi333 (
Insert 2¢)
12:06, 7 August 2018 (UTC)
Metabolic pathways of
estradiol in humans
|
References
- Hi Seppi. Per your request, I have added estrone glucuronide to the upper righthand corner of
File:Estradiol metabolism structures.svg.
Boghog (
talk)
12:20, 7 August 2018 (UTC)
- Much appreciated. Seppi333 ( Insert 2¢) 12:21, 7 August 2018 (UTC)
- Hi Seppi. Per your request, I have added estrone glucuronide to the upper righthand corner of
File:Estradiol metabolism structures.svg.
Boghog (
talk)
12:20, 7 August 2018 (UTC)
I think I'm more-or-less done. Anyone have any feedback/suggestions? Seppi333 ( Insert 2¢) 22:06, 10 August 2018 (UTC)