| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | ← | Archive 5 | Archive 6 | Archive 7 | Archive 8 | Archive 9 | Archive 10 |
- Can this resource be used for linking on WP pages? I am asking because one of the contributors removes linking to TCDB classification on the grounds that are not entirely clear to me (i.e. like here). We need a consensus about this. Thank you. My very best wishes ( talk) 21:41, 24 April 2016 (UTC)
- WP:LINKVIO is the complaint, so it seems to me. Some articles that were copied from that group's website are up at WP:CP because that group apparently copies text from non-free third-party sources. Jo-Jo Eumerus ( talk, contributions) 22:05, 24 April 2016 (UTC)
- The diff cited is an edit by me, and yes, that is correct, that WP:LINKVIO was my reason for that edit. I also considered just deleting the citations and adding "citation needed" tags to everything, but I really do not think that there will be alternative sourcing. Additionally, I will point out that there are WP:UNDUE issues about the material I deleted. It really should not be presented by Wikipedia as though it were the established nomenclature system used in the preponderance of sources. However, I think the underlying research could quite reasonably be cited in a possible section about the molecular phylogenetics of the ion channels. -- Tryptofish ( talk) 00:37, 25 April 2016 (UTC)
- Thank you so much for raising this! The issue isn't so much about "linking" as the largescale importing of content from that database into WP. Pinging User:Transporter Guy who has been working on a project to copy lots of content from the database into WP first as member of the lab that created the database, and now as a volunteer still invested in the project; see this discussion I had with them.
- The copy/pasting has been triggering lots of copyright alerts from bots, but the database is appropriately licensed. The problem, as discussed here and here, and here, is that TCDB itself appears to contain copyright violations and those violations are being brought over into Wikipedia sometimes. Two admins, User:Opabinia regalis and User:Diannaa have raised this issue with Transporter Guy but there doesn't seem to be a change in behavior yet.
- See also Wikipedia:Articles for deletion/Voltage-gated ion channel (VIC) family which is about content imported from TCDB into a new article based on an idiosyncratic classification.
- See also edits like this and this where Tranasporter guy over-wrote the existing article with content from TCDB, which they have done several times.
- See also edits like this where Transporter guy created a new article with content copied from TCDB and then blanked and redirected an existing one; they have done this several times too.
- Some folks have appreciation for all the content creation like here and here and this.
- This seems like something the project should discuss and get consensus on before the work continues. Thanks again for raising this MVBW! I am placing notices at a few other WikiProjects that may be interested in the discussion.
Jytdog (
talk)
22:18, 24 April 2016 (UTC)
- Edits like that are fine because he moved this content to appropriate sub-pages. As about TCDB abstracts, it appears that anyone who wants to use them must check that the inserted text does not contain any copyright violations and be responsible for that. Speaking about the links, they are definitely very useful for a reader. I should think about it. My very best wishes ( talk) 22:26, 24 April 2016 (UTC)
- Yes, thanks for starting this discussion, My very best wishes. Jytdog seems to have outlined the nature of the copyright problem – the database itself appears to to be violating copyrights by copying directly from copyright-protected published articles or abstracts. Regardless of what licence the database claims to have, copyvio content from it may not be imported into Wikipedia. Because Transporter Guy has done exactly that in several articles, and continued to do so after being clearly told to stop, I have asked for a contributor copyright investigation, a full review of the copyright status all his contributions to date. Jo-Jo Eumerus is of course right to point out that we should not link in any way to pages of the database that violate other people's copyrights, though that is not the central issue here.
- All attempts to convince Transporter Guy that he should respect our guidelines for
conflict of interest editors have failed and he continues to
WP:REFSPAM links to his employer's database into the project. Voluntarily or not, that needs to stop.
Justlettersandnumbers (
talk)
23:15, 24 April 2016 (UTC)
- It's not clear to me that the effort needs to stop permanently, but yes, paused for now while the community evaluates if it should continue and if so, how. It may well be that it continues with the blessing of the community, in a different way. That could be a happy outcome here. Jytdog ( talk) 23:24, 24 April 2016 (UTC)
- Just want to add that input from folks who haven't thought about this before would be very valuable. Jytdog ( talk) 23:49, 24 April 2016 (UTC)
- Can be used with care per our guidelines. I can see that two admins raised the copyright issue and agree with everything they said here. Let's also check the existing guidelines. There are two slightly different issues. First, if anyone reuses texts from TCDB, he/she must be absolutely sure that the text does not contain any copyright violations and be responsible for that. This is nothing special and applies to any content. Second, we do have certain guidelines for linking. According to them, we should not link to any individual pages that violate copyright, and this is particularly relevant when linking to sites such as Scribd or YouTube, where due care should be taken to avoid linking to material that violates copyright. Hence, the linking to individual YouTube entries is actually allowed, but anyone who makes such link must be sure this particular YouTube entry is not a copyright violation. By the same logic, anyone who makes link to an individual TCDB abstract must be sure this particular abstract had no copyright violations. My very best wishes ( talk) 23:53, 24 April 2016 (UTC)
- One should also realize that TCDB is a major and important database. It is well known to everyone who works in the area of protein bioinformatics or membrane proteins. It is used by every other major protein database, such as Uniprot or Pfam. It implements a classification of transporters approved by the International Union of Biochemistry and Molecular Biology. It should be used here for the benefit of the project. My very best wishes ( talk) 13:53, 25 April 2016 (UTC)
- Based on your comment, I searched for the IUBMB's website listing their recommendations:
[1]. I cannot find TCDB there. --
Tryptofish (
talk)
20:43, 25 April 2016 (UTC)
- This is in every textbook on the subject. See for example, Protein Families: Relating Protein Sequence, Structure, and Function by Christine Orengo and Alex Bateman. Here it tells that "Transporter Classification Database TCDB (www.tcdb.org) (Saier et al.,2006, 2009) presents the comprehensive IUBMB (International Union of Biochemistry...) approved and adopted classification system...", and so on. This textbook dedicates an entire chapter to the TCDB. And here is large number of other books that describe TCDB. My very best wishes ( talk) 21:25, 25 April 2016 (UTC)
- See also here - Tryptofish, this appears at your link under "Membrane transport proteins". Opabinia regalis ( talk) 21:33, 25 April 2016 (UTC)
- Thanks for finding that, Opabinia. And yet, reading it, it just gets stranger and stranger. At the top it does indeed say that "Practically all the material presented here is based on the work of" the people who maintain the TCDB website. But then, look at the list itself! It is nothing like the TCDB classification as displayed at the TCDB website and edited onto Wikipedia pages! If one looks at Voltage-gated ion channel#Nomenclature, one sees a supposed VIC superfamily, with 8 subfamilies. I cannot find that on the linked page from IUBMB, where a much more conventional classification is found. But – if one clicks through from that on the 1.A links wherever 1.A occurs (alpha-helical channels, not to be confused with I.A), then there the VIC is! So in fact (way down in the "fine print"), yes the Saier group does have a TCDB listing at IUBMB, but it is not organized that way at the IUBMB website, where instead, they use much more traditional structural and functional parameters for classification. -- Tryptofish ( talk) 21:56, 25 April 2016 (UTC)
- Actually, on looking further, not even that! Here is the 1.A link: [2]. It has "VIC" as a member there, followed by other entries, but those subsequent entries are not subfamilies of VIC. They are just other kinds of alpha-helix-containing molecules, along with VIC, but not subtypes of VIC. Compare that listing and numbering scheme with what had been edited here: [3], where some of them are subtypes. The Wikipedia edits include a "K+ transporter" family that is nowhere at IUBMB, and it uses the highly unconventional "TrK" abbreviation, which is very widely known as something completely different. Not the same at all, no matter where one looks at IUBMB. -- Tryptofish ( talk) 22:15, 25 April 2016 (UTC)
- (
edit conflict) In any case, the book chapter you linked in your first diff was written by the same people who maintain the TCDB website! It's not in a textbook, but rather a compilation book made up of chapters provided by authors. And as for your Google link, I wonder if you even looked at it. The initial returns are just Wikipedia and self-referencing sources by the TCDB authors! --
Tryptofish (
talk)
21:37, 25 April 2016 (UTC)
- Then check another textbook
[4]. The differences appear because TCDB classification is constantly developing. Dr. Saier was the person who developed this widely used classification, and he continue doing this, as reflected in the more recent versions of TCDB. He is basically a
classic in this field, meaning that everyone who works with classification of proteins must be familiar with his work. Now, speaking about "copyright problems", I think they are strongly inflated, and TCDB can be actually used as
a reliable source. I saw many copyright violations in academic journals. This is happening when an author is reusing his own nearly identical texts in a number of different publications (as occasionally do TCDB authors). This is bad, but this does not invalidate any of these academic journals as RS.
My very best wishes (
talk)
22:23, 25 April 2016 (UTC)
- That's not "another" textbook, but it is a textbook. Anyway, it has a table that is just like what is at IUBMB, but not like what you purport is later hot-off-the-press work. If the more recent stuff gets taken up by
independent
secondary sources, then Wikipedia can use it, but
not before that happens, and it hasn't happened yet. And you appear to
not hear other editors when you talk about RS. Nobody is saying that copyright issues are about
WP:RS; it's about
WP:LINKVIO. --
Tryptofish (
talk)
22:34, 25 April 2016 (UTC)
- I am telling that if anyone needs to use TCDB for referencing and provides a link to TCDB in inline citation, he can do it. And remember that WP:RS is a policy, one of five pillars. It overrides guidelines, such ones for linking.
My very best wishes (
talk)
22:42, 25 April 2016 (UTC)
-
WP:C,
WP:CV. --
Tryptofish (
talk)
23:13, 25 April 2016 (UTC)
- No one suggests making copyright violations. Look, I
explained this to you already on your talk page.
My very best wishes (
talk)
23:29, 25 April 2016 (UTC)
-
WP:LINKVIO is part of
WP:C. --
Tryptofish (
talk)
23:33, 25 April 2016 (UTC)
- Let's assume that I found an example of a copyright violation in the
Journal of the American Chemical Society (authors used a couple of paragraphs of text which is a close paraphrase of something they published earlier in another journal; yes, that happens). Would it make any reference and linking to JACS from WP WP:LINKVIO? No, absolutely not. That is what I am talking about. Would it make a reference and linking to the particular article in JACS (where such violation happened) a WP:LINKVIO? I am not sure, but claiming that a person who made such ref was a malicious violator and we must check or remove all references to JACS in WP would be ridiculous.
My very best wishes (
talk)
23:50, 25 April 2016 (UTC)
- The situation here is that there is an external website where there appears to be copyright violation of material published by other persons, not simply authors repeating their own material – which is not a copyright violation anyway, just lazy writing (one does not violate one's own copyright). The copied material is apparently from other authors, not from the authors of the website. No one at Wikipedia has claimed that an editor is a malicious violator. I just reverted the links. As indicated by
WP:C. Which is a policy with legal implications. Reverting something is not calling anyone a malicious violator. On the other hand, beating a dead horse can get rather annoying. --
Tryptofish (
talk)
01:27, 26 April 2016 (UTC)
- I am happy to hear this is not a copyright violation (???) Because in this case it was a previous publication by same authors/lab [5]. Same in this case [6]. People who wrote the patent application probably borrowed text from an old review by the same group, but the review is apparently not accessible through Google search. My very best wishes ( talk) 04:10, 26 April 2016 (UTC)
- The situation here is that there is an external website where there appears to be copyright violation of material published by other persons, not simply authors repeating their own material – which is not a copyright violation anyway, just lazy writing (one does not violate one's own copyright). The copied material is apparently from other authors, not from the authors of the website. No one at Wikipedia has claimed that an editor is a malicious violator. I just reverted the links. As indicated by
WP:C. Which is a policy with legal implications. Reverting something is not calling anyone a malicious violator. On the other hand, beating a dead horse can get rather annoying. --
Tryptofish (
talk)
01:27, 26 April 2016 (UTC)
- Let's assume that I found an example of a copyright violation in the
Journal of the American Chemical Society (authors used a couple of paragraphs of text which is a close paraphrase of something they published earlier in another journal; yes, that happens). Would it make any reference and linking to JACS from WP WP:LINKVIO? No, absolutely not. That is what I am talking about. Would it make a reference and linking to the particular article in JACS (where such violation happened) a WP:LINKVIO? I am not sure, but claiming that a person who made such ref was a malicious violator and we must check or remove all references to JACS in WP would be ridiculous.
My very best wishes (
talk)
23:50, 25 April 2016 (UTC)
-
WP:LINKVIO is part of
WP:C. --
Tryptofish (
talk)
23:33, 25 April 2016 (UTC)
- No one suggests making copyright violations. Look, I
explained this to you already on your talk page.
My very best wishes (
talk)
23:29, 25 April 2016 (UTC)
-
WP:C,
WP:CV. --
Tryptofish (
talk)
23:13, 25 April 2016 (UTC)
- I am telling that if anyone needs to use TCDB for referencing and provides a link to TCDB in inline citation, he can do it. And remember that WP:RS is a policy, one of five pillars. It overrides guidelines, such ones for linking.
My very best wishes (
talk)
22:42, 25 April 2016 (UTC)
- That's not "another" textbook, but it is a textbook. Anyway, it has a table that is just like what is at IUBMB, but not like what you purport is later hot-off-the-press work. If the more recent stuff gets taken up by
independent
secondary sources, then Wikipedia can use it, but
not before that happens, and it hasn't happened yet. And you appear to
not hear other editors when you talk about RS. Nobody is saying that copyright issues are about
WP:RS; it's about
WP:LINKVIO. --
Tryptofish (
talk)
22:34, 25 April 2016 (UTC)
- Then check another textbook
[4]. The differences appear because TCDB classification is constantly developing. Dr. Saier was the person who developed this widely used classification, and he continue doing this, as reflected in the more recent versions of TCDB. He is basically a
classic in this field, meaning that everyone who works with classification of proteins must be familiar with his work. Now, speaking about "copyright problems", I think they are strongly inflated, and TCDB can be actually used as
a reliable source. I saw many copyright violations in academic journals. This is happening when an author is reusing his own nearly identical texts in a number of different publications (as occasionally do TCDB authors). This is bad, but this does not invalidate any of these academic journals as RS.
My very best wishes (
talk)
22:23, 25 April 2016 (UTC)
- Based on your comment, I searched for the IUBMB's website listing their recommendations:
[1]. I cannot find TCDB there. --
Tryptofish (
talk)
20:43, 25 April 2016 (UTC)
- P.S. Many major protein databases, such as UniProt, Pfam and Protein Data Bank, do linking to TCDB. Therefore, I do not see any reason why we can not use such links here. My very best wishes ( talk) 14:29, 26 April 2016 (UTC)
- There is certainly no problem with citing such other websites, as they are secondary sources with respect to TCDB and they presumably do not have copyright issues. So that is a good, and preferable, approach to this content. At the same time, the due weight policy requires us to present TCDB within the context of any of the other classifications present in those secondary sources, rather than presenting TCDB as the only one. -- Tryptofish ( talk) 21:34, 26 April 2016 (UTC)
- In the last phrase I am telling not about sourcing, but about links. Links can not be primary or secondary. If other widely used resources do not see any copyright problems with linking to TCDB (and believe me, those are very serious resources supported by large staff including people who are familiar with copyright), I do not see any problems with linking to TCDB here.
My very best wishes (
talk)
23:12, 26 April 2016 (UTC)
- I know that links cannot be primary or secondary in their own right, but when Wikipedia cites them as sources, then they can be primary or secondary sources. -- Tryptofish ( talk) 23:29, 27 April 2016 (UTC)
- In the last phrase I am telling not about sourcing, but about links. Links can not be primary or secondary. If other widely used resources do not see any copyright problems with linking to TCDB (and believe me, those are very serious resources supported by large staff including people who are familiar with copyright), I do not see any problems with linking to TCDB here.
My very best wishes (
talk)
23:12, 26 April 2016 (UTC)
- For that matter, there is nothing wrong with citing primary research papers from Saier et al., but again only subject to due weight. -- Tryptofish ( talk) 21:36, 26 April 2016 (UTC)
- I think that Jytdog did a very good job of explaining the various problems here, and I agree with him that we ought to try to find a happy outcome. I really am very concerned with the volume of Transporter Guy's edits, for multiple reasons. A very large number of pages have been affected, and there is a spammy quality to the edits, above and beyond the specific copyright issues. The external website seems to have been set up deliberately to promulgate the TCD (thus the licensing), but Wikipedia should not be a partner in it unless there is more in the way of acceptance by independent secondary sources first. I have a feeling that Transporter Guy has been put in a tough position by supervisors who want him to use Wikipedia for purposes that contradict our norms. There really is also a WP:UNDUE problem with much of this content. When one gets into the weeds of the source material, the database is really about genetic homology, but not about functional relationships. As phylogenetic information, that's fine. But ion channels should not be lumped together with ion transporters, for example. I'm not surprised that some editors have thanked Transporter Guy for the content, because, like the database website, it is mostly well-written. But that is all the more reason for editors to be concerned that we must not mislead our readers. I think that there is an awful lot of material that will have to be reverted or deleted, and I really hope that Transporter Guy will be able to just slow down while this gets sorted out. -- Tryptofish ( talk) 00:57, 25 April 2016 (UTC)
- I want to follow up on a point I made briefly above. There is a very widely recognized use of the abbreviation "Trk" to designate
Trk receptors and the like. Here is a PubMed search for "Trk", and it confirms what I just said:
[7]. The Saier lab TCDB website uses the abbreviation instead to designate K+ transporters. I don't see that at the IUBMB website, and I don't think I've seen it anywhere else. If we posit that the TCDB site reflects something more recent than what is at IUBMB and elsewhere, then I really have to question the degree to which we can regard the TCDB site as being peer-reviewed. Taking that observation along with the copying issues, I get the feeling that the TCDB site is probably being edited by junior people in the lab group, and is a sort of continuous work-in-progress (not unlike Wikipedia in that regard). It's not really a "final draft". Given that, and given
WP:CRYSTAL, I think that makes it a less reliable source than are sites such as IUBMB, and makes direct copying from TCDB to Wikipedia undesirable. --
Tryptofish (
talk)
22:24, 26 April 2016 (UTC)
- This isn't totally off-base - they seem to have just picked an exemplar of the family to name it after, and they picked TrkH. For whatever reason tropomyosin receptor kinases and one of the more common bacterial potassium uptake systems inconveniently converged on the same abbreviation. (Though I think the eukaryotes call theirs "track" and the prokaryotes "terk".) Opabinia regalis ( talk) 23:09, 26 April 2016 (UTC)
- Yes, it's just significantly off-base, but not totally (and it's tyrosine rather than tropomyosin). As long as they don't call it "twerk".
 But, seriously, does anyone else call the potassium transporters that? --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
But, seriously, does anyone else call the potassium transporters that? --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
- Yes, it's just significantly off-base, but not totally (and it's tyrosine rather than tropomyosin). As long as they don't call it "twerk".
- This TCDB abbreviation does not matter at all because the family appears as "The K+ Transporter (Trk) Family" in the database. There is no way anyone would think they are tyrosine kinases. Same with names used on WP pages. This is not a nomencalture of human genes or something else based on unique identifiers.
My very best wishes (
talk)
23:43, 26 April 2016 (UTC)
- Actually your argument does not matter at all, because an abbreviation is only useful in contexts where it does not appear with the full name. In any case, I think that it has become very clear that the TCDB website is less of a reliable source for Wikipedia's purposes than are either peer-reviewed papers or the websites that secondarily report on TCDB, and it is also clear that TCDB is one of multiple nomenclature systems, rather than the only one. --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
- Obviously, any combination of three letters has multiple meanings, as reflected on the corresponding disambig. page,
TRK. There is nothing else here, really.
My very best wishes (
talk)
17:02, 28 April 2016 (UTC)
- Gene sequence: Truckee Tahoe Airport. Anyway – I think that it has become very clear that the TCDB website is less of a reliable source for Wikipedia's purposes than are either peer-reviewed papers or the websites that secondarily report on TCDB, and it is also clear that TCDB is one of multiple nomenclature systems, rather than the only one. -- Tryptofish ( talk) 00:48, 29 April 2016 (UTC)
- Obviously, any combination of three letters has multiple meanings, as reflected on the corresponding disambig. page,
TRK. There is nothing else here, really.
My very best wishes (
talk)
17:02, 28 April 2016 (UTC)
- Actually your argument does not matter at all, because an abbreviation is only useful in contexts where it does not appear with the full name. In any case, I think that it has become very clear that the TCDB website is less of a reliable source for Wikipedia's purposes than are either peer-reviewed papers or the websites that secondarily report on TCDB, and it is also clear that TCDB is one of multiple nomenclature systems, rather than the only one. --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
- People criticized contributions by Transporter Guy. After looking at this, I think he actually did very good work. Yes, there were certain copyright problems, but they are relatively minor, easily fixable and I think already fixed. Yes, there is some amount of over-linking and POV (preferential use of only one resource this user is most familiar with), but that can also be easily fixed if anyone cares. I do not. The resource he used (TCDB) is an excellent resource, and its use is highly beneficial for the project. I am telling that as someone with an expertise in this area. My very best wishes ( talk) 12:12, 29 April 2016 (UTC)
- I also checked editing by user Tryptofish on these pages and think he is occasionally engaged in WP:OR by claiming, for example, that KcsA potassium channel does not belong to the family of voltage-gated channels [8]. It is a page about specific protein family known as "voltage-gated channels" rather than about potential-sensitive proteins in general, which would be an entirely different subject. My very best wishes ( talk) 03:54, 4 May 2016 (UTC)
- I take very strong issue with the accusation that I have been violating NOR. In fact, there is an extended discussion about the proper definition of voltage-gated channels at
Talk:Voltage-gated ion channel, and I would very much welcome input from more editors there. (I'm not making anything up, but explicitly citing reliable sources such as a widely respected book by
Bertil Hille, and I understand that ion channels have to be voltage-gated in order to be voltage-gated ion channels. The comment above misrepresents what I said, by making it sound like I want to include other, weakly potential-sensitive, channels, which is the opposite of my position. On the other hand, the other editor wants to include channels that are not voltage-gated on the page about "voltage-gated channels".) --
Tryptofish (
talk)
18:22, 4 May 2016 (UTC)
- Some members of
protein family of "voltage-gated channels" (the subject of the page) are not voltage-gated because they lost or never had function typical for other members of the family. Same goes for other protein families. For example, some members of
G protein–coupled receptor family are not G protein coupled. @Boghog. Is not that correct?
My very best wishes (
talk)
20:08, 4 May 2016 (UTC)
- These exceptions may exist, but why on earth put them in the lead? Concentrate on why the family was defined in the first place. These definitions were originally based on function and later structure. Not the other way around.
Boghog (
talk)
20:18, 4 May 2016 (UTC)
- I definitely agree. That sounds reasonable. Sure. Except that function does not serve to "define" any protein families (there are numerous proteins with the same function which belong to different protein families and completely unrelated). What defines the family is evolutionary relatedness of the proteins. This is not much different from biological taxonomy: it is the common origin what defines the existence of "natural" (as opposed to the "artificial") taxonomic groups. My very best wishes ( talk) 20:20, 4 May 2016 (UTC)
- (
edit conflict) You continue to confound phylogenetics with the actual classification according to the preponderance of reliable sources. It's time to let other editors weigh in. --
Tryptofish (
talk)
20:26, 4 May 2016 (UTC)
- What I am talking about is the "actual", commonly accepted classification of proteins. So far you did not indicated any references or links to alternative (function-based) classifications of proteins.
Here is one of them, but it also places KcsA in the same family.
My very best wishes (
talk)
20:38, 4 May 2016 (UTC)
- Not true, and not true that I haven't provided references. Please, other editors: more eyes are needed at
Talk:Voltage-gated ion channel. --
Tryptofish (
talk)
21:08, 4 May 2016 (UTC)
- If this is not true, please explain what alternative general classifications of proteins are you talking about (authors and references/links)? I am talking about
UniProt (
here is KcsA, for example, and it belongs to the
family of potassium channels, most of which are voltage-gated),
Pfam (KcsA belongs to
this family which belongs to
this superfamily) and about
TCDB.
My very best wishes (
talk)
21:21, 4 May 2016 (UTC)
- You have asked me that same question multiple times before, and I have replied to it multiple times before. At this point, what we need are more editors to comment. --
Tryptofish (
talk)
22:15, 4 May 2016 (UTC)
- You responded by providing a reference that supports what I am telling: KcsA belongs to the family of voltage-gated channels [9]. My very best wishes ( talk) 17:06, 5 May 2016 (UTC)
- You have asked me that same question multiple times before, and I have replied to it multiple times before. At this point, what we need are more editors to comment. --
Tryptofish (
talk)
22:15, 4 May 2016 (UTC)
- If this is not true, please explain what alternative general classifications of proteins are you talking about (authors and references/links)? I am talking about
UniProt (
here is KcsA, for example, and it belongs to the
family of potassium channels, most of which are voltage-gated),
Pfam (KcsA belongs to
this family which belongs to
this superfamily) and about
TCDB.
My very best wishes (
talk)
21:21, 4 May 2016 (UTC)
- Not true, and not true that I haven't provided references. Please, other editors: more eyes are needed at
Talk:Voltage-gated ion channel. --
Tryptofish (
talk)
21:08, 4 May 2016 (UTC)
- What I am talking about is the "actual", commonly accepted classification of proteins. So far you did not indicated any references or links to alternative (function-based) classifications of proteins.
Here is one of them, but it also places KcsA in the same family.
My very best wishes (
talk)
20:38, 4 May 2016 (UTC)
- These exceptions may exist, but why on earth put them in the lead? Concentrate on why the family was defined in the first place. These definitions were originally based on function and later structure. Not the other way around.
Boghog (
talk)
20:18, 4 May 2016 (UTC)
- Some members of
protein family of "voltage-gated channels" (the subject of the page) are not voltage-gated because they lost or never had function typical for other members of the family. Same goes for other protein families. For example, some members of
G protein–coupled receptor family are not G protein coupled. @Boghog. Is not that correct?
My very best wishes (
talk)
20:08, 4 May 2016 (UTC)
- I take very strong issue with the accusation that I have been violating NOR. In fact, there is an extended discussion about the proper definition of voltage-gated channels at
Talk:Voltage-gated ion channel, and I would very much welcome input from more editors there. (I'm not making anything up, but explicitly citing reliable sources such as a widely respected book by
Bertil Hille, and I understand that ion channels have to be voltage-gated in order to be voltage-gated ion channels. The comment above misrepresents what I said, by making it sound like I want to include other, weakly potential-sensitive, channels, which is the opposite of my position. On the other hand, the other editor wants to include channels that are not voltage-gated on the page about "voltage-gated channels".) --
Tryptofish (
talk)
18:22, 4 May 2016 (UTC)
To state that KcsA belongs to the family of voltage gated channels is very misleading since it implies that KcsA is also voltage gated. This IMHO is unacceptable. It is acceptable to state that KcsA is evolutionarily related to voltage gated ion channels, but is not itself voltage gated. A fundamental flaw in the TCDB is that it does not clearly distinguish between structure and function. It is naming evolutionary related protein families after the predominate function of the majority of family members and this causes confusion when referring to members of this family that don't share this function. It would have been better to use function neutral names for these families, for example, seven-transmembrane domain family instead of GPCR. Boghog ( talk) 17:47, 5 May 2016 (UTC)
- All existing and widely accepted protein classifications ( UniProt, Pfam, InterPro, TCDB and others) tell that KcsA belongs to the family of voltage-gated channels (see links above). Please show me classifications that tell it does not belong to the family of voltage-gated channels. You may not like naming of protein families in the literature and classifications, but we should follow WP:Common name. And no, this naming is not a flaw of TCDB. It simply uses common name of the family from the literature. As I already explained, the naming of protein families is chosen to reflect most common/typical function of proteins in the family, but not all of them. Look, this is something obvious. What's the problem? My very best wishes ( talk) 18:26, 5 May 2016 (UTC)
- As I have already stated, if one writes that KcsA is a member of the voltage gated ion channel family without further explanation, this is seriously misleading.
Boghog (
talk)
19:00, 5 May 2016 (UTC)
- Boghog, thank you very much for supporting what I have been saying. Editors may find it useful to look at the diff that My very best wishes provided in replying to me ( [10]), in which I demonstrate the exact opposite of what My very best wishes claims. My very best wishes appears to be insisting that the only reliable sources are these non-peer-reviewed websites that classify proteins based on sequence and phylogeny, which leads to a predetermined outcome – and, when presented with reliable sources that say otherwise, My very best wishes pretends that they say what they do not say. We are unfortunately getting into WP:IDHT territory. -- Tryptofish ( talk) 21:25, 5 May 2016 (UTC)
- Boghog, thank you for supporting what I have been saying. My very best wishes ( talk) 21:36, 5 May 2016 (UTC)
- As I have already stated, if one writes that KcsA is a member of the voltage gated ion channel family without further explanation, this is seriously misleading.
Boghog (
talk)
19:00, 5 May 2016 (UTC)
- I think some people criticize TCDB simply because they are not familiar with the subject. Contrary to the claim by Tryptofish just above ("non-peer-reviewed websites"), not only these classification systems/resources are extensively peer-reviewed and republished many times, as anyone can check on WP pages about them, but they are highly cited in many hundreds, possibly thousands publications and used by many thousands researchers. In addition, repeatedly placing this tag into page indicates that the user apparently believes this is classification according only to TCDB. This is not the case. If someone looks at these proteins in UniProt, this will be essentially the same or very similar classification. As I already noted, classification on the family level in TCDB and all other systems is essentially the same. Yes, a number of superfamilies provided by TCDB can not be found in other systems, however they are usually justified based on publications by various research groups. In other cases, such as voltage-gated superfamily, they are practically the same. It also seems that Tryptofish criticizes the commonly accepted biological approach, according to which proteins (and other relevant biological entities, such as species) must be classified based on their phylogeny (i.e. common biological origin), which is precisely the kind of original research I am talking about. My very best wishes ( talk) 11:37, 6 May 2016 (UTC)
GPCRs as transporters???
@ Transporter Guy and My very best wishes: I find this very disturbing. This is a clear example of ref spam and undue weight. GPCRs are not transporters. Full stop. GPCRs transduce signals, not transport molecules. Any classification system that categorizes GPCRs as transporters is highly suspect. The TCDB seems to be a made up ad hoc classification system. I agree with Tryptofish that the TCDB is only one of several transporter classification systems. I also agree with Tryptofish there are significant copyright concerns with Transporter Guy's contributions. Boghog ( talk) 20:00, 4 May 2016 (UTC)
- No one tells that GPCR are transporters. This edit does not assume that GPCR are transporters. What other several transporter systems are you talking about? My very best wishes ( talk) 12:49, 5 May 2016 (UTC)
- TCDB is only called "transporter database". It includes a number of proteins that are not transporters. That's OK. This is only a name. The only relevant question in the edit by Transporter Guy above if it introduces correct linking to biological databases. If the linking is not correct, let's remove of fix it. If it was correct, the edit is fine. As far as I can see, that was correct linking. At the same time, I do agree that a number of links to TCDB by Transporter Guy was indeed excessive (as I already said above). No objection to removing some of these links. My very best wishes ( talk) 20:17, 4 May 2016 (UTC)
- (
edit conflict) Thank you, Boghog. Of course you are correct, that GPCRs are not transporter molecules, even though it is entirely valid to say that there are sequence homologies that reflect phylogenetic relationships. It seems that I am unable to convince My very best wishes of this, but ion channels are not transporters, and transporters are not ion channels. And ion channels that are not voltage-gated are not voltage-gated ion channels. There are indeed problems with ref spam and with undue weight, and there are also problems with what is becoming an increasingly tendentious unwillingness to drop the stick. --
Tryptofish (
talk)
20:26, 4 May 2016 (UTC)
- One of the reasons why TCDB includes GPCR is obvious: the family of microbial rhodopsins, which is closely related to GPCR (same superfamily), contains a number of proton and ion pumps, i.e. transporters. Ones again, proteins from the same family or the same superfamily frequently have very different biological functions. This is something well known. On the other hand, totally unrelated proteins may have the same biological function, even catalyze exactly the same chemical reaction [11]. My very best wishes ( talk) 01:41, 5 May 2016 (UTC)
AfC help?
Hi. There's a draft over at AfC that it would be a good thing if someone with a bit more knowledge of the subject took a look at: Draft:Chloroplast migration. Any comments/suggestions would be greatly appreciated. Onel5969 TT me 13:30, 7 May 2016 (UTC)
Gene Wiki data migration
@ Andrew Su, I9606, and Julialturner: I am posting here because this issue may be of general interest to the MCB community.
Previously in Gene Wiki articles, only a single accession code was generally displayed for UniProt (reviewed Swiss-Prot only) and RefSeq (experimental NM/NP only) entries. I noticed that the Gene Wiki Infoboxes now display additional UniProt TrEMBL entries as well as predicted XM/XP RefSeq entries (see for example, ETHE1). My question is whether multiple UniProt and RefSeq links should be displayed. My own opinion is that only the Swiss-Prot UniProt and experimental RefSeq links should be displayed if they exist because they are the highest quality and determining which of the several displayed links is the most relevant is not immediately apparent. If Swiss-Prot or RefSeq NM/NP entries do not exist, then it may be appropriate to display TrEMBL or XM/XP links. However I would be interested in hearing how others think, especially those maintaining this data. Cheers. Boghog ( talk) 10:19, 16 April 2016 (UTC)
- @ Boghog, Andrew Su, and Julialturner: We often see issues where there are multiple identifiers that seems equally relevant and, for the moment, went with the strategy of letting the user decide. If you have a set of logic that you think should be followed, it would be pretty easy to adapt the Lua script I think. Julia's talk page has a few examples together to make it easier to see. There, you can see cases where we have multiple NM prefixed RefSeq records for the same gene. I see 2 UniProt records for VIPR1 there, with one being C prefixed and one P prefixed. I guess we should only be showing the P uniprots as these seem to match up with SwissProt records ? As for maintaining the data, I think it makes sense to keep all potentially relevant data accessible in wikidata and to let the application (e.g. the infobox lua script) encode the logic to decide what it wants to display. -- Benjamin Good ( talk) 19:12, 18 April 2016 (UTC)
- @
I9606: Thanks Benjamin. Concerning RefSeq codes, the difference between experimental and predicted entries is very clear. The experimental entries have a NM/NP prefix while the predicted entries have XM/XP prefixes. I am having a difficult time tracking down a definitive definition of the UniProt accession codes. The closest that I can find is
here. It appears that the reviewed Swiss-Prot accession numbers start with the letters [O,P,Q] whereas the [A-N,R-Z] denote a unreviewed TrEMBL entry. Hence my suggestion is
- RefSeq – only display NM/NP prefixed entries if they exist, if not, then display the XM/XP entries
- UniProt – only display [O,P,Q] prefixed entries if they exist, if not, then display the [A-N,R-Z] entries
- Does this make sense? Boghog ( talk) 19:40, 18 April 2016 (UTC)
- yes, +1 --
Benjamin Good (
talk)
21:01, 18 April 2016 (UTC)
- @
I9606:@
Boghog: I should be able to implement these changes pretty easily. I could also display all the RefSeq and Uniprot, but sort so the most relevent data is at the top of the list.
Julialturner (
talk)
20:48, 18 April 2016 (UTC)
- I think selectively showing the most relevant is the way to go here. There is the potential to generate pretty long, uninformative lists if we show them all - unwieldy even if sorted. --
Benjamin Good (
talk)
21:01, 18 April 2016 (UTC)
- @
Boghog,
Andrew Su, and
I9606: The changes discussed above should now be reflected in those gene pages with the infobox_gene template. Only the most relevant items are displayed on the above criteria.
Julialturner (
talk)
18:19, 12 May 2016 (UTC)
- Thanks Julialturner! That is a big improvement. Boghog ( talk) 11:05, 14 May 2016 (UTC)
- @
Boghog,
Andrew Su, and
I9606: The changes discussed above should now be reflected in those gene pages with the infobox_gene template. Only the most relevant items are displayed on the above criteria.
Julialturner (
talk)
18:19, 12 May 2016 (UTC)
- I think selectively showing the most relevant is the way to go here. There is the potential to generate pretty long, uninformative lists if we show them all - unwieldy even if sorted. --
Benjamin Good (
talk)
21:01, 18 April 2016 (UTC)
- @
I9606:@
Boghog: I should be able to implement these changes pretty easily. I could also display all the RefSeq and Uniprot, but sort so the most relevent data is at the top of the list.
Julialturner (
talk)
20:48, 18 April 2016 (UTC)
- @
I9606: Thanks Benjamin. Concerning RefSeq codes, the difference between experimental and predicted entries is very clear. The experimental entries have a NM/NP prefix while the predicted entries have XM/XP prefixes. I am having a difficult time tracking down a definitive definition of the UniProt accession codes. The closest that I can find is
here. It appears that the reviewed Swiss-Prot accession numbers start with the letters [O,P,Q] whereas the [A-N,R-Z] denote a unreviewed TrEMBL entry. Hence my suggestion is
Bot implementing wiki-data driven infobox for genes
See this diff which was made by a bot, User:ProteinBoxBot. Just wanted to make sure you all were aware of this. Jytdog ( talk) 03:50, 20 May 2016 (UTC)
The article states that this deletion syndrome is a protein, which strikes me as odd. Also, there is already an article on the Chromosome 22q11.2 deletion syndrome, called DiGeorge syndrome. -- Arno Matthias ( talk) 17:46, 25 May 2016 (UTC)
- There are a fair amount of genes, loci and proteins that are named after a disease condition. Seems like this is the case here. Jo-Jo Eumerus ( talk, contributions) 18:48, 25 May 2016 (UTC)
- Thanks for the heads up. It is true that many gene/proteins are name after diseases. However in this particular case, DEL22Q11.2 is listed as a gene with unknown type. Hence it apparently is not known if this gene is coding or not. Therefore I have modified the text accordingly. Boghog ( talk) 19:00, 25 May 2016 (UTC)
- I agree with Boghog; it is an ill-understood genetic disorder, and after a brief web search I found no product coded. Cheers, BatteryIncluded ( talk) 19:34, 25 May 2016 (UTC)
Thank you all, but shouldn't there at least be a link to DiGeorge syndrome in the article? Or, if the are the same, only one article? -- Arno Matthias ( talk) 06:44, 26 May 2016 (UTC)
- Yes. I missed that part. The chromosome 22q11.2 deletion syndrome is the same as or causes the DiGeorge syndrome, so they should be merged. I'll go ahead. Cheers, BatteryIncluded ( talk) 13:45, 26 May 2016 (UTC)
(Also posted on project PHARMA)
User:Medgirl131 has recently created {{ Cytokine receptor modulators}} and {{ Nitric oxide signaling}} navboxes and has been adding these to a large number of articles. My major concern is that they combine endogenous proteins and recombinant protein drugs in one infobox which I think is confusing. Additional concerns that I have about these navboxes are their large size and that they violate WP:Bidirectional which makes navigation more difficult. I have highlighted my concerns on their talk page here. What do others think about these navboxes? Boghog ( talk) 06:03, 28 May 2016 (UTC)
Auto-assessment of article classes
Following a recent discussion at WP:VPR, there is consensus for an opt-in bot task that automatically assesses the class of articles based on classes listed for other project templates on the same page. In other words, if WikiProject A has evaluated an article to be C-class and WikiProject B hasn't evaluated the article at all, such a bot task would automatically evaluate the article as C-class for WikiProject B.
If you think auto-assessment might benefit this project, consider discussing it with other members here. For more information or to request an auto-assessment run, please visit User:BU RoBOT/autoassess. This is a one-time message to alert projects with over 1,000 unassessed articles to this possibility. ~ Rob Talk 01:16, 4 June 2016 (UTC)
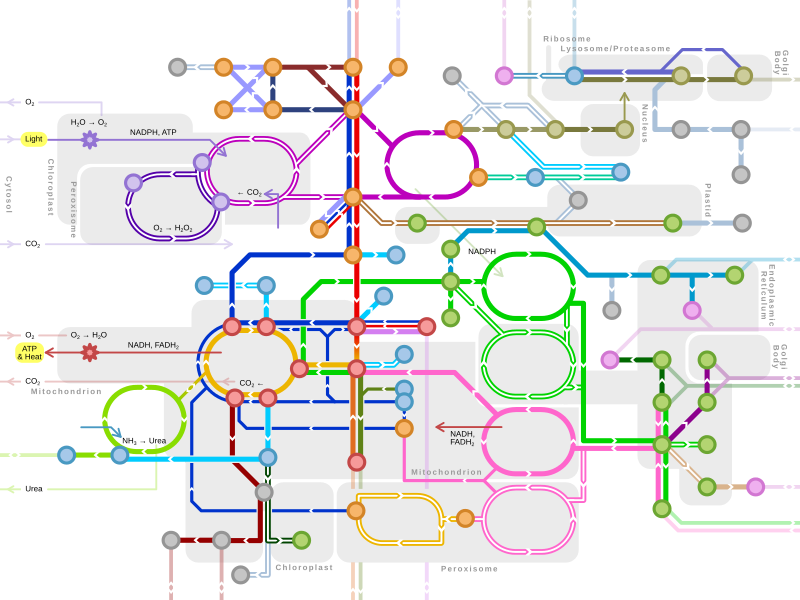
New map of Metabolic Pathways


I've made a new metro-style map of the major metabolic pathways, which was inspired by the already existing one ( Template:Metabolic_pathways) and other excellent online resources (e.g. [12], [13]). The map encompasses most of the biochem pathways in human, but also include important ones in plants / fungi / microbes (double lines). The metabolites are grouped as general terms (e.g. "Hexose Phosphates") instead of the specific ones in the existing map. I'm also going to make it a clickable map.
Some questions:
- Is it suitable for wikipedia, in terms of style and complexity?
- Is it good to replace the existing map with this one (that would change all the pages that includes Template:Metabolic_pathways) or make it a separate work?
- Any comment on the map as an individual work?
Thanks! (also posted at Wikipedia:WikiProject_Molecular_and_Cell_Biology/Metabolic_Pathways_task_force/proposals#New_map_of_Metabolic_Pathways) Chakazul ( talk) 17:30, 30 May 2016 (UTC)
- Great work
Chakazul! I'm a big fan of clear diagrams. I would say that it's an improvement over the image currently used in
Template:Metabolic_pathways. If you're interested, I'd be happy to help you out with
{{ annotated image 4}}to convert it into an interactive diagram (like Template:Glycolysis summary). I have a few suggestions for improvements:- It doesn't really need a title within the image itself (since it'll have a caption etc)
- The legend seems to be spread all around the sides currently. I realise it was for aesthetic reasons to best use the available space, however I think it aids readability to have it all together somewhere.
- Do the node sizes have any significance? How about the overlapping nodes? Again, either of these may be useful to mention in the legend
- Overall, a great diagram, and I'm sure it'll find wide use around Wikipedia. T.Shafee(Evo﹠Evo) talk 00:57, 31 May 2016 (UTC)
- Great work
Chakazul! I'm a big fan of clear diagrams. I would say that it's an improvement over the image currently used in
Template:Metabolic_pathways. If you're interested, I'd be happy to help you out with
- Thanks
Evolution and evolvability! It would be great if you can help to make it interactive. There seems no way to extract the bounding box information form SVG, so need to set the clickable areas one-by-one?
- Let me try make a version with one single legend (I was trying to avoid the use of legend, though)
- Smaller node means one kind of metabolite (e.g. Acetyl-CoA) and a larger one means a group (e.g. Amino Acids). Double nodes allow both of the terms like "Glycoproteins & Proteoglycans" clickable, clicking the left node will link to Glycoprotein and clicking the right one will link to Proteoglycan.
- Chakazul ( talk) 10:27, 31 May 2016 (UTC)
- Thanks
Evolution and evolvability! It would be great if you can help to make it interactive. There seems no way to extract the bounding box information form SVG, so need to set the clickable areas one-by-one?
- @
Chakazul: You could do a version with no legend at all in the image (I actually didn't put any in
Template:Glycolysis summary), and just write a text caption. The
{{ annotated image 4}}template only allows adding wikilinked text over the image. Adding click-able field shapes / image maps tends to be trickier. T.Shafee(Evo﹠Evo) talk 10:38, 31 May 2016 (UTC)
- @
Chakazul: You could do a version with no legend at all in the image (I actually didn't put any in
Template:Glycolysis summary), and just write a text caption. The
- @ Evolution and evolvability: There's a no-legend version above. Yes, image map is troublesome, links overlay would be more suitable in wikipedia. I wonder if the links can be transparent, otherwise I need to remove all the texts. Chakazul ( talk) 18:00, 31 May 2016 (UTC)
- @ Chakazul: The no-legend version look good! I'm afraid you're correct that you'll have to make a no-text version. It's a bother. It's possible to make the wikilink text any colour, but not transparent (as far as I know). I recommend making a no-text version only once you're certain that you're happy with the layout etc sp that you only have to do it once. On that note, it might be worth drafting the caption now. I think it'd be best to make a separate new template rather than replace the image in Template:Metabolic_pathways, so that the older image is kept as an archive. This page lists every article that transcludes Metabolic_pathways and can have the new "Template:metabolic metro" added (or whatever you decide to call it). T.Shafee(Evo﹠Evo) talk 00:00, 1 June 2016 (UTC)
- There actually is a way to create a transparent wikilink/image map-type hyperlinks using
{{ Annotated image 4}}, but it'd probably be a little more tedious to set up relative to annotating text onto an image without pre-existing text. Basically, all you have to do is create a piped wikilink to a bunch of non-breaking spaces (produced via the text ). Adding a space between non-breaking spaces will take up less room in the source code (i.e., use instead of ). As an example, the next line contains an invisible link to the MCB project page and the line after that contains an invisible link to a redirect to this page:
← link is here
← link is here
When you mouse over it, you should see the blue underlining from the HTML link. With the annotated image template, you'd need to adjust the number of non-breaking spaces so that it spans the full length of the word(s) that you want to link to. That may require some trial and error with the number of non-breaking spaces for each term that you want to link to get the length right. Seppi333 ( Insert 2¢) 03:35, 1 June 2016 (UTC)
Edit: I just noticed that the diagram has line breaks in the image text. If invisible wikilinks are annotated onto the diagram, some of the annotated wikilinks would need to have<br />included in the piped link when the text in the diagram is on different lines.
For example,[[wikitext|this wikilink<br />spans several lines via 2 forced<br />line breaks.]]produces: this wikilink
spans several lines via 2 forced
line breaks.
It works the same way if you replace the letters with non-breaking spaces. Seppi333 ( Insert 2¢) 04:07, 1 June 2016 (UTC)
- There actually is a way to create a transparent wikilink/image map-type hyperlinks using
@ Evolution and evolvability and Seppi333: Thanks for the suggestions! Here is a trial version of the template: User:Chakazul/sandbox.
The first one is the diagram without text, added two wikilinks for testing; the second one is the original diagram, using Seppi's method. Both methods have their own merits. Style 1 is a bit easier to match the positions, and can show the availability of the wiki page (blue or red). Style 2 gives more control on the color (most texts are in black BTW), but matching the white spaces needs more work. So Evo&Evo it's up to you to choose which method for annotation. Chakazul ( talk) 18:56, 1 June 2016 (UTC)
Beautiful! A few suggestions:
- The grey boxes are invisible when I face straight towards my monitor. I can only see them if I look from an angle. Maybe make them a bit darker, or put a thin black line around them.
- I find the cellular respiration colour difficult to distinguish from the carbohydrate metabolism and messengers colours. (I can distinguish those two, though.)
- Given that the "feeders to gluconeogenesis" tree is large and convoluted, it might be helpful to dedicate dark blue solely to gluconeogenesis and its feeders, and use a different colour(s) for glycogenesis etc.
- In a few places you've written NADH2 instead of NADH, or NADPH2 instead of NADPH.
- Maybe move MVA to the left a bit to separate it from NADPH from the PPP.
- Maybe write "Energy Metabolism" in black or indicate it with a box or bracket or something, because I'd been looking for arrows of the corresponding colour in the map.
- I'm not sure why there's a red line between pyruvate and lactate, as this reaction is not part of glycolysis.
Awesome work!
![]() Adrian J. Hunter(
talk•
contribs)
14:08, 31 May 2016 (UTC)
Adrian J. Hunter(
talk•
contribs)
14:08, 31 May 2016 (UTC)
- @
Adrian J. Hunter: Thanks for the comments! New version is updated.
- I decided to make glycogenesis / glycogenolysis darker colors. The blue and red lines from lactate to pyruvate (and also those from glycerol to triose phosphates i.e. DHAP & GAP) are technically side ways entering gluconeogenesis and glycolysis, so I make them thin lines much like the "feeders".
- The NADH2 (also NADPH2) were meant to be NADH2+ as sometimes seen on other diagrams as a shorthand of NADH+H+, but that would be confusing, I changed them to NADH & NADPH.
- Chakazul ( talk) 18:00, 31 May 2016 (UTC)
- An additional link that I find helpful is colorbrewer2.org. If you select "qualitative" as your colourscheme, it will give you a set of colours (up to 12) that are the maximally easy to tell apart. I've started to use it since I'm colour-blind so often pick odd colour palettes otherwise! T.Shafee(Evo﹠Evo) talk 08:24, 1 June 2016 (UTC)
- @
Adrian J. Hunter: The color scheme link is very useful! Too bad that they don't have colorblind safe scheme with more than 4 colors. A few of my friends are colorblind (I already guessed you're one) so I often have colorblind-friendliness in mind, but I always wanted to make my diagrams colorful and that became a dilemma.
Chakazul (
talk)
18:56, 1 June 2016 (UTC)
- I think this comment was meant for
Evolution and evolvability :-). Changes look good to me.
Adrian J. Hunter(
talk•
contribs)
12:03, 2 June 2016 (UTC)
- Yes I meant Evo&evo. Chakazul ( talk) 17:15, 2 June 2016 (UTC)
- I think this comment was meant for
Evolution and evolvability :-). Changes look good to me.
Adrian J. Hunter(
talk•
contribs)
12:03, 2 June 2016 (UTC)
- @
Adrian J. Hunter: The color scheme link is very useful! Too bad that they don't have colorblind safe scheme with more than 4 colors. A few of my friends are colorblind (I already guessed you're one) so I often have colorblind-friendliness in mind, but I always wanted to make my diagrams colorful and that became a dilemma.
Chakazul (
talk)
18:56, 1 June 2016 (UTC)
Template:Metabolic_metro now created. My sandbox can be used for drafting. @ Evolution and evolvability:. Chakazul ( talk) 17:15, 2 June 2016 (UTC)
- @
Chakazul: The template's looking great! It would be great to see sub-section versions in the same style of it for some of the related pages.
T.Shafee(Evo﹠Evo)
talk
05:31, 11 June 2016 (UTC)
- @ Evolution and evolvability: Thanks for the color dots. I've corrected the color of blue dot from violet to blue (they looks similar?). Now see if it's suitable to use this template in other pages.
- I do think of making detailed version for individual pathways, or other language versions. Chakazul ( talk) 08:25, 11 June 2016 (UTC)
| Major metabolic pathways | |
|---|---|
|
Flow-through test - article up for speedy deletion
Esteemed members of this project may be best placed to work out whether the speedy deletion tag on Flow-through test should stay or go ... the subject matter appears to be within your sphere of interest. Grateful if someone would take a look (preferably before it gets speedied) - it's an article which popped up on the Wikipedia:Conflict of interest/Noticeboard. thanks -- Tagishsimon (talk) 00:53, 30 June 2016 (UTC)
Conversation on controversial changes to epistasis article being discussed on the WP:GEN talkpage. T.Shafee(Evo﹠Evo) talk 00:28, 4 July 2016 (UTC)
Can someone with some experience help double-check on FAM163A? The title is FAM163A but the infobox is FAM136A. It seemed like it was previously clear with this version but this edit flipped the name but not the infobox and changed none of the citations which is mighty odd. Does a history split need to be done and other fixes? See also User:Flameep/sandbox. -- Ricky81682 ( talk) 03:12, 14 July 2016 (UTC)
- I think this is OK. The addition of the FAM136A infobox to FAM163A was a mistake that was later corrected in this . FAM163A also contained some original research which I have removed. Boghog ( talk) 06:13, 14 July 2016 (UTC)
WikiFactMine: Proposed Wiki Grant for biological Wikidata
We are proposing a Wikimedia Project Grant (WikiFactMine) which will automatically scan the daily peer-reviewed scientific literature (up to 10000 articles /day) and extract biological entities (definable in Wikidata). We concentrate on genes and species which have high precision and recall, but with Wikidata-based dictionaries this can be extended to other well defined entities (e.g. cell types or diseases). These, with their citations, are then offered to Wikidata editors for potential inclusion/update. These can also alert Wikipedia editors to new citations. The project includes a Wikimedian in residence in the University of Cambridge. We'd be grateful for comments, endorsement and offers of help. Petermr ( talk) 10:37, 2 August 2016 (UTC)
Gasdermin
Hi... can anyone start writing about Gasdermins? -- محمود ( talk) 21:50, 26 August 2016 (UTC)
- I just created
GSDMA,
GSDMB,
GSDMC, and
GSDMD. Anything in particular you would like to know about Gasdermin?
Boghog (
talk)
22:07, 26 August 2016 (UTC)
- @ Boghog: That's great. Actually, I was editing in some articles and catched Gasdermin D in Inflammasome article. I thought that it's something deserve an article in Wikipedia -- محمود ( talk) 22:34, 26 August 2016 (UTC)
This project's feedback would be appreciated in this discussion, as this could greatly (and positively) affect biological citations! Headbomb { talk / contribs / physics / books} 21:53, 7 September 2016 (UTC)
There is a redirect discussion of which members of this project could help.
Wikipedia:Redirects_for_discussion/Log/2016_October_2#Misfolded is ongoing. We are not sure whether "misfold" is a purely biochemical term or if it could WP:SURPRISE people who were looking for info about folding blankets. Your input would be much appreciated.-- Mr. Guye ( talk) 19:54, 2 October 2016 (UTC)
Portal talk:Molecular and cell biology move discussion
- See move discussion at Portal talk:Molecular and cell biology#Requested move 20 October 2016. Anthony Appleyard ( talk) 20:41, 20 October 2016 (UTC)
review an article
Can someone review/assess the Phosphatidylserine article? I hope this is the correct place to request this. VeniVidiVicipedia ( talk) 12:56, 25 October 2016 (UTC)
Can someone review/assess the Cyclopentenone prostaglandins article? The article has recently been greatly expanded. Although it is currently categorized as a pharmacy and pharmacology article, it may be better categorized in the Molecular and Cellular biology section. Very similar prostaglandins are so categorized (e.g. see prostaglandin). Suggestions on its improvement would be appreciated. Also, the article as currently formatted is correctly redirected from 15-deoxy-Δ12,14-prostaglandin J2 (15-d-Δ12,14-PGJ2), a principal cycloentenone prostaglandin. Is it possible to similarly redirect it from other cyclopenentone prostaglandins viz., Δ12-PGJ2, PGJ2, PGA2, and PGA1, discussed in the article (but not given separate Wikipedia pages elsewhere) and, if so, how do I do that? Thanks, ( User talk:joflaher) 12:01 P.M. 29 Oct 2016.
A request for comment has been made at the above link. Your input is welcome. Boghog ( talk) 11:50, 30 October 2016 (UTC)
CRISPR article status
I noticed in the CRISPR article the following sentence in the lead section: "The CRISPR interference technique has many potential applications, including altering the germline of humans, animals, and food crops." My background is chemistry and I am confident that that statement is incorrect. That can be a very time-consuming error for the reader to sort out. I think the article should mention something about the Cas9-gRNA complex, since it is the synthetic gRNA that makes it into a gene editing tool and a Breakthrough. The press just calls it "CRISPR", but we should try to sort it out for the reader. I think the article should be split, splitting out the engineering technology from the native phenomena (that is some work). It would be nice to note that Cas9 corresponds to CAS III in the diagram at the top of the page. See [14] for some details. Can any of you experts help out?-- 172.56.0.153 ( talk) 09:12, 29 October 2016 (UTC)
- These sources [1] [2] [3] make it very clear that germline modifications with CRISPR is possible and in fact has already successfully carried out in animals. [4] Guide RNA (gRNA) is already mentioned several times in the article. Boghog ( talk) 09:31, 29 October 2016 (UTC)
References
- ^ Evitt NH, Mascharak S, Altman RB (2015). "Human Germline CRISPR-Cas Modification: Toward a Regulatory Framework". The American Journal of Bioethics : AJOB. 15 (12): 25–9. doi: 10.1080/15265161.2015.1104160. PMC 4699477. PMID 26632357.
- ^ Porteus M (2016). "Genome Editing: A New Approach to Human Therapeutics". Annual Review of Pharmacology and Toxicology. 56: 163–90. doi: 10.1146/annurev-pharmtox-010814-124454. PMID 26566154.
- ^ Strong A, Musunuru K (2016). "Genome editing in cardiovascular diseases". Nature Reviews. Cardiology. 14 (1): 11–20. doi: 10.1038/nrcardio.2016.139. PMID 27609628. S2CID 3157463.
- ^ Mou H, Kennedy Z, Anderson DG, Yin H, Xue W (2015). "Precision cancer mouse models through genome editing with CRISPR-Cas9". Genome Medicine. 7 (1): 53. doi: 10.1186/s13073-015-0178-7. PMC 4460969. PMID 26060510.
---Reply---
- SIGH. PLEASE pay attention. The problematic sentence starts out "The
CRISPR interference technique ....." The CRISPR INTERFERENCE technique is a variation on the "CRISPR technique". It uses dCas9 (dead/deactivated Cas9) to reduce gene expression, NOT to achieve genome editting. Normal Cas9 goes to the genome loci and CUTS while dCas9 goes to the same loci and just sits there, thus "interfering" with gene translation/expression.--
172.56.32.19 (
talk)
13:03, 29 October 2016 (UTC)
- Your original post didn't clearly state what the problem was nor why it was a problem. You only stated that you were confident that it was a problem. The immediate problem that you raised is easily solved by a simple edit. {{
sofixit}}.
Boghog (
talk)
13:38, 29 October 2016 (UTC)
- I believe the OP is the sock of an indeffed user; see
Talk:CRISPR#SPLIT
Jytdog (
talk)
17:22, 29 October 2016 (UTC)
- I was asking for the assistance of experts, not Essjays. Sigh. So the article will never be useful.-- 172.56.32.19 ( talk) 17:34, 29 October 2016 (UTC)
- I believe the OP is the sock of an indeffed user; see
Talk:CRISPR#SPLIT
Jytdog (
talk)
17:22, 29 October 2016 (UTC)
- Your original post didn't clearly state what the problem was nor why it was a problem. You only stated that you were confident that it was a problem. The immediate problem that you raised is easily solved by a simple edit. {{
sofixit}}.
Boghog (
talk)
13:38, 29 October 2016 (UTC)
- SIGH. PLEASE pay attention. The problematic sentence starts out "The
CRISPR interference technique ....." The CRISPR INTERFERENCE technique is a variation on the "CRISPR technique". It uses dCas9 (dead/deactivated Cas9) to reduce gene expression, NOT to achieve genome editting. Normal Cas9 goes to the genome loci and CUTS while dCas9 goes to the same loci and just sits there, thus "interfering" with gene translation/expression.--
172.56.32.19 (
talk)
13:03, 29 October 2016 (UTC)
![]() Done Not sure what the history is here. But made the more specific requested change. Thanks for pointing that out!
Ajpolino (
talk)
21:30, 29 October 2016 (UTC)
Done Not sure what the history is here. But made the more specific requested change. Thanks for pointing that out!
Ajpolino (
talk)
21:30, 29 October 2016 (UTC)
- Thank you, Ajpolino. Maybe someday the Essjays of this site will lose interest in using that article to chase persons and the article might reach Good Article status. Someday.--
172.56.1.126 (
talk)
04:18, 31 October 2016 (UTC)
- As you, not me are claiming expertise in this area, why didn't you fix it yourself with a clear edit summary? Your original post was clear as mud. Your second post was much clearer. Boghog ( talk) 07:35, 31 October 2016 (UTC)
On a related note, given that it is one of the most viewed articles within this project's scope, I was thinking that it might be a good candidate for doing some sort of joint publication, where we make a concerted effort to bring it up to high-standard and then submit to an academic journal for peer review. The lure of a possible publication could entice some academics with greater experience in the area. T.Shafee(Evo&Evo) talk 06:18, 31 October 2016 (UTC)
Should this redlink be redirected to microfilament, actin, cytoskeleton, or related article, or does it merit its own independent article? I can't really tell since I'm not that familiar with the topic. There's 4 incoming links from articles to that redlink. Seppi333 ( Insert 2¢) 04:15, 11 November 2016 (UTC)
- It appears
microfilament (cytoskeletal fibers formed by polymerization of monomeric globular (G) actin) is synonymous with actin cytoskeleton.
Boghog (
talk)
04:59, 11 November 2016 (UTC)
- Actin is a protein; cytoskeleton is a structure made of actin. Does that help? Cheers, BatteryIncluded ( talk) 05:22, 11 November 2016 (UTC)
- I guess I'll redirect it to microfilament; I figured that one was the most relevant article. Seppi333 ( Insert 2¢) 05:36, 11 November 2016 (UTC)
Microscopy help
Hey folks! Super-resolution microscopy has had a "clarification-needed" tag for about a year now in the section on Structured illumination microscopy. I find I'm a bit out of my depth on this one. Any chance someone with a bit more experience in this kind of thing could take a peek and clarify the paragraph in question? Thanks a bunch! Ajpolino ( talk) 17:38, 9 November 2016 (UTC)
- It has to do with Phase (waves)#Phase shift on 2 superimposed images. However, I am unable to explain it to Wikipedia standards. I just added an intralink to Phase (waves) and left the 'clarify' tag intact. BatteryIncluded ( talk) 05:56, 11 November 2016 (UTC)
Coverage of God gene hypothesis in the VMAT2 article
This has come up before on the talk page ( Talk:VMAT2#God Gene does not help understanding of VMAT2 function), but I figured I'd raise the issue here to attract more input. God gene hypothesis is a fringe theory that suggests that a neurotransmitter transporter which is located on synaptic vesicles in monoamine neurons ( VMAT2) has some form of significant role in spirituality. I've removed content which covers the god gene hypothesis from the VMAT2 article twice now - once recently and once a while ago - per WP:ONEWAY. The question I have for you all is this: should this article cover a fringe theory because the theory is covered in popular culture? Please respond here if you have any input: Talk:VMAT2#Re: covering the God gene hypothesis. Seppi333 ( Insert 2¢) 19:30, 18 November 2016 (UTC)
Proposed merger of Neto 1 and NETO1
I have proposed that the article Neto 1 be merged into NETO1 as both articles appear to be on the same topic and "NETO1" is currently a more thorough discourse of the topic. I have created a proposed merger Talk Space on the NETO1 page and listed what I need help with.
I am not a scientist, biologist, etc., so I am not sure which article title is correct. I would appreciate any pertinent information that you can provide. If someone can assist with rewriting the article after the merger is complete that would be appreciated. Currently both articles are lacking substance and incoming links. I'm being pretty bold proposing a merger after being on the site for less than two months, so your patience while I learn more about editing on Wikipedia is appreciated.
Thanks for your help! Zcarstvnz ( talk) 22:50, 20 November 2016 (UTC)
Splitting endogenous molecules used as drugs
See Wikipedia_talk:WikiProject_Medicine#Splitting_articles_about_endogenous_molecules_used_as_drugs Jytdog ( talk) 23:04, 22 November 2016 (UTC)
maybe i have missed it, but i have never found a page that gives an overview of reagents. I just added a tiny bit at that article but it is a chicken scratch. Is there a master artcle somewhere that talks about reagents and their importance? Jytdog ( talk) 06:05, 23 November 2016 (UTC)
- Not as far as I know. I looked about for one back when I updated substrate (chemistry) but didn't find anything sensible then. T.Shafee(Evo&Evo) talk 00:53, 24 November 2016 (UTC)
A request for comment has been made at the above link. Your input is welcome. Boghog ( talk) 16:45, 27 November 2016 (UTC)
Trivial naming question
Trivial difference that's been bothering me: Translation (biology) vs Transcription (genetics). Would it make sense to harmonise them both to XYZ (genetics)? I can't imagine it actually causes any confusion, it just seems untidy. T.Shafee(Evo&Evo) talk 08:47, 6 November 2016 (UTC)
- Good eye! I had never noticed that. While I agree this isn't super critical, for what it's worth my vote is harmonizing to XYZ (biology). My guess is that's more in line with how the average person would think of it.
Ajpolino (
talk)
15:51, 6 November 2016 (UTC)
- Wow, I never noticed that either. Yep, makes sense to make them parallel; don't really care which one we keep.
Opabinia regalis (
talk)
20:56, 6 November 2016 (UTC)
- I'm also learning towards XYZ (biology). They're as relevant to molecular biology as to genetics, and they're things students learn about in general biology courses, rather than specialist genetics courses.
Adrian J. Hunter(
talk•
contribs)
03:19, 7 November 2016 (UTC)
- Actually, I agree that "(biology)" is the sensible disambig, seeing as there are no other biological interpretations of "translation". T.Shafee(Evo&Evo) talk 02:56, 11 November 2016 (UTC)
- I'm also learning towards XYZ (biology). They're as relevant to molecular biology as to genetics, and they're things students learn about in general biology courses, rather than specialist genetics courses.
Adrian J. Hunter(
talk•
contribs)
03:19, 7 November 2016 (UTC)
- Wow, I never noticed that either. Yep, makes sense to make them parallel; don't really care which one we keep.
Opabinia regalis (
talk)
20:56, 6 November 2016 (UTC)
2016 Community Wishlist Survey Proposal to Revive Popular Pages

Greetings WikiProject Molecular Biology/Molecular and Cell Biology/Archive 7 Members!
This is a one-time-only message to inform you about a technical proposal to revive your Popular Pages list in the 2016 Community Wishlist Survey that I think you may be interested in reviewing and perhaps even voting for:
If the above proposal gets in the Top 10 based on the votes, there is a high likelihood of this bot being restored so your project will again see monthly updates of popular pages.
Further, there are over 260 proposals in all to review and vote for, across many aspects of wikis.
Thank you for your consideration. Please note that voting for proposals continues through December 12, 2016.
Best regards, Stevietheman — Delivered: 18:04, 7 December 2016 (UTC)
This template has surprisingly few fields. I wanted to add alt names to Histidine—tRNA ligase and the only place I could do it was in the "name" field which seems like clutter. Shouldn't this have most everything in Template:Infobox protein? Jytdog ( talk) 06:18, 12 December 2016 (UTC)
- The enzyme infobox is meant to represent information about an
Enzyme Commission number (EC number). Strictly speaking, an EC number represents the reaction catalyzed by an enzyme, not the enzyme itself and therefore there is not a strict one to one relationship between EC numbers and proteins. Several structurally unrelated proteins may catalyze the same reaction and a single protein may catalyze more than one reaction. Hence several otherwise unrelated proteins may be assigned the same EC number and a single protein may be assigned more than one EC number. Hence most of the fields in {{
infobox protein}} are not appropriate for {{
infobox enzyme}} and vice versa. I agree that
|AltNames=might be useful, hence I have added this as an optional parameter to the template. Boghog ( talk) 06:58, 12 December 2016 (UTC)- Thanks! Hm! So if the enzyme box is for a reaction, then shouldn't there always be a) a protein box; and b) as many enzyme boxes as there are reactions that the protein is involved in? Hm.
Jytdog (
talk)
07:10, 12 December 2016 (UTC)
- The rule that I have been following is if there is a strict one-to-one relationship between EC number and human gene/protein, then there should be a single article that covers both the gene and the enzyme. If there is not a one-to-one relationship, it is much cleaner to have separate gene and enzyme articles. Due to their large size, it is not practical to place more than one {{ Infobox gene}} in the same article. Because {{ Infobox protein}} is more compact, sometimes more than one of these is placed in articles about protein families. Finally, the SOP that WP:MCBMOS and Gene Wiki follow is one gene, one article. Boghog ( talk) 07:35, 12 December 2016 (UTC)
- Ideally
|ECnumber=in {{ Infobox protein}} should be able to list more multiple EC numbers. I will look into this. Boghog ( talk) 07:40, 12 December 2016 (UTC)- I am really clueless here. :) When do you use infobox: gene vs infobox: protein? Sorry to bother you but neither template says when you use one vs the other.
Jytdog (
talk)
07:48, 12 December 2016 (UTC)
- It depends on what is the scope of the article. If there is a one-to-one relationship between the gene and the enzyme, then use both infoboxes in same article. If there are several genes that encode proteins with the same enzymatic activity, focus on enzymology and use {{ Infobox enzyme}}. If there are a limited number of genes associated with a specific enzymatic activity, it also might make sense to include a limited number of {{ Infobox protein}} as well (see for example Trypsin). If there is a single protein with more than one enzymatic activity, then {{ Infobox protein}} or {{ Infobox gene}} listing the multiple EC numbers would make sense. Basically it comes down to common sense and editorial judgement. To give you some background, most of these enzyme and gene articles were created independently of each other. The enzyme articles systematically by a bot run by User:WillowW and the protein articles by User:ProteinBoxBot. Some of these have since been merged where it made sense. Boghog ( talk) 12:04, 12 December 2016 (UTC)
- I am really clueless here. :) When do you use infobox: gene vs infobox: protein? Sorry to bother you but neither template says when you use one vs the other.
Jytdog (
talk)
07:48, 12 December 2016 (UTC)
- Thanks! Hm! So if the enzyme box is for a reaction, then shouldn't there always be a) a protein box; and b) as many enzyme boxes as there are reactions that the protein is involved in? Hm.
Jytdog (
talk)
07:10, 12 December 2016 (UTC)
Featured article nomination for beta-Hydroxy beta-methylbutyric acid
I recently renominated this article at FAC ( Wikipedia:Featured article candidates/Beta-Hydroxy beta-methylbutyric acid/archive2) and was wondering if anyone from this WikiProject would be interested in doing a review of this article's pharmacology content. Every statement in this section is supported by at least 1 WP:MEDRS-quality review.
Since this compound's immediate biomolecular targets aren't known, there's a large amount of material on its downstream signaling pathways (primarily through mTORC1 and its interactions with the ubiquitin-proteasome system) in the pharmacodynamics section; this content is mostly based upon clinical trials in humans, but there's some contextual information on its signaling mechanisms which comes from in vitro and animal studies (I've clearly indicated what statements are supported by human in vivo research vs in vitro or animal research in the article text). The pharmacokinetics section covers its biosynthetic and metabolic pathways in humans. Consequently, it would be ideal for someone with a pharmacology and/or molecular biology background to take on a review of this material.
I would really appreciate it if someone from this WikiProject would take on a review of this article content at FAC. Seppi333 ( Insert 2¢) 20:42, 25 October 2016 (UTC)
- Still need a reviewer for this content; the current FAC nomination is located at Wikipedia:Featured article candidates/Beta-Hydroxy beta-methylbutyric acid/archive3. Seppi333 ( Insert 2¢) 17:48, 1 January 2017 (UTC)
Missing topics list
My list of missing topics about metabolism is updated - Skysmith ( talk) 15:11, 8 January 2017 (UTC)
User needs help with new article with DYK potential
Hi there. A new user created testosterone-cortisol ratio through AfC and requested some help improving it. I did a copyedit and formatted some references but have no knowledge about the subject itself (or molecular biology for that matter). Maybe someone here can help this user out? The topic sounds interesting enough to be featured in T:DYK if the article were expanded a bit more. Regards So Why 13:17, 17 January 2017 (UTC)
- oy content is almost entirely WP:Biomedical information sourced to refs that fail WP:MEDRS. Am not sure this article will survive. Will work on it. Jytdog ( talk) 20:23, 17 January 2017 (UTC)
WikiJournal of Medicine promotion

 |
The WikiJournal of Medicine is a free, peer reviewed academic journal which aims to provide a new mechanism for ensuring the accuracy of Wikipedia's biomedical content. We started it as a way of bridging the Wikipedia-academia gap. [1] It is also part of a WikiJournal User Group with other WikiJournals under development. [2] The journal is still starting out and not yet well known, so we are advertising ourselves to WikiProjects that might be interested. |
Engaging Wikipedians
- Original articles on topics that don't yet have a Wikipedia page, or only a stub/start
- Wikipedia articles that you are willing to see through external peer review (either solo or as in a group, process analogous to GA / FA review)
- Image articles, based around an important medical image or summary diagram
Engaging non-Wikipedians
We hope that an academic journal format may also encourage non-Wikipedians to contribute who would otherwise not. Therefore, please consider:
- Printing off the advertisement poster and distribute in tearooms & noticeboards at your place of work
- Emailing around the pdf through contact networks or mailing lists (suggested wording)
If you want to know more, we recently published an editorial describing how the journal developed. [3] Alternatively, check out the journal's About or Discussion pages.
- ^ Masukume, G; Kipersztok, L; Das, D; Shafee, T; Laurent, M; Heilman, J (November 2016). "Medical journals and Wikipedia: a global health matter". The Lancet Global Health. 4 (11): e791. doi: 10.1016/S2214-109X(16)30254-6. PMID 27765289.
- ^ "Wikiversity Journal: A new user group". The Signpost. 2016-06-15.
- ^ Shafee, T; Das, D; Masukume, G; Häggström, M (2017). "WikiJournal of Medicine, the first Wikipedia-integrated academic journal". WikiJournal of Medicine. 4. doi: 10.15347/wjm/2017.001.

Additionally, the
WikiJournal of Science is just starting up under a similar model and looking for contributors. Firstly it is seeking editors to guide submissions through external academic peer review and format accepted articles. It is also encouraging submission of articles in the same format as Wiki.J.Med. If you're interested, please come and discuss the project on the
journal's talk page, or the
general discussion page for the WikiJournal User group.
T.Shafee(Evo&Evo)
talk
10:33, 19 January 2017 (UTC)
Orphaned Gene/Protein Articles
Hi, I've been going through the backlog at CAT:ORPHAN lately, and I keep running across articles like EAF1, E2F7, etc etc for various genes and proteins. There's a ton of them and they're all orphans, almost all created by User:ProteinBoxBot, all the way back from 2009 and earlier. I don't know the first thing about molecular biology, so I have no way to determine if these articles are correct, useful, or notable...really anything about them. I would like to be able to de-orphan them in order to clear out the backlog there, but I don't want to make a mess of anything given my lack of knowledge.
Is there any policy or guideline covering the notability of proteins/genes as separate articles? Is there a central list article they could be linked at for the purposes of de-orphaning? Or even a group of such articles, like "genes on human chromosome 3" or something?
Any input would be a huge help, like I said I'd love to do anything that can clear out the backlog, but I don't want to break anything out of ignorance. ♠ PMC♠ (talk) 04:07, 24 January 2017 (UTC)
- Thanks for your note. Notability can normally be established by following the "Human PubMed Reference" link. Both genes have been mentioned in a number of peer-reviewed articles, hence I think they pass the notability requirement. Unfortunately there does not appear a uniform way to deophanize these articles and each one must be handled separately. EAF1 (along with EAF2, CDK9, MLLT3/AF9, AFF ( AFF1 or AFF4), the P-TEFb complex and ELL ( ELL, ELL2 or ELL3)) is a component of the super elongation complex. Creating the second article would deorphanize the former. E2F7 interacts with E2F8. Boghog ( talk) 06:47, 24 January 2017 (UTC)
- Thanks for the fast response! Okay, so, broadly speaking, if a gene/protein name returns at least a couple of results on the Human PubMed Reference then it would be considered reasonably notable. That makes sense. I don't have the technical knowledge to go creating articles like
super elongation complex, but would it be useful if I made a list of any orphaned gene articles that I found, to bring them to the attention of this WikiProject? I don't want to step on any toes or clutter anything up. ♠
PMC♠
(talk)
07:08, 24 January 2017 (UTC)
- If you post any of them here I think people would be happy to help de-orphan them.
Ajpolino (
talk)
23:40, 24 January 2017 (UTC)
- Sweet. I've dropped the first set (A-C from February 2009) onto my personal sandbox. I can always move them elsewhere if that would be better. ♠ PMC♠ (talk) 07:42, 25 January 2017 (UTC)
- If you post any of them here I think people would be happy to help de-orphan them.
Ajpolino (
talk)
23:40, 24 January 2017 (UTC)
- Thanks for the fast response! Okay, so, broadly speaking, if a gene/protein name returns at least a couple of results on the Human PubMed Reference then it would be considered reasonably notable. That makes sense. I don't have the technical knowledge to go creating articles like
super elongation complex, but would it be useful if I made a list of any orphaned gene articles that I found, to bring them to the attention of this WikiProject? I don't want to step on any toes or clutter anything up. ♠
PMC♠
(talk)
07:08, 24 January 2017 (UTC)
This section is poorly written with misleading information. This requires serious editing. Andrzej Slominski, MD, PhD Endowed Professor of Dermatology Professor of Pathology Program Director of Dermatopathology Fellowship Senior Scientist, Comprehensive Cancer Center Cancer Chemoprevention Program Member of the Editorial Boards of Exp Dermatol, J Pineal Research, PLoS ONE, Dermato-Endocrinology, Int J Mol Sci, Melanoma Management, Pol J Path Department of Dermatology University of Alabama at Birmingham Volker Hall 476C Box 109 1720 2nd Avenue South VH 476C Birmingham, AL 35294 Phone: 205.934.5245 e-mail: aslominski@uabmc.edu — Preceding unsigned comment added by 64.134.187.167 ( talk) 14:28, 3 February 2017 (UTC)
- Disclaimer: I wrote most of the RAR-related orphan receptor article. Most of the content was written ten years ago and a lot has happened since then. Precisely which part of the article is poorly written with misleading information? Your comments are appreciated but not very helpful. Per {{ sofixit}}, it would be much more constructive to provide specific suggestions on how to improve the article and even better, improve the article yourself. Is the problem with the sources supplied or the interpretation of those sources? Even less useful is your excessive description of your qualifications which does not carry much weight around here. Boghog ( talk) 22:21, 3 February 2017 (UTC)
- I have made an effort to update the article based on more recent reviews. I hope this is an improvement. Is there any information that is still misleading? Boghog ( talk) 18:42, 4 February 2017 (UTC)
Notice to participants at this page about adminship
Many participants here create a lot of content, may have to evaluate whether or not a subject is notable, decide if content complies with BLP policy, and much more. Well, these are just some of the skills considered at Wikipedia:Requests for adminship.
So, please consider taking a look at and watchlisting this page:
You could be very helpful in evaluating potential candidates, and even finding out if you would be a suitable RfA candidate.
Many thanks and best wishes,
Anna Frodesiak ( talk) 01:11, 10 February 2017 (UTC)
Nicotinamide adenine dinucleotide (NAD)
Two weeks ago I tried to make a meaningful contribution to the NAD article on Wikipedia.
It was labeled as «promotional garbage» by user:Jytdog. The last part of his edit summary was also pure and intended nonsense, and had nothing to do with the content of my original edit.
Within a couple of hours or so I was «edit warring», guilty of «sock puppetry» and given a free, one week sabbatical from Wikipedia.
My original edit, a short passage, is now on display at the NAD talk page.
Feedback from other users is appreciated. Thank you. Clowns und Kinder ( talk) 18:53, 24 February 2017 (UTC)
Wikidata and drug info
Please see Wikipedia_talk:WikiProject_Medicine#Wikidata_again. This is like playing whackamole. Jytdog ( talk) 21:36, 27 February 2017 (UTC)
Opinions requested
Opinions are requested at Talk:Expanded genetic code#mouse code about whether certain content is an acceptable use of primary sources. Thanks, Adrian J. Hunter( talk• contribs) 06:11, 13 March 2017 (UTC)
Merge proposal
Thoughts on this merge proposal (or other ideas on what to do with History of penicillin) would be much appreciated. Thanks! Ajpolino ( talk) 04:51, 14 March 2017 (UTC)
Proposal of using full size image for RNA expression pattern
Hello WP:MCB! In infoboxes of gene/protein articles, there are small thumbnail images of RNA expression pattern. I proposed switching these small thumbnail to full size image, because Wikipedia/Mediawiki image system was changed. Since image data are recalled from Wikidata, I post the proposal at Wikidata page ( wikidata:Property talk:P692#How about using full size image instead of small thumbnail?). I would like to get your thoughts on that. Thank you. -- Was a bee ( talk) 06:28, 18 March 2017 (UTC)
WP:CHEMISTRY/ WP:CHEMICALS shorcut updated
Note that per this RFC, the shortcuts to WP:CHEMISTRY/ WP:CHEMICALS have been updated.
- WP:CHEM now refer to WP:CHEMISTRY
- WP:CHEMS now refer to WP:CHEMICALS
- WP:CHM is deprecated
Old discussions have had their shortcuts updated already. If I have made a mistake during an update, feel free to revert. Headbomb { talk / contribs / physics / books} 16:04, 23 March 2017 (UTC)
Critiquing "Mitochondrial protein-transporting ATPase" Article
The article "Mitochondrial protein-transporting ATPase" there are only four citations present. Three out of four of the citations used are 20 or more years old. To make this Article on Wiki better more recent citations should be used. Also more Citations should be referenced as well. Dintamang ( talk) 19:23, 24 March 2017 (UTC)
Extension of 'Topic Page' review articles from PLOS Computational Biology to PLOS Genetics

The journal group PLOS is extending its ' Topic Page' review format that was spearheaded by PLOS Computational Biology to also include PLOS Genetics. In this format, accepted articles are dual-published both in the journal, and as Wikipedia pages (see Wikipedia category).
Suitable topics must either currently lack a Wikipedia page, or have only stub/start class contents. If you you would like to submit such a review article, see
these guidelines. If you have any recommendations for topics to be commissioned, feel free to let any of the involved editors know:
T Shafee (PLOS Gen),
D Mietchen (PLOS Comp Biol).
T.Shafee(Evo&Evo)
talk
12:27, 25 March 2017 (UTC)
Infobox for enzyme vs protein
While diagnosing how
alcohol dehydrogenase displayed compared to
inositol oxygenase, the former has a
BRENDA link autogenerated when |EC_number= is present as a feature of {{
infobox enzyme}}. The latter does have an EC number passed, but no such link, which it turns out is because {{
infobox protein}} does not have the BRENDA-link feature. But I also notice that this infobox uses a different parameter for the EC number (|ECnumber=, note lack of underscore). So three concerns:
- It's confusing that these two related templates use different parameter names for the same data.
- Is {{ infobox enzyme}} just a more specific infobox, or is there a reason one would want to use {{ infobox protein}} for an enzyme?
- And objection to cloning the BRENDA link magic into {{ infobox protein}}?
I'll post pointers from both templates' talkpages to here momentarily. DMacks ( talk) 14:56, 3 April 2017 (UTC)
- My
twothree cents:
- No objection to standardizing parameters names between related templates. I like underscores since IMHO they make the parameter name more readable.
- Strictly speaking, an EC number does not represent an enzyme, but rather a reaction catalyzed by an enzyme. More than one (iso)enzyme can catalyze the same reaction. Hence {{ infobox enzyme}} is not a more specific version of {{ infobox protein}}, they serve two distinct purposes.
- No objections to adding a BRENDA link into {{ infobox protein}}. Boghog ( talk) 15:53, 3 April 2017 (UTC)
- IMO, adding a BRENDA link to infobox protein would be useful. Seppi333 ( Insert 2¢) 17:48, 3 April 2017 (UTC)
Scope of Light-independent reactions
Hello. After the RM results at Talk:Light-independent reactions, the article " Light-independent reactions" needs attention. The scope of this article is confusing as the article suffered from naming and scope issues. I think participants here may be interested in this. -- George Ho ( talk) 13:12, 9 April 2017 (UTC)
- I left some suggestions on the talk page. This article sure does need cleanup. Adrian J. Hunter( talk• contribs) 00:58, 10 April 2017 (UTC)
Requests for comment on Microscope article
Please comment on a few requests for comments on the Microscope article.
Talk:Microscope#Request_for_comment_on_how_a_scanning_electron_microscope_works
Talk:Microscope#Request_for_comment_on_ultramicroscope
Talk:Microscope#RFC_should_article_focus_on_instrument.2C_microscope.2C_or_technique.2C_microscopy — Preceding unsigned comment added by 2600:387:6:807:0:0:0:C2 ( talk) 14:23, 27 April 2017 (UTC)
Thank you, -- 2601:648:8503:4467:F4B3:6D6C:9DCC:DC06 ( talk) 21:09, 1 April 2017 (UTC)
TfD: {{ Steroid metabolism modulators}} and {{ Cytochrome P450 modulators}}
The above navboxes are being proposed for deletion. Your comments are welcome here. Boghog ( talk) 20:46, 29 April 2017 (UTC)
| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | ← | Archive 5 | Archive 6 | Archive 7 | Archive 8 | Archive 9 | Archive 10 |
- Can this resource be used for linking on WP pages? I am asking because one of the contributors removes linking to TCDB classification on the grounds that are not entirely clear to me (i.e. like here). We need a consensus about this. Thank you. My very best wishes ( talk) 21:41, 24 April 2016 (UTC)
- WP:LINKVIO is the complaint, so it seems to me. Some articles that were copied from that group's website are up at WP:CP because that group apparently copies text from non-free third-party sources. Jo-Jo Eumerus ( talk, contributions) 22:05, 24 April 2016 (UTC)
- The diff cited is an edit by me, and yes, that is correct, that WP:LINKVIO was my reason for that edit. I also considered just deleting the citations and adding "citation needed" tags to everything, but I really do not think that there will be alternative sourcing. Additionally, I will point out that there are WP:UNDUE issues about the material I deleted. It really should not be presented by Wikipedia as though it were the established nomenclature system used in the preponderance of sources. However, I think the underlying research could quite reasonably be cited in a possible section about the molecular phylogenetics of the ion channels. -- Tryptofish ( talk) 00:37, 25 April 2016 (UTC)
- Thank you so much for raising this! The issue isn't so much about "linking" as the largescale importing of content from that database into WP. Pinging User:Transporter Guy who has been working on a project to copy lots of content from the database into WP first as member of the lab that created the database, and now as a volunteer still invested in the project; see this discussion I had with them.
- The copy/pasting has been triggering lots of copyright alerts from bots, but the database is appropriately licensed. The problem, as discussed here and here, and here, is that TCDB itself appears to contain copyright violations and those violations are being brought over into Wikipedia sometimes. Two admins, User:Opabinia regalis and User:Diannaa have raised this issue with Transporter Guy but there doesn't seem to be a change in behavior yet.
- See also Wikipedia:Articles for deletion/Voltage-gated ion channel (VIC) family which is about content imported from TCDB into a new article based on an idiosyncratic classification.
- See also edits like this and this where Tranasporter guy over-wrote the existing article with content from TCDB, which they have done several times.
- See also edits like this where Transporter guy created a new article with content copied from TCDB and then blanked and redirected an existing one; they have done this several times too.
- Some folks have appreciation for all the content creation like here and here and this.
- This seems like something the project should discuss and get consensus on before the work continues. Thanks again for raising this MVBW! I am placing notices at a few other WikiProjects that may be interested in the discussion.
Jytdog (
talk)
22:18, 24 April 2016 (UTC)
- Edits like that are fine because he moved this content to appropriate sub-pages. As about TCDB abstracts, it appears that anyone who wants to use them must check that the inserted text does not contain any copyright violations and be responsible for that. Speaking about the links, they are definitely very useful for a reader. I should think about it. My very best wishes ( talk) 22:26, 24 April 2016 (UTC)
- Yes, thanks for starting this discussion, My very best wishes. Jytdog seems to have outlined the nature of the copyright problem – the database itself appears to to be violating copyrights by copying directly from copyright-protected published articles or abstracts. Regardless of what licence the database claims to have, copyvio content from it may not be imported into Wikipedia. Because Transporter Guy has done exactly that in several articles, and continued to do so after being clearly told to stop, I have asked for a contributor copyright investigation, a full review of the copyright status all his contributions to date. Jo-Jo Eumerus is of course right to point out that we should not link in any way to pages of the database that violate other people's copyrights, though that is not the central issue here.
- All attempts to convince Transporter Guy that he should respect our guidelines for
conflict of interest editors have failed and he continues to
WP:REFSPAM links to his employer's database into the project. Voluntarily or not, that needs to stop.
Justlettersandnumbers (
talk)
23:15, 24 April 2016 (UTC)
- It's not clear to me that the effort needs to stop permanently, but yes, paused for now while the community evaluates if it should continue and if so, how. It may well be that it continues with the blessing of the community, in a different way. That could be a happy outcome here. Jytdog ( talk) 23:24, 24 April 2016 (UTC)
- Just want to add that input from folks who haven't thought about this before would be very valuable. Jytdog ( talk) 23:49, 24 April 2016 (UTC)
- Can be used with care per our guidelines. I can see that two admins raised the copyright issue and agree with everything they said here. Let's also check the existing guidelines. There are two slightly different issues. First, if anyone reuses texts from TCDB, he/she must be absolutely sure that the text does not contain any copyright violations and be responsible for that. This is nothing special and applies to any content. Second, we do have certain guidelines for linking. According to them, we should not link to any individual pages that violate copyright, and this is particularly relevant when linking to sites such as Scribd or YouTube, where due care should be taken to avoid linking to material that violates copyright. Hence, the linking to individual YouTube entries is actually allowed, but anyone who makes such link must be sure this particular YouTube entry is not a copyright violation. By the same logic, anyone who makes link to an individual TCDB abstract must be sure this particular abstract had no copyright violations. My very best wishes ( talk) 23:53, 24 April 2016 (UTC)
- One should also realize that TCDB is a major and important database. It is well known to everyone who works in the area of protein bioinformatics or membrane proteins. It is used by every other major protein database, such as Uniprot or Pfam. It implements a classification of transporters approved by the International Union of Biochemistry and Molecular Biology. It should be used here for the benefit of the project. My very best wishes ( talk) 13:53, 25 April 2016 (UTC)
- Based on your comment, I searched for the IUBMB's website listing their recommendations:
[1]. I cannot find TCDB there. --
Tryptofish (
talk)
20:43, 25 April 2016 (UTC)
- This is in every textbook on the subject. See for example, Protein Families: Relating Protein Sequence, Structure, and Function by Christine Orengo and Alex Bateman. Here it tells that "Transporter Classification Database TCDB (www.tcdb.org) (Saier et al.,2006, 2009) presents the comprehensive IUBMB (International Union of Biochemistry...) approved and adopted classification system...", and so on. This textbook dedicates an entire chapter to the TCDB. And here is large number of other books that describe TCDB. My very best wishes ( talk) 21:25, 25 April 2016 (UTC)
- See also here - Tryptofish, this appears at your link under "Membrane transport proteins". Opabinia regalis ( talk) 21:33, 25 April 2016 (UTC)
- Thanks for finding that, Opabinia. And yet, reading it, it just gets stranger and stranger. At the top it does indeed say that "Practically all the material presented here is based on the work of" the people who maintain the TCDB website. But then, look at the list itself! It is nothing like the TCDB classification as displayed at the TCDB website and edited onto Wikipedia pages! If one looks at Voltage-gated ion channel#Nomenclature, one sees a supposed VIC superfamily, with 8 subfamilies. I cannot find that on the linked page from IUBMB, where a much more conventional classification is found. But – if one clicks through from that on the 1.A links wherever 1.A occurs (alpha-helical channels, not to be confused with I.A), then there the VIC is! So in fact (way down in the "fine print"), yes the Saier group does have a TCDB listing at IUBMB, but it is not organized that way at the IUBMB website, where instead, they use much more traditional structural and functional parameters for classification. -- Tryptofish ( talk) 21:56, 25 April 2016 (UTC)
- Actually, on looking further, not even that! Here is the 1.A link: [2]. It has "VIC" as a member there, followed by other entries, but those subsequent entries are not subfamilies of VIC. They are just other kinds of alpha-helix-containing molecules, along with VIC, but not subtypes of VIC. Compare that listing and numbering scheme with what had been edited here: [3], where some of them are subtypes. The Wikipedia edits include a "K+ transporter" family that is nowhere at IUBMB, and it uses the highly unconventional "TrK" abbreviation, which is very widely known as something completely different. Not the same at all, no matter where one looks at IUBMB. -- Tryptofish ( talk) 22:15, 25 April 2016 (UTC)
- (
edit conflict) In any case, the book chapter you linked in your first diff was written by the same people who maintain the TCDB website! It's not in a textbook, but rather a compilation book made up of chapters provided by authors. And as for your Google link, I wonder if you even looked at it. The initial returns are just Wikipedia and self-referencing sources by the TCDB authors! --
Tryptofish (
talk)
21:37, 25 April 2016 (UTC)
- Then check another textbook
[4]. The differences appear because TCDB classification is constantly developing. Dr. Saier was the person who developed this widely used classification, and he continue doing this, as reflected in the more recent versions of TCDB. He is basically a
classic in this field, meaning that everyone who works with classification of proteins must be familiar with his work. Now, speaking about "copyright problems", I think they are strongly inflated, and TCDB can be actually used as
a reliable source. I saw many copyright violations in academic journals. This is happening when an author is reusing his own nearly identical texts in a number of different publications (as occasionally do TCDB authors). This is bad, but this does not invalidate any of these academic journals as RS.
My very best wishes (
talk)
22:23, 25 April 2016 (UTC)
- That's not "another" textbook, but it is a textbook. Anyway, it has a table that is just like what is at IUBMB, but not like what you purport is later hot-off-the-press work. If the more recent stuff gets taken up by
independent
secondary sources, then Wikipedia can use it, but
not before that happens, and it hasn't happened yet. And you appear to
not hear other editors when you talk about RS. Nobody is saying that copyright issues are about
WP:RS; it's about
WP:LINKVIO. --
Tryptofish (
talk)
22:34, 25 April 2016 (UTC)
- I am telling that if anyone needs to use TCDB for referencing and provides a link to TCDB in inline citation, he can do it. And remember that WP:RS is a policy, one of five pillars. It overrides guidelines, such ones for linking.
My very best wishes (
talk)
22:42, 25 April 2016 (UTC)
-
WP:C,
WP:CV. --
Tryptofish (
talk)
23:13, 25 April 2016 (UTC)
- No one suggests making copyright violations. Look, I
explained this to you already on your talk page.
My very best wishes (
talk)
23:29, 25 April 2016 (UTC)
-
WP:LINKVIO is part of
WP:C. --
Tryptofish (
talk)
23:33, 25 April 2016 (UTC)
- Let's assume that I found an example of a copyright violation in the
Journal of the American Chemical Society (authors used a couple of paragraphs of text which is a close paraphrase of something they published earlier in another journal; yes, that happens). Would it make any reference and linking to JACS from WP WP:LINKVIO? No, absolutely not. That is what I am talking about. Would it make a reference and linking to the particular article in JACS (where such violation happened) a WP:LINKVIO? I am not sure, but claiming that a person who made such ref was a malicious violator and we must check or remove all references to JACS in WP would be ridiculous.
My very best wishes (
talk)
23:50, 25 April 2016 (UTC)
- The situation here is that there is an external website where there appears to be copyright violation of material published by other persons, not simply authors repeating their own material – which is not a copyright violation anyway, just lazy writing (one does not violate one's own copyright). The copied material is apparently from other authors, not from the authors of the website. No one at Wikipedia has claimed that an editor is a malicious violator. I just reverted the links. As indicated by
WP:C. Which is a policy with legal implications. Reverting something is not calling anyone a malicious violator. On the other hand, beating a dead horse can get rather annoying. --
Tryptofish (
talk)
01:27, 26 April 2016 (UTC)
- I am happy to hear this is not a copyright violation (???) Because in this case it was a previous publication by same authors/lab [5]. Same in this case [6]. People who wrote the patent application probably borrowed text from an old review by the same group, but the review is apparently not accessible through Google search. My very best wishes ( talk) 04:10, 26 April 2016 (UTC)
- The situation here is that there is an external website where there appears to be copyright violation of material published by other persons, not simply authors repeating their own material – which is not a copyright violation anyway, just lazy writing (one does not violate one's own copyright). The copied material is apparently from other authors, not from the authors of the website. No one at Wikipedia has claimed that an editor is a malicious violator. I just reverted the links. As indicated by
WP:C. Which is a policy with legal implications. Reverting something is not calling anyone a malicious violator. On the other hand, beating a dead horse can get rather annoying. --
Tryptofish (
talk)
01:27, 26 April 2016 (UTC)
- Let's assume that I found an example of a copyright violation in the
Journal of the American Chemical Society (authors used a couple of paragraphs of text which is a close paraphrase of something they published earlier in another journal; yes, that happens). Would it make any reference and linking to JACS from WP WP:LINKVIO? No, absolutely not. That is what I am talking about. Would it make a reference and linking to the particular article in JACS (where such violation happened) a WP:LINKVIO? I am not sure, but claiming that a person who made such ref was a malicious violator and we must check or remove all references to JACS in WP would be ridiculous.
My very best wishes (
talk)
23:50, 25 April 2016 (UTC)
-
WP:LINKVIO is part of
WP:C. --
Tryptofish (
talk)
23:33, 25 April 2016 (UTC)
- No one suggests making copyright violations. Look, I
explained this to you already on your talk page.
My very best wishes (
talk)
23:29, 25 April 2016 (UTC)
-
WP:C,
WP:CV. --
Tryptofish (
talk)
23:13, 25 April 2016 (UTC)
- I am telling that if anyone needs to use TCDB for referencing and provides a link to TCDB in inline citation, he can do it. And remember that WP:RS is a policy, one of five pillars. It overrides guidelines, such ones for linking.
My very best wishes (
talk)
22:42, 25 April 2016 (UTC)
- That's not "another" textbook, but it is a textbook. Anyway, it has a table that is just like what is at IUBMB, but not like what you purport is later hot-off-the-press work. If the more recent stuff gets taken up by
independent
secondary sources, then Wikipedia can use it, but
not before that happens, and it hasn't happened yet. And you appear to
not hear other editors when you talk about RS. Nobody is saying that copyright issues are about
WP:RS; it's about
WP:LINKVIO. --
Tryptofish (
talk)
22:34, 25 April 2016 (UTC)
- Then check another textbook
[4]. The differences appear because TCDB classification is constantly developing. Dr. Saier was the person who developed this widely used classification, and he continue doing this, as reflected in the more recent versions of TCDB. He is basically a
classic in this field, meaning that everyone who works with classification of proteins must be familiar with his work. Now, speaking about "copyright problems", I think they are strongly inflated, and TCDB can be actually used as
a reliable source. I saw many copyright violations in academic journals. This is happening when an author is reusing his own nearly identical texts in a number of different publications (as occasionally do TCDB authors). This is bad, but this does not invalidate any of these academic journals as RS.
My very best wishes (
talk)
22:23, 25 April 2016 (UTC)
- Based on your comment, I searched for the IUBMB's website listing their recommendations:
[1]. I cannot find TCDB there. --
Tryptofish (
talk)
20:43, 25 April 2016 (UTC)
- P.S. Many major protein databases, such as UniProt, Pfam and Protein Data Bank, do linking to TCDB. Therefore, I do not see any reason why we can not use such links here. My very best wishes ( talk) 14:29, 26 April 2016 (UTC)
- There is certainly no problem with citing such other websites, as they are secondary sources with respect to TCDB and they presumably do not have copyright issues. So that is a good, and preferable, approach to this content. At the same time, the due weight policy requires us to present TCDB within the context of any of the other classifications present in those secondary sources, rather than presenting TCDB as the only one. -- Tryptofish ( talk) 21:34, 26 April 2016 (UTC)
- In the last phrase I am telling not about sourcing, but about links. Links can not be primary or secondary. If other widely used resources do not see any copyright problems with linking to TCDB (and believe me, those are very serious resources supported by large staff including people who are familiar with copyright), I do not see any problems with linking to TCDB here.
My very best wishes (
talk)
23:12, 26 April 2016 (UTC)
- I know that links cannot be primary or secondary in their own right, but when Wikipedia cites them as sources, then they can be primary or secondary sources. -- Tryptofish ( talk) 23:29, 27 April 2016 (UTC)
- In the last phrase I am telling not about sourcing, but about links. Links can not be primary or secondary. If other widely used resources do not see any copyright problems with linking to TCDB (and believe me, those are very serious resources supported by large staff including people who are familiar with copyright), I do not see any problems with linking to TCDB here.
My very best wishes (
talk)
23:12, 26 April 2016 (UTC)
- For that matter, there is nothing wrong with citing primary research papers from Saier et al., but again only subject to due weight. -- Tryptofish ( talk) 21:36, 26 April 2016 (UTC)
- I think that Jytdog did a very good job of explaining the various problems here, and I agree with him that we ought to try to find a happy outcome. I really am very concerned with the volume of Transporter Guy's edits, for multiple reasons. A very large number of pages have been affected, and there is a spammy quality to the edits, above and beyond the specific copyright issues. The external website seems to have been set up deliberately to promulgate the TCD (thus the licensing), but Wikipedia should not be a partner in it unless there is more in the way of acceptance by independent secondary sources first. I have a feeling that Transporter Guy has been put in a tough position by supervisors who want him to use Wikipedia for purposes that contradict our norms. There really is also a WP:UNDUE problem with much of this content. When one gets into the weeds of the source material, the database is really about genetic homology, but not about functional relationships. As phylogenetic information, that's fine. But ion channels should not be lumped together with ion transporters, for example. I'm not surprised that some editors have thanked Transporter Guy for the content, because, like the database website, it is mostly well-written. But that is all the more reason for editors to be concerned that we must not mislead our readers. I think that there is an awful lot of material that will have to be reverted or deleted, and I really hope that Transporter Guy will be able to just slow down while this gets sorted out. -- Tryptofish ( talk) 00:57, 25 April 2016 (UTC)
- I want to follow up on a point I made briefly above. There is a very widely recognized use of the abbreviation "Trk" to designate
Trk receptors and the like. Here is a PubMed search for "Trk", and it confirms what I just said:
[7]. The Saier lab TCDB website uses the abbreviation instead to designate K+ transporters. I don't see that at the IUBMB website, and I don't think I've seen it anywhere else. If we posit that the TCDB site reflects something more recent than what is at IUBMB and elsewhere, then I really have to question the degree to which we can regard the TCDB site as being peer-reviewed. Taking that observation along with the copying issues, I get the feeling that the TCDB site is probably being edited by junior people in the lab group, and is a sort of continuous work-in-progress (not unlike Wikipedia in that regard). It's not really a "final draft". Given that, and given
WP:CRYSTAL, I think that makes it a less reliable source than are sites such as IUBMB, and makes direct copying from TCDB to Wikipedia undesirable. --
Tryptofish (
talk)
22:24, 26 April 2016 (UTC)
- This isn't totally off-base - they seem to have just picked an exemplar of the family to name it after, and they picked TrkH. For whatever reason tropomyosin receptor kinases and one of the more common bacterial potassium uptake systems inconveniently converged on the same abbreviation. (Though I think the eukaryotes call theirs "track" and the prokaryotes "terk".) Opabinia regalis ( talk) 23:09, 26 April 2016 (UTC)
- Yes, it's just significantly off-base, but not totally (and it's tyrosine rather than tropomyosin). As long as they don't call it "twerk".
 But, seriously, does anyone else call the potassium transporters that? --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
But, seriously, does anyone else call the potassium transporters that? --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
- Yes, it's just significantly off-base, but not totally (and it's tyrosine rather than tropomyosin). As long as they don't call it "twerk".
- This TCDB abbreviation does not matter at all because the family appears as "The K+ Transporter (Trk) Family" in the database. There is no way anyone would think they are tyrosine kinases. Same with names used on WP pages. This is not a nomencalture of human genes or something else based on unique identifiers.
My very best wishes (
talk)
23:43, 26 April 2016 (UTC)
- Actually your argument does not matter at all, because an abbreviation is only useful in contexts where it does not appear with the full name. In any case, I think that it has become very clear that the TCDB website is less of a reliable source for Wikipedia's purposes than are either peer-reviewed papers or the websites that secondarily report on TCDB, and it is also clear that TCDB is one of multiple nomenclature systems, rather than the only one. --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
- Obviously, any combination of three letters has multiple meanings, as reflected on the corresponding disambig. page,
TRK. There is nothing else here, really.
My very best wishes (
talk)
17:02, 28 April 2016 (UTC)
- Gene sequence: Truckee Tahoe Airport. Anyway – I think that it has become very clear that the TCDB website is less of a reliable source for Wikipedia's purposes than are either peer-reviewed papers or the websites that secondarily report on TCDB, and it is also clear that TCDB is one of multiple nomenclature systems, rather than the only one. -- Tryptofish ( talk) 00:48, 29 April 2016 (UTC)
- Obviously, any combination of three letters has multiple meanings, as reflected on the corresponding disambig. page,
TRK. There is nothing else here, really.
My very best wishes (
talk)
17:02, 28 April 2016 (UTC)
- Actually your argument does not matter at all, because an abbreviation is only useful in contexts where it does not appear with the full name. In any case, I think that it has become very clear that the TCDB website is less of a reliable source for Wikipedia's purposes than are either peer-reviewed papers or the websites that secondarily report on TCDB, and it is also clear that TCDB is one of multiple nomenclature systems, rather than the only one. --
Tryptofish (
talk)
23:29, 27 April 2016 (UTC)
- People criticized contributions by Transporter Guy. After looking at this, I think he actually did very good work. Yes, there were certain copyright problems, but they are relatively minor, easily fixable and I think already fixed. Yes, there is some amount of over-linking and POV (preferential use of only one resource this user is most familiar with), but that can also be easily fixed if anyone cares. I do not. The resource he used (TCDB) is an excellent resource, and its use is highly beneficial for the project. I am telling that as someone with an expertise in this area. My very best wishes ( talk) 12:12, 29 April 2016 (UTC)
- I also checked editing by user Tryptofish on these pages and think he is occasionally engaged in WP:OR by claiming, for example, that KcsA potassium channel does not belong to the family of voltage-gated channels [8]. It is a page about specific protein family known as "voltage-gated channels" rather than about potential-sensitive proteins in general, which would be an entirely different subject. My very best wishes ( talk) 03:54, 4 May 2016 (UTC)
- I take very strong issue with the accusation that I have been violating NOR. In fact, there is an extended discussion about the proper definition of voltage-gated channels at
Talk:Voltage-gated ion channel, and I would very much welcome input from more editors there. (I'm not making anything up, but explicitly citing reliable sources such as a widely respected book by
Bertil Hille, and I understand that ion channels have to be voltage-gated in order to be voltage-gated ion channels. The comment above misrepresents what I said, by making it sound like I want to include other, weakly potential-sensitive, channels, which is the opposite of my position. On the other hand, the other editor wants to include channels that are not voltage-gated on the page about "voltage-gated channels".) --
Tryptofish (
talk)
18:22, 4 May 2016 (UTC)
- Some members of
protein family of "voltage-gated channels" (the subject of the page) are not voltage-gated because they lost or never had function typical for other members of the family. Same goes for other protein families. For example, some members of
G protein–coupled receptor family are not G protein coupled. @Boghog. Is not that correct?
My very best wishes (
talk)
20:08, 4 May 2016 (UTC)
- These exceptions may exist, but why on earth put them in the lead? Concentrate on why the family was defined in the first place. These definitions were originally based on function and later structure. Not the other way around.
Boghog (
talk)
20:18, 4 May 2016 (UTC)
- I definitely agree. That sounds reasonable. Sure. Except that function does not serve to "define" any protein families (there are numerous proteins with the same function which belong to different protein families and completely unrelated). What defines the family is evolutionary relatedness of the proteins. This is not much different from biological taxonomy: it is the common origin what defines the existence of "natural" (as opposed to the "artificial") taxonomic groups. My very best wishes ( talk) 20:20, 4 May 2016 (UTC)
- (
edit conflict) You continue to confound phylogenetics with the actual classification according to the preponderance of reliable sources. It's time to let other editors weigh in. --
Tryptofish (
talk)
20:26, 4 May 2016 (UTC)
- What I am talking about is the "actual", commonly accepted classification of proteins. So far you did not indicated any references or links to alternative (function-based) classifications of proteins.
Here is one of them, but it also places KcsA in the same family.
My very best wishes (
talk)
20:38, 4 May 2016 (UTC)
- Not true, and not true that I haven't provided references. Please, other editors: more eyes are needed at
Talk:Voltage-gated ion channel. --
Tryptofish (
talk)
21:08, 4 May 2016 (UTC)
- If this is not true, please explain what alternative general classifications of proteins are you talking about (authors and references/links)? I am talking about
UniProt (
here is KcsA, for example, and it belongs to the
family of potassium channels, most of which are voltage-gated),
Pfam (KcsA belongs to
this family which belongs to
this superfamily) and about
TCDB.
My very best wishes (
talk)
21:21, 4 May 2016 (UTC)
- You have asked me that same question multiple times before, and I have replied to it multiple times before. At this point, what we need are more editors to comment. --
Tryptofish (
talk)
22:15, 4 May 2016 (UTC)
- You responded by providing a reference that supports what I am telling: KcsA belongs to the family of voltage-gated channels [9]. My very best wishes ( talk) 17:06, 5 May 2016 (UTC)
- You have asked me that same question multiple times before, and I have replied to it multiple times before. At this point, what we need are more editors to comment. --
Tryptofish (
talk)
22:15, 4 May 2016 (UTC)
- If this is not true, please explain what alternative general classifications of proteins are you talking about (authors and references/links)? I am talking about
UniProt (
here is KcsA, for example, and it belongs to the
family of potassium channels, most of which are voltage-gated),
Pfam (KcsA belongs to
this family which belongs to
this superfamily) and about
TCDB.
My very best wishes (
talk)
21:21, 4 May 2016 (UTC)
- Not true, and not true that I haven't provided references. Please, other editors: more eyes are needed at
Talk:Voltage-gated ion channel. --
Tryptofish (
talk)
21:08, 4 May 2016 (UTC)
- What I am talking about is the "actual", commonly accepted classification of proteins. So far you did not indicated any references or links to alternative (function-based) classifications of proteins.
Here is one of them, but it also places KcsA in the same family.
My very best wishes (
talk)
20:38, 4 May 2016 (UTC)
- These exceptions may exist, but why on earth put them in the lead? Concentrate on why the family was defined in the first place. These definitions were originally based on function and later structure. Not the other way around.
Boghog (
talk)
20:18, 4 May 2016 (UTC)
- Some members of
protein family of "voltage-gated channels" (the subject of the page) are not voltage-gated because they lost or never had function typical for other members of the family. Same goes for other protein families. For example, some members of
G protein–coupled receptor family are not G protein coupled. @Boghog. Is not that correct?
My very best wishes (
talk)
20:08, 4 May 2016 (UTC)
- I take very strong issue with the accusation that I have been violating NOR. In fact, there is an extended discussion about the proper definition of voltage-gated channels at
Talk:Voltage-gated ion channel, and I would very much welcome input from more editors there. (I'm not making anything up, but explicitly citing reliable sources such as a widely respected book by
Bertil Hille, and I understand that ion channels have to be voltage-gated in order to be voltage-gated ion channels. The comment above misrepresents what I said, by making it sound like I want to include other, weakly potential-sensitive, channels, which is the opposite of my position. On the other hand, the other editor wants to include channels that are not voltage-gated on the page about "voltage-gated channels".) --
Tryptofish (
talk)
18:22, 4 May 2016 (UTC)
To state that KcsA belongs to the family of voltage gated channels is very misleading since it implies that KcsA is also voltage gated. This IMHO is unacceptable. It is acceptable to state that KcsA is evolutionarily related to voltage gated ion channels, but is not itself voltage gated. A fundamental flaw in the TCDB is that it does not clearly distinguish between structure and function. It is naming evolutionary related protein families after the predominate function of the majority of family members and this causes confusion when referring to members of this family that don't share this function. It would have been better to use function neutral names for these families, for example, seven-transmembrane domain family instead of GPCR. Boghog ( talk) 17:47, 5 May 2016 (UTC)
- All existing and widely accepted protein classifications ( UniProt, Pfam, InterPro, TCDB and others) tell that KcsA belongs to the family of voltage-gated channels (see links above). Please show me classifications that tell it does not belong to the family of voltage-gated channels. You may not like naming of protein families in the literature and classifications, but we should follow WP:Common name. And no, this naming is not a flaw of TCDB. It simply uses common name of the family from the literature. As I already explained, the naming of protein families is chosen to reflect most common/typical function of proteins in the family, but not all of them. Look, this is something obvious. What's the problem? My very best wishes ( talk) 18:26, 5 May 2016 (UTC)
- As I have already stated, if one writes that KcsA is a member of the voltage gated ion channel family without further explanation, this is seriously misleading.
Boghog (
talk)
19:00, 5 May 2016 (UTC)
- Boghog, thank you very much for supporting what I have been saying. Editors may find it useful to look at the diff that My very best wishes provided in replying to me ( [10]), in which I demonstrate the exact opposite of what My very best wishes claims. My very best wishes appears to be insisting that the only reliable sources are these non-peer-reviewed websites that classify proteins based on sequence and phylogeny, which leads to a predetermined outcome – and, when presented with reliable sources that say otherwise, My very best wishes pretends that they say what they do not say. We are unfortunately getting into WP:IDHT territory. -- Tryptofish ( talk) 21:25, 5 May 2016 (UTC)
- Boghog, thank you for supporting what I have been saying. My very best wishes ( talk) 21:36, 5 May 2016 (UTC)
- As I have already stated, if one writes that KcsA is a member of the voltage gated ion channel family without further explanation, this is seriously misleading.
Boghog (
talk)
19:00, 5 May 2016 (UTC)
- I think some people criticize TCDB simply because they are not familiar with the subject. Contrary to the claim by Tryptofish just above ("non-peer-reviewed websites"), not only these classification systems/resources are extensively peer-reviewed and republished many times, as anyone can check on WP pages about them, but they are highly cited in many hundreds, possibly thousands publications and used by many thousands researchers. In addition, repeatedly placing this tag into page indicates that the user apparently believes this is classification according only to TCDB. This is not the case. If someone looks at these proteins in UniProt, this will be essentially the same or very similar classification. As I already noted, classification on the family level in TCDB and all other systems is essentially the same. Yes, a number of superfamilies provided by TCDB can not be found in other systems, however they are usually justified based on publications by various research groups. In other cases, such as voltage-gated superfamily, they are practically the same. It also seems that Tryptofish criticizes the commonly accepted biological approach, according to which proteins (and other relevant biological entities, such as species) must be classified based on their phylogeny (i.e. common biological origin), which is precisely the kind of original research I am talking about. My very best wishes ( talk) 11:37, 6 May 2016 (UTC)
GPCRs as transporters???
@ Transporter Guy and My very best wishes: I find this very disturbing. This is a clear example of ref spam and undue weight. GPCRs are not transporters. Full stop. GPCRs transduce signals, not transport molecules. Any classification system that categorizes GPCRs as transporters is highly suspect. The TCDB seems to be a made up ad hoc classification system. I agree with Tryptofish that the TCDB is only one of several transporter classification systems. I also agree with Tryptofish there are significant copyright concerns with Transporter Guy's contributions. Boghog ( talk) 20:00, 4 May 2016 (UTC)
- No one tells that GPCR are transporters. This edit does not assume that GPCR are transporters. What other several transporter systems are you talking about? My very best wishes ( talk) 12:49, 5 May 2016 (UTC)
- TCDB is only called "transporter database". It includes a number of proteins that are not transporters. That's OK. This is only a name. The only relevant question in the edit by Transporter Guy above if it introduces correct linking to biological databases. If the linking is not correct, let's remove of fix it. If it was correct, the edit is fine. As far as I can see, that was correct linking. At the same time, I do agree that a number of links to TCDB by Transporter Guy was indeed excessive (as I already said above). No objection to removing some of these links. My very best wishes ( talk) 20:17, 4 May 2016 (UTC)
- (
edit conflict) Thank you, Boghog. Of course you are correct, that GPCRs are not transporter molecules, even though it is entirely valid to say that there are sequence homologies that reflect phylogenetic relationships. It seems that I am unable to convince My very best wishes of this, but ion channels are not transporters, and transporters are not ion channels. And ion channels that are not voltage-gated are not voltage-gated ion channels. There are indeed problems with ref spam and with undue weight, and there are also problems with what is becoming an increasingly tendentious unwillingness to drop the stick. --
Tryptofish (
talk)
20:26, 4 May 2016 (UTC)
- One of the reasons why TCDB includes GPCR is obvious: the family of microbial rhodopsins, which is closely related to GPCR (same superfamily), contains a number of proton and ion pumps, i.e. transporters. Ones again, proteins from the same family or the same superfamily frequently have very different biological functions. This is something well known. On the other hand, totally unrelated proteins may have the same biological function, even catalyze exactly the same chemical reaction [11]. My very best wishes ( talk) 01:41, 5 May 2016 (UTC)
AfC help?
Hi. There's a draft over at AfC that it would be a good thing if someone with a bit more knowledge of the subject took a look at: Draft:Chloroplast migration. Any comments/suggestions would be greatly appreciated. Onel5969 TT me 13:30, 7 May 2016 (UTC)
Gene Wiki data migration
@ Andrew Su, I9606, and Julialturner: I am posting here because this issue may be of general interest to the MCB community.
Previously in Gene Wiki articles, only a single accession code was generally displayed for UniProt (reviewed Swiss-Prot only) and RefSeq (experimental NM/NP only) entries. I noticed that the Gene Wiki Infoboxes now display additional UniProt TrEMBL entries as well as predicted XM/XP RefSeq entries (see for example, ETHE1). My question is whether multiple UniProt and RefSeq links should be displayed. My own opinion is that only the Swiss-Prot UniProt and experimental RefSeq links should be displayed if they exist because they are the highest quality and determining which of the several displayed links is the most relevant is not immediately apparent. If Swiss-Prot or RefSeq NM/NP entries do not exist, then it may be appropriate to display TrEMBL or XM/XP links. However I would be interested in hearing how others think, especially those maintaining this data. Cheers. Boghog ( talk) 10:19, 16 April 2016 (UTC)
- @ Boghog, Andrew Su, and Julialturner: We often see issues where there are multiple identifiers that seems equally relevant and, for the moment, went with the strategy of letting the user decide. If you have a set of logic that you think should be followed, it would be pretty easy to adapt the Lua script I think. Julia's talk page has a few examples together to make it easier to see. There, you can see cases where we have multiple NM prefixed RefSeq records for the same gene. I see 2 UniProt records for VIPR1 there, with one being C prefixed and one P prefixed. I guess we should only be showing the P uniprots as these seem to match up with SwissProt records ? As for maintaining the data, I think it makes sense to keep all potentially relevant data accessible in wikidata and to let the application (e.g. the infobox lua script) encode the logic to decide what it wants to display. -- Benjamin Good ( talk) 19:12, 18 April 2016 (UTC)
- @
I9606: Thanks Benjamin. Concerning RefSeq codes, the difference between experimental and predicted entries is very clear. The experimental entries have a NM/NP prefix while the predicted entries have XM/XP prefixes. I am having a difficult time tracking down a definitive definition of the UniProt accession codes. The closest that I can find is
here. It appears that the reviewed Swiss-Prot accession numbers start with the letters [O,P,Q] whereas the [A-N,R-Z] denote a unreviewed TrEMBL entry. Hence my suggestion is
- RefSeq – only display NM/NP prefixed entries if they exist, if not, then display the XM/XP entries
- UniProt – only display [O,P,Q] prefixed entries if they exist, if not, then display the [A-N,R-Z] entries
- Does this make sense? Boghog ( talk) 19:40, 18 April 2016 (UTC)
- yes, +1 --
Benjamin Good (
talk)
21:01, 18 April 2016 (UTC)
- @
I9606:@
Boghog: I should be able to implement these changes pretty easily. I could also display all the RefSeq and Uniprot, but sort so the most relevent data is at the top of the list.
Julialturner (
talk)
20:48, 18 April 2016 (UTC)
- I think selectively showing the most relevant is the way to go here. There is the potential to generate pretty long, uninformative lists if we show them all - unwieldy even if sorted. --
Benjamin Good (
talk)
21:01, 18 April 2016 (UTC)
- @
Boghog,
Andrew Su, and
I9606: The changes discussed above should now be reflected in those gene pages with the infobox_gene template. Only the most relevant items are displayed on the above criteria.
Julialturner (
talk)
18:19, 12 May 2016 (UTC)
- Thanks Julialturner! That is a big improvement. Boghog ( talk) 11:05, 14 May 2016 (UTC)
- @
Boghog,
Andrew Su, and
I9606: The changes discussed above should now be reflected in those gene pages with the infobox_gene template. Only the most relevant items are displayed on the above criteria.
Julialturner (
talk)
18:19, 12 May 2016 (UTC)
- I think selectively showing the most relevant is the way to go here. There is the potential to generate pretty long, uninformative lists if we show them all - unwieldy even if sorted. --
Benjamin Good (
talk)
21:01, 18 April 2016 (UTC)
- @
I9606:@
Boghog: I should be able to implement these changes pretty easily. I could also display all the RefSeq and Uniprot, but sort so the most relevent data is at the top of the list.
Julialturner (
talk)
20:48, 18 April 2016 (UTC)
- @
I9606: Thanks Benjamin. Concerning RefSeq codes, the difference between experimental and predicted entries is very clear. The experimental entries have a NM/NP prefix while the predicted entries have XM/XP prefixes. I am having a difficult time tracking down a definitive definition of the UniProt accession codes. The closest that I can find is
here. It appears that the reviewed Swiss-Prot accession numbers start with the letters [O,P,Q] whereas the [A-N,R-Z] denote a unreviewed TrEMBL entry. Hence my suggestion is
Bot implementing wiki-data driven infobox for genes
See this diff which was made by a bot, User:ProteinBoxBot. Just wanted to make sure you all were aware of this. Jytdog ( talk) 03:50, 20 May 2016 (UTC)
The article states that this deletion syndrome is a protein, which strikes me as odd. Also, there is already an article on the Chromosome 22q11.2 deletion syndrome, called DiGeorge syndrome. -- Arno Matthias ( talk) 17:46, 25 May 2016 (UTC)
- There are a fair amount of genes, loci and proteins that are named after a disease condition. Seems like this is the case here. Jo-Jo Eumerus ( talk, contributions) 18:48, 25 May 2016 (UTC)
- Thanks for the heads up. It is true that many gene/proteins are name after diseases. However in this particular case, DEL22Q11.2 is listed as a gene with unknown type. Hence it apparently is not known if this gene is coding or not. Therefore I have modified the text accordingly. Boghog ( talk) 19:00, 25 May 2016 (UTC)
- I agree with Boghog; it is an ill-understood genetic disorder, and after a brief web search I found no product coded. Cheers, BatteryIncluded ( talk) 19:34, 25 May 2016 (UTC)
Thank you all, but shouldn't there at least be a link to DiGeorge syndrome in the article? Or, if the are the same, only one article? -- Arno Matthias ( talk) 06:44, 26 May 2016 (UTC)
- Yes. I missed that part. The chromosome 22q11.2 deletion syndrome is the same as or causes the DiGeorge syndrome, so they should be merged. I'll go ahead. Cheers, BatteryIncluded ( talk) 13:45, 26 May 2016 (UTC)
(Also posted on project PHARMA)
User:Medgirl131 has recently created {{ Cytokine receptor modulators}} and {{ Nitric oxide signaling}} navboxes and has been adding these to a large number of articles. My major concern is that they combine endogenous proteins and recombinant protein drugs in one infobox which I think is confusing. Additional concerns that I have about these navboxes are their large size and that they violate WP:Bidirectional which makes navigation more difficult. I have highlighted my concerns on their talk page here. What do others think about these navboxes? Boghog ( talk) 06:03, 28 May 2016 (UTC)
Auto-assessment of article classes
Following a recent discussion at WP:VPR, there is consensus for an opt-in bot task that automatically assesses the class of articles based on classes listed for other project templates on the same page. In other words, if WikiProject A has evaluated an article to be C-class and WikiProject B hasn't evaluated the article at all, such a bot task would automatically evaluate the article as C-class for WikiProject B.
If you think auto-assessment might benefit this project, consider discussing it with other members here. For more information or to request an auto-assessment run, please visit User:BU RoBOT/autoassess. This is a one-time message to alert projects with over 1,000 unassessed articles to this possibility. ~ Rob Talk 01:16, 4 June 2016 (UTC)
New map of Metabolic Pathways


I've made a new metro-style map of the major metabolic pathways, which was inspired by the already existing one ( Template:Metabolic_pathways) and other excellent online resources (e.g. [12], [13]). The map encompasses most of the biochem pathways in human, but also include important ones in plants / fungi / microbes (double lines). The metabolites are grouped as general terms (e.g. "Hexose Phosphates") instead of the specific ones in the existing map. I'm also going to make it a clickable map.
Some questions:
- Is it suitable for wikipedia, in terms of style and complexity?
- Is it good to replace the existing map with this one (that would change all the pages that includes Template:Metabolic_pathways) or make it a separate work?
- Any comment on the map as an individual work?
Thanks! (also posted at Wikipedia:WikiProject_Molecular_and_Cell_Biology/Metabolic_Pathways_task_force/proposals#New_map_of_Metabolic_Pathways) Chakazul ( talk) 17:30, 30 May 2016 (UTC)
- Great work
Chakazul! I'm a big fan of clear diagrams. I would say that it's an improvement over the image currently used in
Template:Metabolic_pathways. If you're interested, I'd be happy to help you out with
{{ annotated image 4}}to convert it into an interactive diagram (like Template:Glycolysis summary). I have a few suggestions for improvements:- It doesn't really need a title within the image itself (since it'll have a caption etc)
- The legend seems to be spread all around the sides currently. I realise it was for aesthetic reasons to best use the available space, however I think it aids readability to have it all together somewhere.
- Do the node sizes have any significance? How about the overlapping nodes? Again, either of these may be useful to mention in the legend
- Overall, a great diagram, and I'm sure it'll find wide use around Wikipedia. T.Shafee(Evo﹠Evo) talk 00:57, 31 May 2016 (UTC)
- Great work
Chakazul! I'm a big fan of clear diagrams. I would say that it's an improvement over the image currently used in
Template:Metabolic_pathways. If you're interested, I'd be happy to help you out with
- Thanks
Evolution and evolvability! It would be great if you can help to make it interactive. There seems no way to extract the bounding box information form SVG, so need to set the clickable areas one-by-one?
- Let me try make a version with one single legend (I was trying to avoid the use of legend, though)
- Smaller node means one kind of metabolite (e.g. Acetyl-CoA) and a larger one means a group (e.g. Amino Acids). Double nodes allow both of the terms like "Glycoproteins & Proteoglycans" clickable, clicking the left node will link to Glycoprotein and clicking the right one will link to Proteoglycan.
- Chakazul ( talk) 10:27, 31 May 2016 (UTC)
- Thanks
Evolution and evolvability! It would be great if you can help to make it interactive. There seems no way to extract the bounding box information form SVG, so need to set the clickable areas one-by-one?
- @
Chakazul: You could do a version with no legend at all in the image (I actually didn't put any in
Template:Glycolysis summary), and just write a text caption. The
{{ annotated image 4}}template only allows adding wikilinked text over the image. Adding click-able field shapes / image maps tends to be trickier. T.Shafee(Evo﹠Evo) talk 10:38, 31 May 2016 (UTC)
- @
Chakazul: You could do a version with no legend at all in the image (I actually didn't put any in
Template:Glycolysis summary), and just write a text caption. The
- @ Evolution and evolvability: There's a no-legend version above. Yes, image map is troublesome, links overlay would be more suitable in wikipedia. I wonder if the links can be transparent, otherwise I need to remove all the texts. Chakazul ( talk) 18:00, 31 May 2016 (UTC)
- @ Chakazul: The no-legend version look good! I'm afraid you're correct that you'll have to make a no-text version. It's a bother. It's possible to make the wikilink text any colour, but not transparent (as far as I know). I recommend making a no-text version only once you're certain that you're happy with the layout etc sp that you only have to do it once. On that note, it might be worth drafting the caption now. I think it'd be best to make a separate new template rather than replace the image in Template:Metabolic_pathways, so that the older image is kept as an archive. This page lists every article that transcludes Metabolic_pathways and can have the new "Template:metabolic metro" added (or whatever you decide to call it). T.Shafee(Evo﹠Evo) talk 00:00, 1 June 2016 (UTC)
- There actually is a way to create a transparent wikilink/image map-type hyperlinks using
{{ Annotated image 4}}, but it'd probably be a little more tedious to set up relative to annotating text onto an image without pre-existing text. Basically, all you have to do is create a piped wikilink to a bunch of non-breaking spaces (produced via the text ). Adding a space between non-breaking spaces will take up less room in the source code (i.e., use instead of ). As an example, the next line contains an invisible link to the MCB project page and the line after that contains an invisible link to a redirect to this page:
← link is here
← link is here
When you mouse over it, you should see the blue underlining from the HTML link. With the annotated image template, you'd need to adjust the number of non-breaking spaces so that it spans the full length of the word(s) that you want to link to. That may require some trial and error with the number of non-breaking spaces for each term that you want to link to get the length right. Seppi333 ( Insert 2¢) 03:35, 1 June 2016 (UTC)
Edit: I just noticed that the diagram has line breaks in the image text. If invisible wikilinks are annotated onto the diagram, some of the annotated wikilinks would need to have<br />included in the piped link when the text in the diagram is on different lines.
For example,[[wikitext|this wikilink<br />spans several lines via 2 forced<br />line breaks.]]produces: this wikilink
spans several lines via 2 forced
line breaks.
It works the same way if you replace the letters with non-breaking spaces. Seppi333 ( Insert 2¢) 04:07, 1 June 2016 (UTC)
- There actually is a way to create a transparent wikilink/image map-type hyperlinks using
@ Evolution and evolvability and Seppi333: Thanks for the suggestions! Here is a trial version of the template: User:Chakazul/sandbox.
The first one is the diagram without text, added two wikilinks for testing; the second one is the original diagram, using Seppi's method. Both methods have their own merits. Style 1 is a bit easier to match the positions, and can show the availability of the wiki page (blue or red). Style 2 gives more control on the color (most texts are in black BTW), but matching the white spaces needs more work. So Evo&Evo it's up to you to choose which method for annotation. Chakazul ( talk) 18:56, 1 June 2016 (UTC)
Beautiful! A few suggestions:
- The grey boxes are invisible when I face straight towards my monitor. I can only see them if I look from an angle. Maybe make them a bit darker, or put a thin black line around them.
- I find the cellular respiration colour difficult to distinguish from the carbohydrate metabolism and messengers colours. (I can distinguish those two, though.)
- Given that the "feeders to gluconeogenesis" tree is large and convoluted, it might be helpful to dedicate dark blue solely to gluconeogenesis and its feeders, and use a different colour(s) for glycogenesis etc.
- In a few places you've written NADH2 instead of NADH, or NADPH2 instead of NADPH.
- Maybe move MVA to the left a bit to separate it from NADPH from the PPP.
- Maybe write "Energy Metabolism" in black or indicate it with a box or bracket or something, because I'd been looking for arrows of the corresponding colour in the map.
- I'm not sure why there's a red line between pyruvate and lactate, as this reaction is not part of glycolysis.
Awesome work!
![]() Adrian J. Hunter(
talk•
contribs)
14:08, 31 May 2016 (UTC)
Adrian J. Hunter(
talk•
contribs)
14:08, 31 May 2016 (UTC)
- @
Adrian J. Hunter: Thanks for the comments! New version is updated.
- I decided to make glycogenesis / glycogenolysis darker colors. The blue and red lines from lactate to pyruvate (and also those from glycerol to triose phosphates i.e. DHAP & GAP) are technically side ways entering gluconeogenesis and glycolysis, so I make them thin lines much like the "feeders".
- The NADH2 (also NADPH2) were meant to be NADH2+ as sometimes seen on other diagrams as a shorthand of NADH+H+, but that would be confusing, I changed them to NADH & NADPH.
- Chakazul ( talk) 18:00, 31 May 2016 (UTC)
- An additional link that I find helpful is colorbrewer2.org. If you select "qualitative" as your colourscheme, it will give you a set of colours (up to 12) that are the maximally easy to tell apart. I've started to use it since I'm colour-blind so often pick odd colour palettes otherwise! T.Shafee(Evo﹠Evo) talk 08:24, 1 June 2016 (UTC)
- @
Adrian J. Hunter: The color scheme link is very useful! Too bad that they don't have colorblind safe scheme with more than 4 colors. A few of my friends are colorblind (I already guessed you're one) so I often have colorblind-friendliness in mind, but I always wanted to make my diagrams colorful and that became a dilemma.
Chakazul (
talk)
18:56, 1 June 2016 (UTC)
- I think this comment was meant for
Evolution and evolvability :-). Changes look good to me.
Adrian J. Hunter(
talk•
contribs)
12:03, 2 June 2016 (UTC)
- Yes I meant Evo&evo. Chakazul ( talk) 17:15, 2 June 2016 (UTC)
- I think this comment was meant for
Evolution and evolvability :-). Changes look good to me.
Adrian J. Hunter(
talk•
contribs)
12:03, 2 June 2016 (UTC)
- @
Adrian J. Hunter: The color scheme link is very useful! Too bad that they don't have colorblind safe scheme with more than 4 colors. A few of my friends are colorblind (I already guessed you're one) so I often have colorblind-friendliness in mind, but I always wanted to make my diagrams colorful and that became a dilemma.
Chakazul (
talk)
18:56, 1 June 2016 (UTC)
Template:Metabolic_metro now created. My sandbox can be used for drafting. @ Evolution and evolvability:. Chakazul ( talk) 17:15, 2 June 2016 (UTC)
- @
Chakazul: The template's looking great! It would be great to see sub-section versions in the same style of it for some of the related pages.
T.Shafee(Evo﹠Evo)
talk
05:31, 11 June 2016 (UTC)
- @ Evolution and evolvability: Thanks for the color dots. I've corrected the color of blue dot from violet to blue (they looks similar?). Now see if it's suitable to use this template in other pages.
- I do think of making detailed version for individual pathways, or other language versions. Chakazul ( talk) 08:25, 11 June 2016 (UTC)
| Major metabolic pathways | |
|---|---|
|
Flow-through test - article up for speedy deletion
Esteemed members of this project may be best placed to work out whether the speedy deletion tag on Flow-through test should stay or go ... the subject matter appears to be within your sphere of interest. Grateful if someone would take a look (preferably before it gets speedied) - it's an article which popped up on the Wikipedia:Conflict of interest/Noticeboard. thanks -- Tagishsimon (talk) 00:53, 30 June 2016 (UTC)
Conversation on controversial changes to epistasis article being discussed on the WP:GEN talkpage. T.Shafee(Evo﹠Evo) talk 00:28, 4 July 2016 (UTC)
Can someone with some experience help double-check on FAM163A? The title is FAM163A but the infobox is FAM136A. It seemed like it was previously clear with this version but this edit flipped the name but not the infobox and changed none of the citations which is mighty odd. Does a history split need to be done and other fixes? See also User:Flameep/sandbox. -- Ricky81682 ( talk) 03:12, 14 July 2016 (UTC)
- I think this is OK. The addition of the FAM136A infobox to FAM163A was a mistake that was later corrected in this . FAM163A also contained some original research which I have removed. Boghog ( talk) 06:13, 14 July 2016 (UTC)
WikiFactMine: Proposed Wiki Grant for biological Wikidata
We are proposing a Wikimedia Project Grant (WikiFactMine) which will automatically scan the daily peer-reviewed scientific literature (up to 10000 articles /day) and extract biological entities (definable in Wikidata). We concentrate on genes and species which have high precision and recall, but with Wikidata-based dictionaries this can be extended to other well defined entities (e.g. cell types or diseases). These, with their citations, are then offered to Wikidata editors for potential inclusion/update. These can also alert Wikipedia editors to new citations. The project includes a Wikimedian in residence in the University of Cambridge. We'd be grateful for comments, endorsement and offers of help. Petermr ( talk) 10:37, 2 August 2016 (UTC)
Gasdermin
Hi... can anyone start writing about Gasdermins? -- محمود ( talk) 21:50, 26 August 2016 (UTC)
- I just created
GSDMA,
GSDMB,
GSDMC, and
GSDMD. Anything in particular you would like to know about Gasdermin?
Boghog (
talk)
22:07, 26 August 2016 (UTC)
- @ Boghog: That's great. Actually, I was editing in some articles and catched Gasdermin D in Inflammasome article. I thought that it's something deserve an article in Wikipedia -- محمود ( talk) 22:34, 26 August 2016 (UTC)
This project's feedback would be appreciated in this discussion, as this could greatly (and positively) affect biological citations! Headbomb { talk / contribs / physics / books} 21:53, 7 September 2016 (UTC)
There is a redirect discussion of which members of this project could help.
Wikipedia:Redirects_for_discussion/Log/2016_October_2#Misfolded is ongoing. We are not sure whether "misfold" is a purely biochemical term or if it could WP:SURPRISE people who were looking for info about folding blankets. Your input would be much appreciated.-- Mr. Guye ( talk) 19:54, 2 October 2016 (UTC)
Portal talk:Molecular and cell biology move discussion
- See move discussion at Portal talk:Molecular and cell biology#Requested move 20 October 2016. Anthony Appleyard ( talk) 20:41, 20 October 2016 (UTC)
review an article
Can someone review/assess the Phosphatidylserine article? I hope this is the correct place to request this. VeniVidiVicipedia ( talk) 12:56, 25 October 2016 (UTC)
Can someone review/assess the Cyclopentenone prostaglandins article? The article has recently been greatly expanded. Although it is currently categorized as a pharmacy and pharmacology article, it may be better categorized in the Molecular and Cellular biology section. Very similar prostaglandins are so categorized (e.g. see prostaglandin). Suggestions on its improvement would be appreciated. Also, the article as currently formatted is correctly redirected from 15-deoxy-Δ12,14-prostaglandin J2 (15-d-Δ12,14-PGJ2), a principal cycloentenone prostaglandin. Is it possible to similarly redirect it from other cyclopenentone prostaglandins viz., Δ12-PGJ2, PGJ2, PGA2, and PGA1, discussed in the article (but not given separate Wikipedia pages elsewhere) and, if so, how do I do that? Thanks, ( User talk:joflaher) 12:01 P.M. 29 Oct 2016.
A request for comment has been made at the above link. Your input is welcome. Boghog ( talk) 11:50, 30 October 2016 (UTC)
CRISPR article status
I noticed in the CRISPR article the following sentence in the lead section: "The CRISPR interference technique has many potential applications, including altering the germline of humans, animals, and food crops." My background is chemistry and I am confident that that statement is incorrect. That can be a very time-consuming error for the reader to sort out. I think the article should mention something about the Cas9-gRNA complex, since it is the synthetic gRNA that makes it into a gene editing tool and a Breakthrough. The press just calls it "CRISPR", but we should try to sort it out for the reader. I think the article should be split, splitting out the engineering technology from the native phenomena (that is some work). It would be nice to note that Cas9 corresponds to CAS III in the diagram at the top of the page. See [14] for some details. Can any of you experts help out?-- 172.56.0.153 ( talk) 09:12, 29 October 2016 (UTC)
- These sources [1] [2] [3] make it very clear that germline modifications with CRISPR is possible and in fact has already successfully carried out in animals. [4] Guide RNA (gRNA) is already mentioned several times in the article. Boghog ( talk) 09:31, 29 October 2016 (UTC)
References
- ^ Evitt NH, Mascharak S, Altman RB (2015). "Human Germline CRISPR-Cas Modification: Toward a Regulatory Framework". The American Journal of Bioethics : AJOB. 15 (12): 25–9. doi: 10.1080/15265161.2015.1104160. PMC 4699477. PMID 26632357.
- ^ Porteus M (2016). "Genome Editing: A New Approach to Human Therapeutics". Annual Review of Pharmacology and Toxicology. 56: 163–90. doi: 10.1146/annurev-pharmtox-010814-124454. PMID 26566154.
- ^ Strong A, Musunuru K (2016). "Genome editing in cardiovascular diseases". Nature Reviews. Cardiology. 14 (1): 11–20. doi: 10.1038/nrcardio.2016.139. PMID 27609628. S2CID 3157463.
- ^ Mou H, Kennedy Z, Anderson DG, Yin H, Xue W (2015). "Precision cancer mouse models through genome editing with CRISPR-Cas9". Genome Medicine. 7 (1): 53. doi: 10.1186/s13073-015-0178-7. PMC 4460969. PMID 26060510.
---Reply---
- SIGH. PLEASE pay attention. The problematic sentence starts out "The
CRISPR interference technique ....." The CRISPR INTERFERENCE technique is a variation on the "CRISPR technique". It uses dCas9 (dead/deactivated Cas9) to reduce gene expression, NOT to achieve genome editting. Normal Cas9 goes to the genome loci and CUTS while dCas9 goes to the same loci and just sits there, thus "interfering" with gene translation/expression.--
172.56.32.19 (
talk)
13:03, 29 October 2016 (UTC)
- Your original post didn't clearly state what the problem was nor why it was a problem. You only stated that you were confident that it was a problem. The immediate problem that you raised is easily solved by a simple edit. {{
sofixit}}.
Boghog (
talk)
13:38, 29 October 2016 (UTC)
- I believe the OP is the sock of an indeffed user; see
Talk:CRISPR#SPLIT
Jytdog (
talk)
17:22, 29 October 2016 (UTC)
- I was asking for the assistance of experts, not Essjays. Sigh. So the article will never be useful.-- 172.56.32.19 ( talk) 17:34, 29 October 2016 (UTC)
- I believe the OP is the sock of an indeffed user; see
Talk:CRISPR#SPLIT
Jytdog (
talk)
17:22, 29 October 2016 (UTC)
- Your original post didn't clearly state what the problem was nor why it was a problem. You only stated that you were confident that it was a problem. The immediate problem that you raised is easily solved by a simple edit. {{
sofixit}}.
Boghog (
talk)
13:38, 29 October 2016 (UTC)
- SIGH. PLEASE pay attention. The problematic sentence starts out "The
CRISPR interference technique ....." The CRISPR INTERFERENCE technique is a variation on the "CRISPR technique". It uses dCas9 (dead/deactivated Cas9) to reduce gene expression, NOT to achieve genome editting. Normal Cas9 goes to the genome loci and CUTS while dCas9 goes to the same loci and just sits there, thus "interfering" with gene translation/expression.--
172.56.32.19 (
talk)
13:03, 29 October 2016 (UTC)
![]() Done Not sure what the history is here. But made the more specific requested change. Thanks for pointing that out!
Ajpolino (
talk)
21:30, 29 October 2016 (UTC)
Done Not sure what the history is here. But made the more specific requested change. Thanks for pointing that out!
Ajpolino (
talk)
21:30, 29 October 2016 (UTC)
- Thank you, Ajpolino. Maybe someday the Essjays of this site will lose interest in using that article to chase persons and the article might reach Good Article status. Someday.--
172.56.1.126 (
talk)
04:18, 31 October 2016 (UTC)
- As you, not me are claiming expertise in this area, why didn't you fix it yourself with a clear edit summary? Your original post was clear as mud. Your second post was much clearer. Boghog ( talk) 07:35, 31 October 2016 (UTC)
On a related note, given that it is one of the most viewed articles within this project's scope, I was thinking that it might be a good candidate for doing some sort of joint publication, where we make a concerted effort to bring it up to high-standard and then submit to an academic journal for peer review. The lure of a possible publication could entice some academics with greater experience in the area. T.Shafee(Evo&Evo) talk 06:18, 31 October 2016 (UTC)
Should this redlink be redirected to microfilament, actin, cytoskeleton, or related article, or does it merit its own independent article? I can't really tell since I'm not that familiar with the topic. There's 4 incoming links from articles to that redlink. Seppi333 ( Insert 2¢) 04:15, 11 November 2016 (UTC)
- It appears
microfilament (cytoskeletal fibers formed by polymerization of monomeric globular (G) actin) is synonymous with actin cytoskeleton.
Boghog (
talk)
04:59, 11 November 2016 (UTC)
- Actin is a protein; cytoskeleton is a structure made of actin. Does that help? Cheers, BatteryIncluded ( talk) 05:22, 11 November 2016 (UTC)
- I guess I'll redirect it to microfilament; I figured that one was the most relevant article. Seppi333 ( Insert 2¢) 05:36, 11 November 2016 (UTC)
Microscopy help
Hey folks! Super-resolution microscopy has had a "clarification-needed" tag for about a year now in the section on Structured illumination microscopy. I find I'm a bit out of my depth on this one. Any chance someone with a bit more experience in this kind of thing could take a peek and clarify the paragraph in question? Thanks a bunch! Ajpolino ( talk) 17:38, 9 November 2016 (UTC)
- It has to do with Phase (waves)#Phase shift on 2 superimposed images. However, I am unable to explain it to Wikipedia standards. I just added an intralink to Phase (waves) and left the 'clarify' tag intact. BatteryIncluded ( talk) 05:56, 11 November 2016 (UTC)
Coverage of God gene hypothesis in the VMAT2 article
This has come up before on the talk page ( Talk:VMAT2#God Gene does not help understanding of VMAT2 function), but I figured I'd raise the issue here to attract more input. God gene hypothesis is a fringe theory that suggests that a neurotransmitter transporter which is located on synaptic vesicles in monoamine neurons ( VMAT2) has some form of significant role in spirituality. I've removed content which covers the god gene hypothesis from the VMAT2 article twice now - once recently and once a while ago - per WP:ONEWAY. The question I have for you all is this: should this article cover a fringe theory because the theory is covered in popular culture? Please respond here if you have any input: Talk:VMAT2#Re: covering the God gene hypothesis. Seppi333 ( Insert 2¢) 19:30, 18 November 2016 (UTC)
Proposed merger of Neto 1 and NETO1
I have proposed that the article Neto 1 be merged into NETO1 as both articles appear to be on the same topic and "NETO1" is currently a more thorough discourse of the topic. I have created a proposed merger Talk Space on the NETO1 page and listed what I need help with.
I am not a scientist, biologist, etc., so I am not sure which article title is correct. I would appreciate any pertinent information that you can provide. If someone can assist with rewriting the article after the merger is complete that would be appreciated. Currently both articles are lacking substance and incoming links. I'm being pretty bold proposing a merger after being on the site for less than two months, so your patience while I learn more about editing on Wikipedia is appreciated.
Thanks for your help! Zcarstvnz ( talk) 22:50, 20 November 2016 (UTC)
Splitting endogenous molecules used as drugs
See Wikipedia_talk:WikiProject_Medicine#Splitting_articles_about_endogenous_molecules_used_as_drugs Jytdog ( talk) 23:04, 22 November 2016 (UTC)
maybe i have missed it, but i have never found a page that gives an overview of reagents. I just added a tiny bit at that article but it is a chicken scratch. Is there a master artcle somewhere that talks about reagents and their importance? Jytdog ( talk) 06:05, 23 November 2016 (UTC)
- Not as far as I know. I looked about for one back when I updated substrate (chemistry) but didn't find anything sensible then. T.Shafee(Evo&Evo) talk 00:53, 24 November 2016 (UTC)
A request for comment has been made at the above link. Your input is welcome. Boghog ( talk) 16:45, 27 November 2016 (UTC)
Trivial naming question
Trivial difference that's been bothering me: Translation (biology) vs Transcription (genetics). Would it make sense to harmonise them both to XYZ (genetics)? I can't imagine it actually causes any confusion, it just seems untidy. T.Shafee(Evo&Evo) talk 08:47, 6 November 2016 (UTC)
- Good eye! I had never noticed that. While I agree this isn't super critical, for what it's worth my vote is harmonizing to XYZ (biology). My guess is that's more in line with how the average person would think of it.
Ajpolino (
talk)
15:51, 6 November 2016 (UTC)
- Wow, I never noticed that either. Yep, makes sense to make them parallel; don't really care which one we keep.
Opabinia regalis (
talk)
20:56, 6 November 2016 (UTC)
- I'm also learning towards XYZ (biology). They're as relevant to molecular biology as to genetics, and they're things students learn about in general biology courses, rather than specialist genetics courses.
Adrian J. Hunter(
talk•
contribs)
03:19, 7 November 2016 (UTC)
- Actually, I agree that "(biology)" is the sensible disambig, seeing as there are no other biological interpretations of "translation". T.Shafee(Evo&Evo) talk 02:56, 11 November 2016 (UTC)
- I'm also learning towards XYZ (biology). They're as relevant to molecular biology as to genetics, and they're things students learn about in general biology courses, rather than specialist genetics courses.
Adrian J. Hunter(
talk•
contribs)
03:19, 7 November 2016 (UTC)
- Wow, I never noticed that either. Yep, makes sense to make them parallel; don't really care which one we keep.
Opabinia regalis (
talk)
20:56, 6 November 2016 (UTC)
2016 Community Wishlist Survey Proposal to Revive Popular Pages

Greetings WikiProject Molecular Biology/Molecular and Cell Biology/Archive 7 Members!
This is a one-time-only message to inform you about a technical proposal to revive your Popular Pages list in the 2016 Community Wishlist Survey that I think you may be interested in reviewing and perhaps even voting for:
If the above proposal gets in the Top 10 based on the votes, there is a high likelihood of this bot being restored so your project will again see monthly updates of popular pages.
Further, there are over 260 proposals in all to review and vote for, across many aspects of wikis.
Thank you for your consideration. Please note that voting for proposals continues through December 12, 2016.
Best regards, Stevietheman — Delivered: 18:04, 7 December 2016 (UTC)
This template has surprisingly few fields. I wanted to add alt names to Histidine—tRNA ligase and the only place I could do it was in the "name" field which seems like clutter. Shouldn't this have most everything in Template:Infobox protein? Jytdog ( talk) 06:18, 12 December 2016 (UTC)
- The enzyme infobox is meant to represent information about an
Enzyme Commission number (EC number). Strictly speaking, an EC number represents the reaction catalyzed by an enzyme, not the enzyme itself and therefore there is not a strict one to one relationship between EC numbers and proteins. Several structurally unrelated proteins may catalyze the same reaction and a single protein may catalyze more than one reaction. Hence several otherwise unrelated proteins may be assigned the same EC number and a single protein may be assigned more than one EC number. Hence most of the fields in {{
infobox protein}} are not appropriate for {{
infobox enzyme}} and vice versa. I agree that
|AltNames=might be useful, hence I have added this as an optional parameter to the template. Boghog ( talk) 06:58, 12 December 2016 (UTC)- Thanks! Hm! So if the enzyme box is for a reaction, then shouldn't there always be a) a protein box; and b) as many enzyme boxes as there are reactions that the protein is involved in? Hm.
Jytdog (
talk)
07:10, 12 December 2016 (UTC)
- The rule that I have been following is if there is a strict one-to-one relationship between EC number and human gene/protein, then there should be a single article that covers both the gene and the enzyme. If there is not a one-to-one relationship, it is much cleaner to have separate gene and enzyme articles. Due to their large size, it is not practical to place more than one {{ Infobox gene}} in the same article. Because {{ Infobox protein}} is more compact, sometimes more than one of these is placed in articles about protein families. Finally, the SOP that WP:MCBMOS and Gene Wiki follow is one gene, one article. Boghog ( talk) 07:35, 12 December 2016 (UTC)
- Ideally
|ECnumber=in {{ Infobox protein}} should be able to list more multiple EC numbers. I will look into this. Boghog ( talk) 07:40, 12 December 2016 (UTC)- I am really clueless here. :) When do you use infobox: gene vs infobox: protein? Sorry to bother you but neither template says when you use one vs the other.
Jytdog (
talk)
07:48, 12 December 2016 (UTC)
- It depends on what is the scope of the article. If there is a one-to-one relationship between the gene and the enzyme, then use both infoboxes in same article. If there are several genes that encode proteins with the same enzymatic activity, focus on enzymology and use {{ Infobox enzyme}}. If there are a limited number of genes associated with a specific enzymatic activity, it also might make sense to include a limited number of {{ Infobox protein}} as well (see for example Trypsin). If there is a single protein with more than one enzymatic activity, then {{ Infobox protein}} or {{ Infobox gene}} listing the multiple EC numbers would make sense. Basically it comes down to common sense and editorial judgement. To give you some background, most of these enzyme and gene articles were created independently of each other. The enzyme articles systematically by a bot run by User:WillowW and the protein articles by User:ProteinBoxBot. Some of these have since been merged where it made sense. Boghog ( talk) 12:04, 12 December 2016 (UTC)
- I am really clueless here. :) When do you use infobox: gene vs infobox: protein? Sorry to bother you but neither template says when you use one vs the other.
Jytdog (
talk)
07:48, 12 December 2016 (UTC)
- Thanks! Hm! So if the enzyme box is for a reaction, then shouldn't there always be a) a protein box; and b) as many enzyme boxes as there are reactions that the protein is involved in? Hm.
Jytdog (
talk)
07:10, 12 December 2016 (UTC)
Featured article nomination for beta-Hydroxy beta-methylbutyric acid
I recently renominated this article at FAC ( Wikipedia:Featured article candidates/Beta-Hydroxy beta-methylbutyric acid/archive2) and was wondering if anyone from this WikiProject would be interested in doing a review of this article's pharmacology content. Every statement in this section is supported by at least 1 WP:MEDRS-quality review.
Since this compound's immediate biomolecular targets aren't known, there's a large amount of material on its downstream signaling pathways (primarily through mTORC1 and its interactions with the ubiquitin-proteasome system) in the pharmacodynamics section; this content is mostly based upon clinical trials in humans, but there's some contextual information on its signaling mechanisms which comes from in vitro and animal studies (I've clearly indicated what statements are supported by human in vivo research vs in vitro or animal research in the article text). The pharmacokinetics section covers its biosynthetic and metabolic pathways in humans. Consequently, it would be ideal for someone with a pharmacology and/or molecular biology background to take on a review of this material.
I would really appreciate it if someone from this WikiProject would take on a review of this article content at FAC. Seppi333 ( Insert 2¢) 20:42, 25 October 2016 (UTC)
- Still need a reviewer for this content; the current FAC nomination is located at Wikipedia:Featured article candidates/Beta-Hydroxy beta-methylbutyric acid/archive3. Seppi333 ( Insert 2¢) 17:48, 1 January 2017 (UTC)
Missing topics list
My list of missing topics about metabolism is updated - Skysmith ( talk) 15:11, 8 January 2017 (UTC)
User needs help with new article with DYK potential
Hi there. A new user created testosterone-cortisol ratio through AfC and requested some help improving it. I did a copyedit and formatted some references but have no knowledge about the subject itself (or molecular biology for that matter). Maybe someone here can help this user out? The topic sounds interesting enough to be featured in T:DYK if the article were expanded a bit more. Regards So Why 13:17, 17 January 2017 (UTC)
- oy content is almost entirely WP:Biomedical information sourced to refs that fail WP:MEDRS. Am not sure this article will survive. Will work on it. Jytdog ( talk) 20:23, 17 January 2017 (UTC)
WikiJournal of Medicine promotion

 |
The WikiJournal of Medicine is a free, peer reviewed academic journal which aims to provide a new mechanism for ensuring the accuracy of Wikipedia's biomedical content. We started it as a way of bridging the Wikipedia-academia gap. [1] It is also part of a WikiJournal User Group with other WikiJournals under development. [2] The journal is still starting out and not yet well known, so we are advertising ourselves to WikiProjects that might be interested. |
Engaging Wikipedians
- Original articles on topics that don't yet have a Wikipedia page, or only a stub/start
- Wikipedia articles that you are willing to see through external peer review (either solo or as in a group, process analogous to GA / FA review)
- Image articles, based around an important medical image or summary diagram
Engaging non-Wikipedians
We hope that an academic journal format may also encourage non-Wikipedians to contribute who would otherwise not. Therefore, please consider:
- Printing off the advertisement poster and distribute in tearooms & noticeboards at your place of work
- Emailing around the pdf through contact networks or mailing lists (suggested wording)
If you want to know more, we recently published an editorial describing how the journal developed. [3] Alternatively, check out the journal's About or Discussion pages.
- ^ Masukume, G; Kipersztok, L; Das, D; Shafee, T; Laurent, M; Heilman, J (November 2016). "Medical journals and Wikipedia: a global health matter". The Lancet Global Health. 4 (11): e791. doi: 10.1016/S2214-109X(16)30254-6. PMID 27765289.
- ^ "Wikiversity Journal: A new user group". The Signpost. 2016-06-15.
- ^ Shafee, T; Das, D; Masukume, G; Häggström, M (2017). "WikiJournal of Medicine, the first Wikipedia-integrated academic journal". WikiJournal of Medicine. 4. doi: 10.15347/wjm/2017.001.

Additionally, the
WikiJournal of Science is just starting up under a similar model and looking for contributors. Firstly it is seeking editors to guide submissions through external academic peer review and format accepted articles. It is also encouraging submission of articles in the same format as Wiki.J.Med. If you're interested, please come and discuss the project on the
journal's talk page, or the
general discussion page for the WikiJournal User group.
T.Shafee(Evo&Evo)
talk
10:33, 19 January 2017 (UTC)
Orphaned Gene/Protein Articles
Hi, I've been going through the backlog at CAT:ORPHAN lately, and I keep running across articles like EAF1, E2F7, etc etc for various genes and proteins. There's a ton of them and they're all orphans, almost all created by User:ProteinBoxBot, all the way back from 2009 and earlier. I don't know the first thing about molecular biology, so I have no way to determine if these articles are correct, useful, or notable...really anything about them. I would like to be able to de-orphan them in order to clear out the backlog there, but I don't want to make a mess of anything given my lack of knowledge.
Is there any policy or guideline covering the notability of proteins/genes as separate articles? Is there a central list article they could be linked at for the purposes of de-orphaning? Or even a group of such articles, like "genes on human chromosome 3" or something?
Any input would be a huge help, like I said I'd love to do anything that can clear out the backlog, but I don't want to break anything out of ignorance. ♠ PMC♠ (talk) 04:07, 24 January 2017 (UTC)
- Thanks for your note. Notability can normally be established by following the "Human PubMed Reference" link. Both genes have been mentioned in a number of peer-reviewed articles, hence I think they pass the notability requirement. Unfortunately there does not appear a uniform way to deophanize these articles and each one must be handled separately. EAF1 (along with EAF2, CDK9, MLLT3/AF9, AFF ( AFF1 or AFF4), the P-TEFb complex and ELL ( ELL, ELL2 or ELL3)) is a component of the super elongation complex. Creating the second article would deorphanize the former. E2F7 interacts with E2F8. Boghog ( talk) 06:47, 24 January 2017 (UTC)
- Thanks for the fast response! Okay, so, broadly speaking, if a gene/protein name returns at least a couple of results on the Human PubMed Reference then it would be considered reasonably notable. That makes sense. I don't have the technical knowledge to go creating articles like
super elongation complex, but would it be useful if I made a list of any orphaned gene articles that I found, to bring them to the attention of this WikiProject? I don't want to step on any toes or clutter anything up. ♠
PMC♠
(talk)
07:08, 24 January 2017 (UTC)
- If you post any of them here I think people would be happy to help de-orphan them.
Ajpolino (
talk)
23:40, 24 January 2017 (UTC)
- Sweet. I've dropped the first set (A-C from February 2009) onto my personal sandbox. I can always move them elsewhere if that would be better. ♠ PMC♠ (talk) 07:42, 25 January 2017 (UTC)
- If you post any of them here I think people would be happy to help de-orphan them.
Ajpolino (
talk)
23:40, 24 January 2017 (UTC)
- Thanks for the fast response! Okay, so, broadly speaking, if a gene/protein name returns at least a couple of results on the Human PubMed Reference then it would be considered reasonably notable. That makes sense. I don't have the technical knowledge to go creating articles like
super elongation complex, but would it be useful if I made a list of any orphaned gene articles that I found, to bring them to the attention of this WikiProject? I don't want to step on any toes or clutter anything up. ♠
PMC♠
(talk)
07:08, 24 January 2017 (UTC)
This section is poorly written with misleading information. This requires serious editing. Andrzej Slominski, MD, PhD Endowed Professor of Dermatology Professor of Pathology Program Director of Dermatopathology Fellowship Senior Scientist, Comprehensive Cancer Center Cancer Chemoprevention Program Member of the Editorial Boards of Exp Dermatol, J Pineal Research, PLoS ONE, Dermato-Endocrinology, Int J Mol Sci, Melanoma Management, Pol J Path Department of Dermatology University of Alabama at Birmingham Volker Hall 476C Box 109 1720 2nd Avenue South VH 476C Birmingham, AL 35294 Phone: 205.934.5245 e-mail: aslominski@uabmc.edu — Preceding unsigned comment added by 64.134.187.167 ( talk) 14:28, 3 February 2017 (UTC)
- Disclaimer: I wrote most of the RAR-related orphan receptor article. Most of the content was written ten years ago and a lot has happened since then. Precisely which part of the article is poorly written with misleading information? Your comments are appreciated but not very helpful. Per {{ sofixit}}, it would be much more constructive to provide specific suggestions on how to improve the article and even better, improve the article yourself. Is the problem with the sources supplied or the interpretation of those sources? Even less useful is your excessive description of your qualifications which does not carry much weight around here. Boghog ( talk) 22:21, 3 February 2017 (UTC)
- I have made an effort to update the article based on more recent reviews. I hope this is an improvement. Is there any information that is still misleading? Boghog ( talk) 18:42, 4 February 2017 (UTC)
Notice to participants at this page about adminship
Many participants here create a lot of content, may have to evaluate whether or not a subject is notable, decide if content complies with BLP policy, and much more. Well, these are just some of the skills considered at Wikipedia:Requests for adminship.
So, please consider taking a look at and watchlisting this page:
You could be very helpful in evaluating potential candidates, and even finding out if you would be a suitable RfA candidate.
Many thanks and best wishes,
Anna Frodesiak ( talk) 01:11, 10 February 2017 (UTC)
Nicotinamide adenine dinucleotide (NAD)
Two weeks ago I tried to make a meaningful contribution to the NAD article on Wikipedia.
It was labeled as «promotional garbage» by user:Jytdog. The last part of his edit summary was also pure and intended nonsense, and had nothing to do with the content of my original edit.
Within a couple of hours or so I was «edit warring», guilty of «sock puppetry» and given a free, one week sabbatical from Wikipedia.
My original edit, a short passage, is now on display at the NAD talk page.
Feedback from other users is appreciated. Thank you. Clowns und Kinder ( talk) 18:53, 24 February 2017 (UTC)
Wikidata and drug info
Please see Wikipedia_talk:WikiProject_Medicine#Wikidata_again. This is like playing whackamole. Jytdog ( talk) 21:36, 27 February 2017 (UTC)
Opinions requested
Opinions are requested at Talk:Expanded genetic code#mouse code about whether certain content is an acceptable use of primary sources. Thanks, Adrian J. Hunter( talk• contribs) 06:11, 13 March 2017 (UTC)
Merge proposal
Thoughts on this merge proposal (or other ideas on what to do with History of penicillin) would be much appreciated. Thanks! Ajpolino ( talk) 04:51, 14 March 2017 (UTC)
Proposal of using full size image for RNA expression pattern
Hello WP:MCB! In infoboxes of gene/protein articles, there are small thumbnail images of RNA expression pattern. I proposed switching these small thumbnail to full size image, because Wikipedia/Mediawiki image system was changed. Since image data are recalled from Wikidata, I post the proposal at Wikidata page ( wikidata:Property talk:P692#How about using full size image instead of small thumbnail?). I would like to get your thoughts on that. Thank you. -- Was a bee ( talk) 06:28, 18 March 2017 (UTC)
WP:CHEMISTRY/ WP:CHEMICALS shorcut updated
Note that per this RFC, the shortcuts to WP:CHEMISTRY/ WP:CHEMICALS have been updated.
- WP:CHEM now refer to WP:CHEMISTRY
- WP:CHEMS now refer to WP:CHEMICALS
- WP:CHM is deprecated
Old discussions have had their shortcuts updated already. If I have made a mistake during an update, feel free to revert. Headbomb { talk / contribs / physics / books} 16:04, 23 March 2017 (UTC)
Critiquing "Mitochondrial protein-transporting ATPase" Article
The article "Mitochondrial protein-transporting ATPase" there are only four citations present. Three out of four of the citations used are 20 or more years old. To make this Article on Wiki better more recent citations should be used. Also more Citations should be referenced as well. Dintamang ( talk) 19:23, 24 March 2017 (UTC)
Extension of 'Topic Page' review articles from PLOS Computational Biology to PLOS Genetics

The journal group PLOS is extending its ' Topic Page' review format that was spearheaded by PLOS Computational Biology to also include PLOS Genetics. In this format, accepted articles are dual-published both in the journal, and as Wikipedia pages (see Wikipedia category).
Suitable topics must either currently lack a Wikipedia page, or have only stub/start class contents. If you you would like to submit such a review article, see
these guidelines. If you have any recommendations for topics to be commissioned, feel free to let any of the involved editors know:
T Shafee (PLOS Gen),
D Mietchen (PLOS Comp Biol).
T.Shafee(Evo&Evo)
talk
12:27, 25 March 2017 (UTC)
Infobox for enzyme vs protein
While diagnosing how
alcohol dehydrogenase displayed compared to
inositol oxygenase, the former has a
BRENDA link autogenerated when |EC_number= is present as a feature of {{
infobox enzyme}}. The latter does have an EC number passed, but no such link, which it turns out is because {{
infobox protein}} does not have the BRENDA-link feature. But I also notice that this infobox uses a different parameter for the EC number (|ECnumber=, note lack of underscore). So three concerns:
- It's confusing that these two related templates use different parameter names for the same data.
- Is {{ infobox enzyme}} just a more specific infobox, or is there a reason one would want to use {{ infobox protein}} for an enzyme?
- And objection to cloning the BRENDA link magic into {{ infobox protein}}?
I'll post pointers from both templates' talkpages to here momentarily. DMacks ( talk) 14:56, 3 April 2017 (UTC)
- My
twothree cents:
- No objection to standardizing parameters names between related templates. I like underscores since IMHO they make the parameter name more readable.
- Strictly speaking, an EC number does not represent an enzyme, but rather a reaction catalyzed by an enzyme. More than one (iso)enzyme can catalyze the same reaction. Hence {{ infobox enzyme}} is not a more specific version of {{ infobox protein}}, they serve two distinct purposes.
- No objections to adding a BRENDA link into {{ infobox protein}}. Boghog ( talk) 15:53, 3 April 2017 (UTC)
- IMO, adding a BRENDA link to infobox protein would be useful. Seppi333 ( Insert 2¢) 17:48, 3 April 2017 (UTC)
Scope of Light-independent reactions
Hello. After the RM results at Talk:Light-independent reactions, the article " Light-independent reactions" needs attention. The scope of this article is confusing as the article suffered from naming and scope issues. I think participants here may be interested in this. -- George Ho ( talk) 13:12, 9 April 2017 (UTC)
- I left some suggestions on the talk page. This article sure does need cleanup. Adrian J. Hunter( talk• contribs) 00:58, 10 April 2017 (UTC)
Requests for comment on Microscope article
Please comment on a few requests for comments on the Microscope article.
Talk:Microscope#Request_for_comment_on_how_a_scanning_electron_microscope_works
Talk:Microscope#Request_for_comment_on_ultramicroscope
Talk:Microscope#RFC_should_article_focus_on_instrument.2C_microscope.2C_or_technique.2C_microscopy — Preceding unsigned comment added by 2600:387:6:807:0:0:0:C2 ( talk) 14:23, 27 April 2017 (UTC)
Thank you, -- 2601:648:8503:4467:F4B3:6D6C:9DCC:DC06 ( talk) 21:09, 1 April 2017 (UTC)
TfD: {{ Steroid metabolism modulators}} and {{ Cytochrome P450 modulators}}
The above navboxes are being proposed for deletion. Your comments are welcome here. Boghog ( talk) 20:46, 29 April 2017 (UTC)