The transactivation domain or trans-activating domain (TAD) is a transcription factor scaffold domain which contains binding sites for other proteins such as transcription coregulators. These binding sites are frequently referred to as activation functions (AFs). [1] TADs are named after their amino acid composition. These amino acids are either essential for the activity or simply the most abundant in the TAD. Transactivation by the Gal4 transcription factor is mediated by acidic amino acids, whereas hydrophobic residues in Gcn4 play a similar role. Hence, the TADs in Gal4 and Gcn4 are referred to as acidic or hydrophobic, respectively. [2] [3] [4] [5] [6] [7] [8] [9]
In general we can distinguish four classes of TADs: [10]
- acidic domains (called also “acid blobs” or “negative noodles", rich in D and E amino acids, present in Gal4, Gcn4 and VP16). [11]
- glutamine-rich domains (contains multiple repetitions like "QQQXXXQQQ", present in SP1) [12]
- proline-rich domains (contains repetitions like "PPPXXXPPP" present in c-jun, AP2 and Oct-2) [13]
- isoleucine-rich domains (repetitions "IIXXII", present in NTF-1) [14]
Alternatively, since similar amino acid compositions does not necessarily mean similar activation pathways, TADs can be grouped by the process they stimulate, either initiation or elongation. [15]
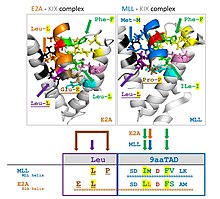
Nine-amino-acid transactivation domain (9aaTAD) defines a domain common to a large superfamily of eukaryotic transcription factors represented by Gal4, Oaf1, Leu3, Rtg3, Pho4, Gln3, Gcn4 in yeast, and by p53, NFAT, NF-κB and VP16 in mammals. The definition largely overlaps with an "acidic" family definition. A 9aaTAD prediction tool is available. [16] 9aaTADs tend to have an associated 3-aa hydrophobic (usually Leu-rich) region immediately to its N-terminal. [17]
9aaTAD transcription factors p53, VP16, MLL, E2A, HSF1, NF-IL6, NFAT1 and NF-κB interact directly with the general coactivators TAF9 and CBP/p300. [16] [18] [19] [20] [21] [22] [23] [24] [25] [26] [27] [28] [29] p53 9aaTADs interact with TAF9, GCN5 and with multiple domains of CBP/p300 (KIX, TAZ1,TAZ2 and IBiD). [30] [31] [32] [33] [34]
The KIX domain of general coactivators Med15(Gal11) interacts with 9aaTAD transcription factors Gal4, Pdr1, Oaf1, Gcn4, VP16, Pho4, Msn2, Ino2 and P201. Positions 1, 3-4, and 7 of the 9aaTAD are the main residues that interact with KIX. [35] [36] [37] [38] [39] [40] [41] [42] [43] [44] [45] [46] [47] [48] [49] [50] Interactions of Gal4, Pdr1 and Gcn4 with Taf9 have been observed. [8] [51] [52] 9aaTAD is a common transactivation domain which recruits multiple general coactivators TAF9, MED15, CBP/p300 and GCN5. [16]
| Source | 9aaTAD | Peptide-KIX interaction (NMR) |
|---|---|---|
| p53 TAD1 | E TFSD LWKL | LSPEETFSDLWKLPE |
| p53 TAD2 | D DIEQ WFTE | QAMDDLMLSPDDIEQWFTEDPGPD |
| MLL | S DIMD FVLK | DCGNILPSDIMDFVLKNTP |
| E2A | D LLDF SMMF | PVGTDKELSDLLDFSMMFPLPVT |
| Rtg3 | E TLDF SLVT | E2A homolog |
| CREB | R KILN DLSS | RREILSRRPSYRKILNDLSSDAP |
| CREBaB6 | E AILA ELKK | CREB-mutant binding to KIX |
| Gli3 | D DVVQ YLNS | TAD homology to CREB/KIX |
| Gal4 | D DVYN YLFD | Pdr1 and Oaf1 homolog |
| Oaf1 | D LFDY DFLV | DLFDYDFLV |
| Pip2 | D FFDY DLLF | Oafl homolog |
| Pdr1 | E DLYS ILWS | EDLYSILWSDWY |
| Pdr3 | T DLYH TLWN | Pdr1 homolog |
Glutamine (Q)-rich TADs are found in POU2F1 (Oct1), POU2F2 (Oct2), and Sp1 (see also Sp/KLF family). [12] Although such is not the case for every Q-rich TAD, Sp1 is shown to interact with TAF4 (TAFII 130), a part of the TFIID assembly. [15] [53]
- ^ Wärnmark A, Treuter E, Wright AP, Gustafsson JA (Oct 2003). "Activation functions 1 and 2 of nuclear receptors: molecular strategies for transcriptional activation". Molecular Endocrinology. 17 (10): 1901–9. doi: 10.1210/me.2002-0384. PMID 12893880.
- ^ Ma J, Ptashne M (Oct 1987). "A new class of yeast transcriptional activators". Cell. 51 (1): 113–9. doi: 10.1016/0092-8674(87)90015-8. PMID 3115591.
- ^ Sadowski I, Ma J, Triezenberg S, Ptashne M (Oct 1988). "GAL4-VP16 is an unusually potent transcriptional activator". Nature. 335 (6190): 563–4. Bibcode: 1988Natur.335..563S. doi: 10.1038/335563a0. PMID 3047590. S2CID 4276393.
- ^ Sullivan SM, Horn PJ, Olson VA, Koop AH, Niu W, Ebright RH, Triezenberg SJ (Oct 1998). "Mutational analysis of a transcriptional activation region of the VP16 protein of herpes simplex virus". Nucleic Acids Research. 26 (19): 4487–96. doi: 10.1093/nar/26.19.4487. PMC 147869. PMID 9742254.
- ^ Gill G, Ptashne M (Oct 1987). "Mutants of GAL4 protein altered in an activation function". Cell. 51 (1): 121–6. doi: 10.1016/0092-8674(87)90016-X. PMID 3115592.
- ^ Hope IA, Mahadevan S, Struhl K (Jun 1988). "Structural and functional characterization of the short acidic transcriptional activation region of yeast GCN4 protein". Nature. 333 (6174): 635–40. Bibcode: 1988Natur.333..635H. doi: 10.1038/333635a0. PMID 3287180. S2CID 2635634.
- ^ Hope IA, Struhl K (Sep 1986). "Functional dissection of a eukaryotic transcriptional activator protein, GCN4 of yeast". Cell. 46 (6): 885–94. doi: 10.1016/0092-8674(86)90070-X. PMID 3530496. S2CID 40730692.
- ^ a b Drysdale CM, Dueñas E, Jackson BM, Reusser U, Braus GH, Hinnebusch AG (Mar 1995). "The transcriptional activator GCN4 contains multiple activation domains that are critically dependent on hydrophobic amino acids". Molecular and Cellular Biology. 15 (3): 1220–33. doi: 10.1128/mcb.15.3.1220. PMC 230345. PMID 7862116.
- ^ Regier JL, Shen F, Triezenberg SJ (Feb 1993). "Pattern of aromatic and hydrophobic amino acids critical for one of two subdomains of the VP16 transcriptional activator". Proceedings of the National Academy of Sciences of the United States of America. 90 (3): 883–7. Bibcode: 1993PNAS...90..883R. doi: 10.1073/pnas.90.3.883. PMC 45774. PMID 8381535.
- ^ Mitchell PJ, Tjian R (July 1989). "Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins". Science. 245 (4916): 371–8. Bibcode: 1989Sci...245..371M. doi: 10.1126/science.2667136. PMID 2667136.
- ^ Sadowski I, Ma J, Triezenberg S, Ptashne M (October 1988). "GAL4-VP16 is an unusually potent transcriptional activator". Nature. 335 (6190): 563–4. Bibcode: 1988Natur.335..563S. doi: 10.1038/335563a0. PMID 3047590. S2CID 4276393.
- ^ a b Courey AJ, Holtzman DA, Jackson SP, Tjian R (December 1989). "Synergistic activation by the glutamine-rich domains of human transcription factor Sp1". Cell. 59 (5): 827–36. doi: 10.1016/0092-8674(89)90606-5. PMID 2512012. S2CID 2910480.
- ^ Mermod N, O'Neill EA, Kelly TJ, Tjian R (August 1989). "The proline-rich transcriptional activator of CTF/NF-I is distinct from the replication and DNA binding domain". Cell. 58 (4): 741–53. doi: 10.1016/0092-8674(89)90108-6. PMID 2504497. S2CID 22817940.
- ^ Attardi LD, Tjian R (July 1993). "Drosophila tissue-specific transcription factor NTF-1 contains a novel isoleucine-rich activation motif". Genes & Development. 7 (7B): 1341–53. doi: 10.1101/gad.7.7b.1341. PMID 8330738.
- ^ a b Frietze S, Farnham PJ (14 April 2011). "Transcription Factor Effector Domains". A Handbook of Transcription Factors. Subcellular Biochemistry. Vol. 52. pp. 261–277. doi: 10.1007/978-90-481-9069-0_12. ISBN 978-90-481-9068-3. PMC 4151296. PMID 21557087.
- ^ a b c Piskacek S, Gregor M, Nemethova M, Grabner M, Kovarik P, Piskacek M (Jun 2007). "Nine-amino-acid transactivation domain: establishment and prediction utilities". Genomics. 89 (6): 756–68. doi: 10.1016/j.ygeno.2007.02.003. PMID 17467953.
- ^ a b Piskacek M, Havelka M, Rezacova M, Knight A (12 September 2016). "The 9aaTAD Transactivation Domains: From Gal4 to p53". PLOS ONE. 11 (9): e0162842. Bibcode: 2016PLoSO..1162842P. doi: 10.1371/journal.pone.0162842. PMC 5019370. PMID 27618436.
- ^ Uesugi M, Verdine GL (Dec 1999). "The alpha-helical FXXPhiPhi motif in p53: TAF interaction and discrimination by MDM2". Proceedings of the National Academy of Sciences of the United States of America. 96 (26): 14801–6. Bibcode: 1999PNAS...9614801U. doi: 10.1073/pnas.96.26.14801. PMC 24728. PMID 10611293.
- ^ Uesugi M, Nyanguile O, Lu H, Levine AJ, Verdine GL (Aug 1997). "Induced alpha helix in the VP16 activation domain upon binding to a human TAF". Science. 277 (5330): 1310–3. doi: 10.1126/science.277.5330.1310. PMID 9271577.
- ^ Choi Y, Asada S, Uesugi M (May 2000). "Divergent hTAFII31-binding motifs hidden in activation domains". The Journal of Biological Chemistry. 275 (21): 15912–6. doi: 10.1074/jbc.275.21.15912. PMID 10821850.
- ^ Lee CW, Arai M, Martinez-Yamout MA, Dyson HJ, Wright PE (Mar 2009). "Mapping the interactions of the p53 transactivation domain with the KIX domain of CBP". Biochemistry. 48 (10): 2115–24. doi: 10.1021/bi802055v. PMC 2765525. PMID 19220000.
- ^ Goto NK, Zor T, Martinez-Yamout M, Dyson HJ, Wright PE (Nov 2002). "Cooperativity in transcription factor binding to the coactivator CREB-binding protein (CBP). The mixed lineage leukemia protein (MLL) activation domain binds to an allosteric site on the KIX domain". The Journal of Biological Chemistry. 277 (45): 43168–74. CiteSeerX 10.1.1.615.9401. doi: 10.1074/jbc.M207660200. PMID 12205094.
- ^ Radhakrishnan I, Pérez-Alvarado GC, Parker D, Dyson HJ, Montminy MR, Wright PE (Dec 1997). "Solution structure of the KIX domain of CBP bound to the transactivation domain of CREB: a model for activator:coactivator interactions". Cell. 91 (6): 741–52. doi: 10.1016/S0092-8674(00)80463-8. PMID 9413984. S2CID 17268267.
- ^ Zor T, Mayr BM, Dyson HJ, Montminy MR, Wright PE (Nov 2002). "Roles of phosphorylation and helix propensity in the binding of the KIX domain of CREB-binding protein by constitutive (c-Myb) and inducible (CREB) activators". The Journal of Biological Chemistry. 277 (44): 42241–8. doi: 10.1074/jbc.M207361200. PMID 12196545.
- ^ Brüschweiler S, Schanda P, Kloiber K, Brutscher B, Kontaxis G, Konrat R, Tollinger M (Mar 2009). "Direct observation of the dynamic process underlying allosteric signal transmission". Journal of the American Chemical Society. 131 (8): 3063–8. doi: 10.1021/ja809947w. PMID 19203263.
- ^ Liu GH, Qu J, Shen X (May 2008). "NF-kappaB/p65 antagonizes Nrf2-ARE pathway by depriving CBP from Nrf2 and facilitating recruitment of HDAC3 to MafK". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1783 (5): 713–27. doi: 10.1016/j.bbamcr.2008.01.002. PMID 18241676.
- ^ Bayly R, Murase T, Hyndman BD, Savage R, Nurmohamed S, Munro K, Casselman R, Smith SP, LeBrun DP (Sep 2006). "Critical role for a single leucine residue in leukemia induction by E2A-PBX1". Molecular and Cellular Biology. 26 (17): 6442–52. doi: 10.1128/MCB.02025-05. PMC 1592826. PMID 16914730.
- ^ García-Rodríguez C, Rao A (Jun 1998). "Nuclear factor of activated T cells (NFAT)-dependent transactivation regulated by the coactivators p300/CREB-binding protein (CBP)". The Journal of Experimental Medicine. 187 (12): 2031–6. doi: 10.1084/jem.187.12.2031. PMC 2212364. PMID 9625762.
- ^ Mink S, Haenig B, Klempnauer KH (Nov 1997). "Interaction and functional collaboration of p300 and C/EBPbeta". Molecular and Cellular Biology. 17 (11): 6609–17. doi: 10.1128/mcb.17.11.6609. PMC 232514. PMID 9343424.
- ^ Teufel DP, Freund SM, Bycroft M, Fersht AR (Apr 2007). "Four domains of p300 each bind tightly to a sequence spanning both transactivation subdomains of p53". Proceedings of the National Academy of Sciences of the United States of America. 104 (17): 7009–14. Bibcode: 2007PNAS..104.7009T. doi: 10.1073/pnas.0702010104. PMC 1855428. PMID 17438265.
- ^ Teufel DP, Bycroft M, Fersht AR (May 2009). "Regulation by phosphorylation of the relative affinities of the N-terminal transactivation domains of p53 for p300 domains and Mdm2". Oncogene. 28 (20): 2112–8. doi: 10.1038/onc.2009.71. PMC 2685776. PMID 19363523.
- ^ Feng H, Jenkins LM, Durell SR, Hayashi R, Mazur SJ, Cherry S, Tropea JE, Miller M, Wlodawer A, Appella E, Bai Y (Feb 2009). "Structural basis for p300 Taz2-p53 TAD1 binding and modulation by phosphorylation". Structure. 17 (2): 202–10. doi: 10.1016/j.str.2008.12.009. PMC 2705179. PMID 19217391.
- ^ Ferreon JC, Lee CW, Arai M, Martinez-Yamout MA, Dyson HJ, Wright PE (Apr 2009). "Cooperative regulation of p53 by modulation of ternary complex formation with CBP/p300 and HDM2". Proceedings of the National Academy of Sciences of the United States of America. 106 (16): 6591–6. Bibcode: 2009PNAS..106.6591F. doi: 10.1073/pnas.0811023106. PMC 2672497. PMID 19357310.
- ^ Gamper AM, Roeder RG (Apr 2008). "Multivalent binding of p53 to the STAGA complex mediates coactivator recruitment after UV damage". Molecular and Cellular Biology. 28 (8): 2517–27. doi: 10.1128/MCB.01461-07. PMC 2293101. PMID 18250150.
- ^ Fukasawa T, Fukuma M, Yano K, Sakurai H (Feb 2001). "A genome-wide analysis of transcriptional effect of Gal11 in Saccharomyces cerevisiae: an application of "mini-array hybridization technique"". DNA Research. 8 (1): 23–31. doi: 10.1093/dnares/8.1.23. PMID 11258797.
- ^ Badi L, Barberis A (Aug 2001). "Proteins that genetically interact with the Saccharomyces cerevisiae transcription factor Gal11p emphasize its role in the initiation-elongation transition". Molecular Genetics and Genomics. 265 (6): 1076–86. doi: 10.1007/s004380100505. PMID 11523780. S2CID 19287634.
- ^ Kim YJ, Björklund S, Li Y, Sayre MH, Kornberg RD (May 1994). "A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II". Cell. 77 (4): 599–608. doi: 10.1016/0092-8674(94)90221-6. PMID 8187178. S2CID 5002125.
- ^ Suzuki Y, Nogi Y, Abe A, Fukasawa T (Nov 1988). "GAL11 protein, an auxiliary transcription activator for genes encoding galactose-metabolizing enzymes in Saccharomyces cerevisiae". Molecular and Cellular Biology. 8 (11): 4991–9. doi: 10.1128/mcb.8.11.4991. PMC 365593. PMID 3062377.
- ^ Fassler JS, Winston F (Dec 1989). "The Saccharomyces cerevisiae SPT13/GAL11 gene has both positive and negative regulatory roles in transcription". Molecular and Cellular Biology. 9 (12): 5602–9. doi: 10.1128/mcb.9.12.5602. PMC 363730. PMID 2685570.
- ^ Park JM, Kim HS, Han SJ, Hwang MS, Lee YC, Kim YJ (Dec 2000). "In vivo requirement of activator-specific binding targets of mediator". Molecular and Cellular Biology. 20 (23): 8709–19. doi: 10.1128/mcb.20.23.8709-8719.2000. PMC 86488. PMID 11073972.
- ^ Lu Z, Ansari AZ, Lu X, Ogirala A, Ptashne M (Jun 2002). "A target essential for the activity of a nonacidic yeast transcriptional activator". Proceedings of the National Academy of Sciences of the United States of America. 99 (13): 8591–6. Bibcode: 2002PNAS...99.8591L. doi: 10.1073/pnas.092263499. PMC 124323. PMID 12084920.
- ^ Swanson MJ, Qiu H, Sumibcay L, Krueger A, Kim SJ, Natarajan K, Yoon S, Hinnebusch AG (Apr 2003). "A multiplicity of coactivators is required by Gcn4p at individual promoters in vivo". Molecular and Cellular Biology. 23 (8): 2800–20. doi: 10.1128/MCB.23.8.2800-2820.2003. PMC 152555. PMID 12665580.
- ^ Bryant GO, Ptashne M (May 2003). "Independent recruitment in vivo by Gal4 of two complexes required for transcription". Molecular Cell. 11 (5): 1301–9. doi: 10.1016/S1097-2765(03)00144-8. PMID 12769853.
- ^ Fishburn J, Mohibullah N, Hahn S (Apr 2005). "Function of a eukaryotic transcription activator during the transcription cycle". Molecular Cell. 18 (3): 369–78. doi: 10.1016/j.molcel.2005.03.029. PMID 15866178.
- ^ Lim MK, Tang V, Le Saux A, Schüller J, Bongards C, Lehming N (Nov 2007). "Gal11p dosage-compensates transcriptional activator deletions via Taf14p". Journal of Molecular Biology. 374 (1): 9–23. doi: 10.1016/j.jmb.2007.09.013. PMID 17919657.
- ^ Lallet S, Garreau H, Garmendia-Torres C, Szestakowska D, Boy-Marcotte E, Quevillon-Chéruel S, Jacquet M (Oct 2006). "Role of Gal11, a component of the RNA polymerase II mediator in stress-induced hyperphosphorylation of Msn2 in Saccharomyces cerevisiae". Molecular Microbiology. 62 (2): 438–52. doi: 10.1111/j.1365-2958.2006.05363.x. PMID 17020582.
- ^ Dietz M, Heyken WT, Hoppen J, Geburtig S, Schüller HJ (May 2003). "TFIIB and subunits of the SAGA complex are involved in transcriptional activation of phospholipid biosynthetic genes by the regulatory protein Ino2 in the yeast Saccharomyces cerevisiae". Molecular Microbiology. 48 (4): 1119–30. doi: 10.1046/j.1365-2958.2003.03501.x. PMID 12753200.
- ^ Mizuno T, Harashima S (Apr 2003). "Gal11 is a general activator of basal transcription, whose activity is regulated by the general repressor Sin4 in yeast". Molecular Genetics and Genomics. 269 (1): 68–77. doi: 10.1007/s00438-003-0810-x. PMID 12715155. S2CID 882139.
- ^ Thakur JK, Arthanari H, Yang F, Pan SJ, Fan X, Breger J, Frueh DP, Gulshan K, Li DK, Mylonakis E, Struhl K, Moye-Rowley WS, Cormack BP, Wagner G, Näär AM (Apr 2008). "A nuclear receptor-like pathway regulating multidrug resistance in fungi". Nature. 452 (7187): 604–9. Bibcode: 2008Natur.452..604T. doi: 10.1038/nature06836. PMID 18385733. S2CID 205212715.
- ^ Thakur JK, Arthanari H, Yang F, Chau KH, Wagner G, Näär AM (Feb 2009). "Mediator subunit Gal11p/MED15 is required for fatty acid-dependent gene activation by yeast transcription factor Oaf1p". The Journal of Biological Chemistry. 284 (7): 4422–8. doi: 10.1074/jbc.M808263200. PMC 3837390. PMID 19056732.
- ^ Klein J, Nolden M, Sanders SL, Kirchner J, Weil PA, Melcher K (Feb 2003). "Use of a genetically introduced cross-linker to identify interaction sites of acidic activators within native transcription factor IID and SAGA". The Journal of Biological Chemistry. 278 (9): 6779–86. doi: 10.1074/jbc.M212514200. PMID 12501245.
- ^ Milgrom E, West RW, Gao C, Shen WC (Nov 2005). "TFIID and Spt-Ada-Gcn5-acetyltransferase functions probed by genome-wide synthetic genetic array analysis using a Saccharomyces cerevisiae taf9-ts allele". Genetics. 171 (3): 959–73. doi: 10.1534/genetics.105.046557. PMC 1456853. PMID 16118188.
- ^ Hibino E, Inoue R, Sugiyama M, Kuwahara J, Matsuzaki K, Hoshino M (November 2016). "Interaction between intrinsically disordered regions in transcription factors Sp1 and TAF4". Protein Science. 25 (11): 2006–2017. doi: 10.1002/pro.3013. PMC 5079245. PMID 27515574.
The transactivation domain or trans-activating domain (TAD) is a transcription factor scaffold domain which contains binding sites for other proteins such as transcription coregulators. These binding sites are frequently referred to as activation functions (AFs). [1] TADs are named after their amino acid composition. These amino acids are either essential for the activity or simply the most abundant in the TAD. Transactivation by the Gal4 transcription factor is mediated by acidic amino acids, whereas hydrophobic residues in Gcn4 play a similar role. Hence, the TADs in Gal4 and Gcn4 are referred to as acidic or hydrophobic, respectively. [2] [3] [4] [5] [6] [7] [8] [9]
In general we can distinguish four classes of TADs: [10]
- acidic domains (called also “acid blobs” or “negative noodles", rich in D and E amino acids, present in Gal4, Gcn4 and VP16). [11]
- glutamine-rich domains (contains multiple repetitions like "QQQXXXQQQ", present in SP1) [12]
- proline-rich domains (contains repetitions like "PPPXXXPPP" present in c-jun, AP2 and Oct-2) [13]
- isoleucine-rich domains (repetitions "IIXXII", present in NTF-1) [14]
Alternatively, since similar amino acid compositions does not necessarily mean similar activation pathways, TADs can be grouped by the process they stimulate, either initiation or elongation. [15]

Nine-amino-acid transactivation domain (9aaTAD) defines a domain common to a large superfamily of eukaryotic transcription factors represented by Gal4, Oaf1, Leu3, Rtg3, Pho4, Gln3, Gcn4 in yeast, and by p53, NFAT, NF-κB and VP16 in mammals. The definition largely overlaps with an "acidic" family definition. A 9aaTAD prediction tool is available. [16] 9aaTADs tend to have an associated 3-aa hydrophobic (usually Leu-rich) region immediately to its N-terminal. [17]
9aaTAD transcription factors p53, VP16, MLL, E2A, HSF1, NF-IL6, NFAT1 and NF-κB interact directly with the general coactivators TAF9 and CBP/p300. [16] [18] [19] [20] [21] [22] [23] [24] [25] [26] [27] [28] [29] p53 9aaTADs interact with TAF9, GCN5 and with multiple domains of CBP/p300 (KIX, TAZ1,TAZ2 and IBiD). [30] [31] [32] [33] [34]
The KIX domain of general coactivators Med15(Gal11) interacts with 9aaTAD transcription factors Gal4, Pdr1, Oaf1, Gcn4, VP16, Pho4, Msn2, Ino2 and P201. Positions 1, 3-4, and 7 of the 9aaTAD are the main residues that interact with KIX. [35] [36] [37] [38] [39] [40] [41] [42] [43] [44] [45] [46] [47] [48] [49] [50] Interactions of Gal4, Pdr1 and Gcn4 with Taf9 have been observed. [8] [51] [52] 9aaTAD is a common transactivation domain which recruits multiple general coactivators TAF9, MED15, CBP/p300 and GCN5. [16]
| Source | 9aaTAD | Peptide-KIX interaction (NMR) |
|---|---|---|
| p53 TAD1 | E TFSD LWKL | LSPEETFSDLWKLPE |
| p53 TAD2 | D DIEQ WFTE | QAMDDLMLSPDDIEQWFTEDPGPD |
| MLL | S DIMD FVLK | DCGNILPSDIMDFVLKNTP |
| E2A | D LLDF SMMF | PVGTDKELSDLLDFSMMFPLPVT |
| Rtg3 | E TLDF SLVT | E2A homolog |
| CREB | R KILN DLSS | RREILSRRPSYRKILNDLSSDAP |
| CREBaB6 | E AILA ELKK | CREB-mutant binding to KIX |
| Gli3 | D DVVQ YLNS | TAD homology to CREB/KIX |
| Gal4 | D DVYN YLFD | Pdr1 and Oaf1 homolog |
| Oaf1 | D LFDY DFLV | DLFDYDFLV |
| Pip2 | D FFDY DLLF | Oafl homolog |
| Pdr1 | E DLYS ILWS | EDLYSILWSDWY |
| Pdr3 | T DLYH TLWN | Pdr1 homolog |
Glutamine (Q)-rich TADs are found in POU2F1 (Oct1), POU2F2 (Oct2), and Sp1 (see also Sp/KLF family). [12] Although such is not the case for every Q-rich TAD, Sp1 is shown to interact with TAF4 (TAFII 130), a part of the TFIID assembly. [15] [53]
- ^ Wärnmark A, Treuter E, Wright AP, Gustafsson JA (Oct 2003). "Activation functions 1 and 2 of nuclear receptors: molecular strategies for transcriptional activation". Molecular Endocrinology. 17 (10): 1901–9. doi: 10.1210/me.2002-0384. PMID 12893880.
- ^ Ma J, Ptashne M (Oct 1987). "A new class of yeast transcriptional activators". Cell. 51 (1): 113–9. doi: 10.1016/0092-8674(87)90015-8. PMID 3115591.
- ^ Sadowski I, Ma J, Triezenberg S, Ptashne M (Oct 1988). "GAL4-VP16 is an unusually potent transcriptional activator". Nature. 335 (6190): 563–4. Bibcode: 1988Natur.335..563S. doi: 10.1038/335563a0. PMID 3047590. S2CID 4276393.
- ^ Sullivan SM, Horn PJ, Olson VA, Koop AH, Niu W, Ebright RH, Triezenberg SJ (Oct 1998). "Mutational analysis of a transcriptional activation region of the VP16 protein of herpes simplex virus". Nucleic Acids Research. 26 (19): 4487–96. doi: 10.1093/nar/26.19.4487. PMC 147869. PMID 9742254.
- ^ Gill G, Ptashne M (Oct 1987). "Mutants of GAL4 protein altered in an activation function". Cell. 51 (1): 121–6. doi: 10.1016/0092-8674(87)90016-X. PMID 3115592.
- ^ Hope IA, Mahadevan S, Struhl K (Jun 1988). "Structural and functional characterization of the short acidic transcriptional activation region of yeast GCN4 protein". Nature. 333 (6174): 635–40. Bibcode: 1988Natur.333..635H. doi: 10.1038/333635a0. PMID 3287180. S2CID 2635634.
- ^ Hope IA, Struhl K (Sep 1986). "Functional dissection of a eukaryotic transcriptional activator protein, GCN4 of yeast". Cell. 46 (6): 885–94. doi: 10.1016/0092-8674(86)90070-X. PMID 3530496. S2CID 40730692.
- ^ a b Drysdale CM, Dueñas E, Jackson BM, Reusser U, Braus GH, Hinnebusch AG (Mar 1995). "The transcriptional activator GCN4 contains multiple activation domains that are critically dependent on hydrophobic amino acids". Molecular and Cellular Biology. 15 (3): 1220–33. doi: 10.1128/mcb.15.3.1220. PMC 230345. PMID 7862116.
- ^ Regier JL, Shen F, Triezenberg SJ (Feb 1993). "Pattern of aromatic and hydrophobic amino acids critical for one of two subdomains of the VP16 transcriptional activator". Proceedings of the National Academy of Sciences of the United States of America. 90 (3): 883–7. Bibcode: 1993PNAS...90..883R. doi: 10.1073/pnas.90.3.883. PMC 45774. PMID 8381535.
- ^ Mitchell PJ, Tjian R (July 1989). "Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins". Science. 245 (4916): 371–8. Bibcode: 1989Sci...245..371M. doi: 10.1126/science.2667136. PMID 2667136.
- ^ Sadowski I, Ma J, Triezenberg S, Ptashne M (October 1988). "GAL4-VP16 is an unusually potent transcriptional activator". Nature. 335 (6190): 563–4. Bibcode: 1988Natur.335..563S. doi: 10.1038/335563a0. PMID 3047590. S2CID 4276393.
- ^ a b Courey AJ, Holtzman DA, Jackson SP, Tjian R (December 1989). "Synergistic activation by the glutamine-rich domains of human transcription factor Sp1". Cell. 59 (5): 827–36. doi: 10.1016/0092-8674(89)90606-5. PMID 2512012. S2CID 2910480.
- ^ Mermod N, O'Neill EA, Kelly TJ, Tjian R (August 1989). "The proline-rich transcriptional activator of CTF/NF-I is distinct from the replication and DNA binding domain". Cell. 58 (4): 741–53. doi: 10.1016/0092-8674(89)90108-6. PMID 2504497. S2CID 22817940.
- ^ Attardi LD, Tjian R (July 1993). "Drosophila tissue-specific transcription factor NTF-1 contains a novel isoleucine-rich activation motif". Genes & Development. 7 (7B): 1341–53. doi: 10.1101/gad.7.7b.1341. PMID 8330738.
- ^ a b Frietze S, Farnham PJ (14 April 2011). "Transcription Factor Effector Domains". A Handbook of Transcription Factors. Subcellular Biochemistry. Vol. 52. pp. 261–277. doi: 10.1007/978-90-481-9069-0_12. ISBN 978-90-481-9068-3. PMC 4151296. PMID 21557087.
- ^ a b c Piskacek S, Gregor M, Nemethova M, Grabner M, Kovarik P, Piskacek M (Jun 2007). "Nine-amino-acid transactivation domain: establishment and prediction utilities". Genomics. 89 (6): 756–68. doi: 10.1016/j.ygeno.2007.02.003. PMID 17467953.
- ^ a b Piskacek M, Havelka M, Rezacova M, Knight A (12 September 2016). "The 9aaTAD Transactivation Domains: From Gal4 to p53". PLOS ONE. 11 (9): e0162842. Bibcode: 2016PLoSO..1162842P. doi: 10.1371/journal.pone.0162842. PMC 5019370. PMID 27618436.
- ^ Uesugi M, Verdine GL (Dec 1999). "The alpha-helical FXXPhiPhi motif in p53: TAF interaction and discrimination by MDM2". Proceedings of the National Academy of Sciences of the United States of America. 96 (26): 14801–6. Bibcode: 1999PNAS...9614801U. doi: 10.1073/pnas.96.26.14801. PMC 24728. PMID 10611293.
- ^ Uesugi M, Nyanguile O, Lu H, Levine AJ, Verdine GL (Aug 1997). "Induced alpha helix in the VP16 activation domain upon binding to a human TAF". Science. 277 (5330): 1310–3. doi: 10.1126/science.277.5330.1310. PMID 9271577.
- ^ Choi Y, Asada S, Uesugi M (May 2000). "Divergent hTAFII31-binding motifs hidden in activation domains". The Journal of Biological Chemistry. 275 (21): 15912–6. doi: 10.1074/jbc.275.21.15912. PMID 10821850.
- ^ Lee CW, Arai M, Martinez-Yamout MA, Dyson HJ, Wright PE (Mar 2009). "Mapping the interactions of the p53 transactivation domain with the KIX domain of CBP". Biochemistry. 48 (10): 2115–24. doi: 10.1021/bi802055v. PMC 2765525. PMID 19220000.
- ^ Goto NK, Zor T, Martinez-Yamout M, Dyson HJ, Wright PE (Nov 2002). "Cooperativity in transcription factor binding to the coactivator CREB-binding protein (CBP). The mixed lineage leukemia protein (MLL) activation domain binds to an allosteric site on the KIX domain". The Journal of Biological Chemistry. 277 (45): 43168–74. CiteSeerX 10.1.1.615.9401. doi: 10.1074/jbc.M207660200. PMID 12205094.
- ^ Radhakrishnan I, Pérez-Alvarado GC, Parker D, Dyson HJ, Montminy MR, Wright PE (Dec 1997). "Solution structure of the KIX domain of CBP bound to the transactivation domain of CREB: a model for activator:coactivator interactions". Cell. 91 (6): 741–52. doi: 10.1016/S0092-8674(00)80463-8. PMID 9413984. S2CID 17268267.
- ^ Zor T, Mayr BM, Dyson HJ, Montminy MR, Wright PE (Nov 2002). "Roles of phosphorylation and helix propensity in the binding of the KIX domain of CREB-binding protein by constitutive (c-Myb) and inducible (CREB) activators". The Journal of Biological Chemistry. 277 (44): 42241–8. doi: 10.1074/jbc.M207361200. PMID 12196545.
- ^ Brüschweiler S, Schanda P, Kloiber K, Brutscher B, Kontaxis G, Konrat R, Tollinger M (Mar 2009). "Direct observation of the dynamic process underlying allosteric signal transmission". Journal of the American Chemical Society. 131 (8): 3063–8. doi: 10.1021/ja809947w. PMID 19203263.
- ^ Liu GH, Qu J, Shen X (May 2008). "NF-kappaB/p65 antagonizes Nrf2-ARE pathway by depriving CBP from Nrf2 and facilitating recruitment of HDAC3 to MafK". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1783 (5): 713–27. doi: 10.1016/j.bbamcr.2008.01.002. PMID 18241676.
- ^ Bayly R, Murase T, Hyndman BD, Savage R, Nurmohamed S, Munro K, Casselman R, Smith SP, LeBrun DP (Sep 2006). "Critical role for a single leucine residue in leukemia induction by E2A-PBX1". Molecular and Cellular Biology. 26 (17): 6442–52. doi: 10.1128/MCB.02025-05. PMC 1592826. PMID 16914730.
- ^ García-Rodríguez C, Rao A (Jun 1998). "Nuclear factor of activated T cells (NFAT)-dependent transactivation regulated by the coactivators p300/CREB-binding protein (CBP)". The Journal of Experimental Medicine. 187 (12): 2031–6. doi: 10.1084/jem.187.12.2031. PMC 2212364. PMID 9625762.
- ^ Mink S, Haenig B, Klempnauer KH (Nov 1997). "Interaction and functional collaboration of p300 and C/EBPbeta". Molecular and Cellular Biology. 17 (11): 6609–17. doi: 10.1128/mcb.17.11.6609. PMC 232514. PMID 9343424.
- ^ Teufel DP, Freund SM, Bycroft M, Fersht AR (Apr 2007). "Four domains of p300 each bind tightly to a sequence spanning both transactivation subdomains of p53". Proceedings of the National Academy of Sciences of the United States of America. 104 (17): 7009–14. Bibcode: 2007PNAS..104.7009T. doi: 10.1073/pnas.0702010104. PMC 1855428. PMID 17438265.
- ^ Teufel DP, Bycroft M, Fersht AR (May 2009). "Regulation by phosphorylation of the relative affinities of the N-terminal transactivation domains of p53 for p300 domains and Mdm2". Oncogene. 28 (20): 2112–8. doi: 10.1038/onc.2009.71. PMC 2685776. PMID 19363523.
- ^ Feng H, Jenkins LM, Durell SR, Hayashi R, Mazur SJ, Cherry S, Tropea JE, Miller M, Wlodawer A, Appella E, Bai Y (Feb 2009). "Structural basis for p300 Taz2-p53 TAD1 binding and modulation by phosphorylation". Structure. 17 (2): 202–10. doi: 10.1016/j.str.2008.12.009. PMC 2705179. PMID 19217391.
- ^ Ferreon JC, Lee CW, Arai M, Martinez-Yamout MA, Dyson HJ, Wright PE (Apr 2009). "Cooperative regulation of p53 by modulation of ternary complex formation with CBP/p300 and HDM2". Proceedings of the National Academy of Sciences of the United States of America. 106 (16): 6591–6. Bibcode: 2009PNAS..106.6591F. doi: 10.1073/pnas.0811023106. PMC 2672497. PMID 19357310.
- ^ Gamper AM, Roeder RG (Apr 2008). "Multivalent binding of p53 to the STAGA complex mediates coactivator recruitment after UV damage". Molecular and Cellular Biology. 28 (8): 2517–27. doi: 10.1128/MCB.01461-07. PMC 2293101. PMID 18250150.
- ^ Fukasawa T, Fukuma M, Yano K, Sakurai H (Feb 2001). "A genome-wide analysis of transcriptional effect of Gal11 in Saccharomyces cerevisiae: an application of "mini-array hybridization technique"". DNA Research. 8 (1): 23–31. doi: 10.1093/dnares/8.1.23. PMID 11258797.
- ^ Badi L, Barberis A (Aug 2001). "Proteins that genetically interact with the Saccharomyces cerevisiae transcription factor Gal11p emphasize its role in the initiation-elongation transition". Molecular Genetics and Genomics. 265 (6): 1076–86. doi: 10.1007/s004380100505. PMID 11523780. S2CID 19287634.
- ^ Kim YJ, Björklund S, Li Y, Sayre MH, Kornberg RD (May 1994). "A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II". Cell. 77 (4): 599–608. doi: 10.1016/0092-8674(94)90221-6. PMID 8187178. S2CID 5002125.
- ^ Suzuki Y, Nogi Y, Abe A, Fukasawa T (Nov 1988). "GAL11 protein, an auxiliary transcription activator for genes encoding galactose-metabolizing enzymes in Saccharomyces cerevisiae". Molecular and Cellular Biology. 8 (11): 4991–9. doi: 10.1128/mcb.8.11.4991. PMC 365593. PMID 3062377.
- ^ Fassler JS, Winston F (Dec 1989). "The Saccharomyces cerevisiae SPT13/GAL11 gene has both positive and negative regulatory roles in transcription". Molecular and Cellular Biology. 9 (12): 5602–9. doi: 10.1128/mcb.9.12.5602. PMC 363730. PMID 2685570.
- ^ Park JM, Kim HS, Han SJ, Hwang MS, Lee YC, Kim YJ (Dec 2000). "In vivo requirement of activator-specific binding targets of mediator". Molecular and Cellular Biology. 20 (23): 8709–19. doi: 10.1128/mcb.20.23.8709-8719.2000. PMC 86488. PMID 11073972.
- ^ Lu Z, Ansari AZ, Lu X, Ogirala A, Ptashne M (Jun 2002). "A target essential for the activity of a nonacidic yeast transcriptional activator". Proceedings of the National Academy of Sciences of the United States of America. 99 (13): 8591–6. Bibcode: 2002PNAS...99.8591L. doi: 10.1073/pnas.092263499. PMC 124323. PMID 12084920.
- ^ Swanson MJ, Qiu H, Sumibcay L, Krueger A, Kim SJ, Natarajan K, Yoon S, Hinnebusch AG (Apr 2003). "A multiplicity of coactivators is required by Gcn4p at individual promoters in vivo". Molecular and Cellular Biology. 23 (8): 2800–20. doi: 10.1128/MCB.23.8.2800-2820.2003. PMC 152555. PMID 12665580.
- ^ Bryant GO, Ptashne M (May 2003). "Independent recruitment in vivo by Gal4 of two complexes required for transcription". Molecular Cell. 11 (5): 1301–9. doi: 10.1016/S1097-2765(03)00144-8. PMID 12769853.
- ^ Fishburn J, Mohibullah N, Hahn S (Apr 2005). "Function of a eukaryotic transcription activator during the transcription cycle". Molecular Cell. 18 (3): 369–78. doi: 10.1016/j.molcel.2005.03.029. PMID 15866178.
- ^ Lim MK, Tang V, Le Saux A, Schüller J, Bongards C, Lehming N (Nov 2007). "Gal11p dosage-compensates transcriptional activator deletions via Taf14p". Journal of Molecular Biology. 374 (1): 9–23. doi: 10.1016/j.jmb.2007.09.013. PMID 17919657.
- ^ Lallet S, Garreau H, Garmendia-Torres C, Szestakowska D, Boy-Marcotte E, Quevillon-Chéruel S, Jacquet M (Oct 2006). "Role of Gal11, a component of the RNA polymerase II mediator in stress-induced hyperphosphorylation of Msn2 in Saccharomyces cerevisiae". Molecular Microbiology. 62 (2): 438–52. doi: 10.1111/j.1365-2958.2006.05363.x. PMID 17020582.
- ^ Dietz M, Heyken WT, Hoppen J, Geburtig S, Schüller HJ (May 2003). "TFIIB and subunits of the SAGA complex are involved in transcriptional activation of phospholipid biosynthetic genes by the regulatory protein Ino2 in the yeast Saccharomyces cerevisiae". Molecular Microbiology. 48 (4): 1119–30. doi: 10.1046/j.1365-2958.2003.03501.x. PMID 12753200.
- ^ Mizuno T, Harashima S (Apr 2003). "Gal11 is a general activator of basal transcription, whose activity is regulated by the general repressor Sin4 in yeast". Molecular Genetics and Genomics. 269 (1): 68–77. doi: 10.1007/s00438-003-0810-x. PMID 12715155. S2CID 882139.
- ^ Thakur JK, Arthanari H, Yang F, Pan SJ, Fan X, Breger J, Frueh DP, Gulshan K, Li DK, Mylonakis E, Struhl K, Moye-Rowley WS, Cormack BP, Wagner G, Näär AM (Apr 2008). "A nuclear receptor-like pathway regulating multidrug resistance in fungi". Nature. 452 (7187): 604–9. Bibcode: 2008Natur.452..604T. doi: 10.1038/nature06836. PMID 18385733. S2CID 205212715.
- ^ Thakur JK, Arthanari H, Yang F, Chau KH, Wagner G, Näär AM (Feb 2009). "Mediator subunit Gal11p/MED15 is required for fatty acid-dependent gene activation by yeast transcription factor Oaf1p". The Journal of Biological Chemistry. 284 (7): 4422–8. doi: 10.1074/jbc.M808263200. PMC 3837390. PMID 19056732.
- ^ Klein J, Nolden M, Sanders SL, Kirchner J, Weil PA, Melcher K (Feb 2003). "Use of a genetically introduced cross-linker to identify interaction sites of acidic activators within native transcription factor IID and SAGA". The Journal of Biological Chemistry. 278 (9): 6779–86. doi: 10.1074/jbc.M212514200. PMID 12501245.
- ^ Milgrom E, West RW, Gao C, Shen WC (Nov 2005). "TFIID and Spt-Ada-Gcn5-acetyltransferase functions probed by genome-wide synthetic genetic array analysis using a Saccharomyces cerevisiae taf9-ts allele". Genetics. 171 (3): 959–73. doi: 10.1534/genetics.105.046557. PMC 1456853. PMID 16118188.
- ^ Hibino E, Inoue R, Sugiyama M, Kuwahara J, Matsuzaki K, Hoshino M (November 2016). "Interaction between intrinsically disordered regions in transcription factors Sp1 and TAF4". Protein Science. 25 (11): 2006–2017. doi: 10.1002/pro.3013. PMC 5079245. PMID 27515574.