| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | Archive 2 | Archive 3 |
Deleted text?
Something is missing in this sentence: "Because they use RNA to store In 2006, French researchers came across a puzzling example of RNA-mediated inheritance in mice." Perhaps someone could fix? Jimjamjak ( talk) 15:38, 11 April 2012 (UTC)
- It was an edit in March that broke the text, and I have restored the original. Johnuniq ( talk) 02:05, 12 April 2012 (UTC)
2012 GA Review
| GA toolbox |
|---|
| Reviewing |
- This review is transcluded from Talk:Gene/GA1. The edit link for this section can be used to add comments to the review.
Reviewer: Sasata ( talk · contribs) 20:23, 19 August 2012 (UTC)
I'll review this. Sasata ( talk) 20:23, 19 August 2012 (UTC)
Before we start, could you please ensure that all paragraphs, and end-of-paragraph sentences have citations? This will help me (and other readers) verify the material. The material is pretty basic, and so a general text like Genes (Lewin) or perhaps a middle-level university genetics text would work nicely for this purpose. Sasata ( talk) 23:27, 19 August 2012 (UTC)
- There's been no editing activity on this article by the nominator, so I'm going to close this GAN now. Quite a bit a work needs to be with sourcing to meet the GA criteria. Sasata ( talk) 15:35, 4 September 2012 (UTC)
Gene exactly?
"though there still are controversies about what plays the role of the genetic material.[1]" Strikes me as a very dubious opening. DNA & RNA are all that need be mentioned (in the beginning) for genes. Prions and epigenetic factors can be left for later. I tried to find out about ref 1, without buying it, by reading articles by the editors. Plutynski is a normal biologist. Sarkar is an anti-reductionist, but his webpage fails to provide any links to his articles. How can a wiki throw doubt about DNA being a genetic material in the introduction? It is similar to starting an AIDS article with a discussion of HIV denialists. OK at the end, not at the beginning. Peggy hopper ( talk) 03:23, 13 April 2013 (UTC) "
- Yes, this is terrible in the opening. Perhaps something like "Although information is transmitted from parent to offspring in many ways, a gene is... " It has been a while since I have looked at this article, maybe I'll come back to this.
Abductive (
reasoning)
04:33, 13 April 2013 (UTC)
- I've deleted this considering there is no known controversy about DNA being the genetic material and considering there were no responses to this post giving any reason to keep it. I can see how it might be handy to have mention of other mechanisms of inheritance such as epigenetics somewhere else in this article but I'll leave that for other people if they so choose. 2403:7900:ADE1:A1DE:250:56FF:FEA6:3B1F ( talk) 09:09, 19 March 2014 (UTC)
"The genetic code is nearly the same for all known organisms." is false. Human mtDNA has a different genetic code than human nuclear DNA. This discovery was crucial in proving the symbiotic origin of eukaryotes. (ref. Lynn Margulis) (Comp Biochem Physiol B. 1993 Nov;106(3):489-94. Evolutionary changes in the genetic code. Jukes TH, Osawa S.) Peggy hopper ( talk) 04:03, 13 April 2013 (UTC)
- Here "nearly" encapsulates that fact. Readers can click on genetic code to find out more. Abductive ( reasoning) 04:33, 13 April 2013 (UTC)
discontinuing inheritance
The history section uses the expression discontinuing inheritance. I guess this means that a phenotypic trait can be observable in one generation, "disappear" i a following generation and then reappear in an even later generation. Regardless of whether this or something else is meant, it needs to be explained in a way that is more understandable for the general reader. — Preceding unsigned comment added by Ettrig ( talk • contribs)
- Should be 'discontinuous', and you're right. There's a lot of clunky prose in this article at the moment. Opabinia regalis ( talk) 09:12, 7 April 2015 (UTC)
2015 GA Review (April)
| GA toolbox |
|---|
| Reviewing |
- This review is transcluded from Talk:Gene/GA2. The edit link for this section can be used to add comments to the review.
Reviewer: Cerebellum ( talk · contribs) 01:25, 10 April 2015 (UTC)
Hello! I will be reviewing this article. --
Cerebellum (
talk)
01:25, 10 April 2015 (UTC)
- It is reasonably well written.
- a (prose, no copyvios, spelling and grammar):
 b (
MoS for
lead,
layout,
word choice,
fiction, and
lists):
b (
MoS for
lead,
layout,
word choice,
fiction, and
lists): 
- In the "Translation" section you used the word "ligates," which I do not know. Is there a more common word you could use there? Also, the first sentence of the "Gene targeting and implications" section left me confused. You say that gene targeting provides "mouse models for studying the roles of individual genes," but the sentence seems abrupt and I'm not really sure why we are talking about mice all of a sudden or what mouse models are. Maybe start off with a broad intro sentence to transition from the previous section on genes in evolution, and explain why we are now talking about mice whereas the rest of the article seemed to be about genes in all forms of life. As far as layout goes, I recommend putting the two sections on the concept of a gene ("Changing concept" and "evolutionary concept of a gene") in sequence, right now they are split up by the gene targeting paragraph.
- a (prose, no copyvios, spelling and grammar):
- It is factually accurate and
verifiable.
- a (
reference section):
 b (citations to
reliable sources):
b (citations to
reliable sources):  c (
OR):
c (
OR): 
- I think the 2012 review was too harsh on this but some of the same criticism applies. Per WP:SCG you don't need a ref for every paragraph if the information all comes from a basic textbook, but it's not clear to me where some information comes from. For example, the section on Mendelian inheritance has no references. I assume most of the information in later sections can be found in Molecular Biology of the Cell, but is that the case for Mendelian inheritance as well? Please add at least one reference to that section. Also, you always need to cite direct quotations, so please provide a reference for the Williams definition of a gene in the "Evolutionary concept of a gene" section.
- a (
reference section):
- It is broad in its coverage.
- a (
major aspects):
 b (
focused):
b (
focused): 
- a (
major aspects):
- It follows the
neutral point of view policy.
- Fair representation without bias:

- Fair representation without bias:
- It is stable.
- No edit wars, etc.:

- No edit wars, etc.:
- It is illustrated by
images and other media, where possible and appropriate.
- a (images are tagged and non-free content have
fair use rationales):
 b (
appropriate use with
suitable captions):
b (
appropriate use with
suitable captions): 
- Great use of diagrams!
- a (images are tagged and non-free content have
fair use rationales):
- Overall:
- Pass/Fail:

Good article, but I'm putting it on hold for now so you can make some tweaks. Cerebellum ( talk) 17:15, 11 April 2015 (UTC)
- Pass/Fail:
- Thanks everyone for your work on this article this week, in particular BlueMoonset for identifying the copyright issue, Opabinia regalis for correcting it, and Evolution and evolvability for fixes throughout. Unfortunately some of the issues from the review, in particular regarding sourcing, have not been fixed. Because of that I have to fail the article for now, but please let me know if you renominate it and I'll take another look. -- Cerebellum ( talk) 03:26, 19 April 2015 (UTC)
Comment
I thought I'd take a look after recent exchanges about the nomination on the GAN talk page, which ended by noting that the nomination was under review. It's an impressive article, but at the moment does not meet some of the criteria
It seems to me that the article has a classic violation of
WP:LEAD, a GA criterion: Apart from trivial basic facts, significant information should not appear in the lead if it is not covered in the remainder of the article.
The lead's third paragraph is solely about a topic, "big genes", that is only mentioned there; I could not find the phrase "big genes" anywhere in the body.
I'm completely at sea as to the sourcing, since this is a very long article where so much of the material is not sourced; I don't see how any assumption can be made about what came from the textbook (which is only cited twice) and what didn't. (Also, textbooks are big: these citations should be to a page or page range, not to an immense tome where it's impractical to find the information being referenced.) I think that every subsection should be sourced, not merely every section, and more than a source for a parenthetical comment at that (as in "Genetic code", which is the only citation in the entire section). Genes are complicated and involve technical explanations, as is plain in this article; there needs to be concomitant sourcing. As it says, Complex, current, or controversial subjects may require many citations
in the lead; this is even more true of the article body of a very complex scientific topic.
BlueMoonset (
talk)
23:46, 12 April 2015 (UTC)
- I just ran Earwig's Copyvio Detector on the article, and found significant copyvio in the article, enough to stop this nomination in its tracks until a thorough check and cleanup job has been completed. For example, a great deal of the "Gene targeting and implications" section is taken from the FN24 (www.biolsci.org) source; the second paragraph is copied almost verbatim in its entirety, as you can see in this report, and much of the first paragraph is very like FN25. BlueMoonset ( talk) 03:23, 13 April 2015 (UTC)
- Cleaned up the gene-targeting section (which was also rather undue). I had planned/promised to work on the text of this article awhile back but just haven't had the time. Images look fantastic though! Opabinia regalis ( talk) 19:41, 15 April 2015 (UTC)
Prose suggestions
Below are the main issues that I've spotted or now. Hopefully they're logically laid out. Let me know if you agree or disagree with any. T.Shafee(Evo﹠Evo) talk 00:02, 4 April 2015 (UTC)
RNA genes
Certainly an important topic but it is currently brought up in several places.
- History
- Physical definitions#RNA genes and genomes in the world (before description of protein-coding genes)
- Changing concept
This leads to it being massively over-weighted in the article when really it needs only to be a single paragraph total. RNA genes are also not referred to in the images at all. We also need to clearly distinguish between a gene encoded by RNA (e.g. genes in an RNA virus) versus a gene that encodes a functional RNA product (e.g. genes for tRNAs or siRNAs).
History and concepts
Perhaps the evolutionary concept and changing concept sections can be folded into the history section? That way the History section is structured:
- How genes were understood before we knew about DNA
- The understanding that DNA genes that encode proteins
- Minor expansion of the simple model of #2 (RNA genomes, functional RNAs, splice variants etc).
Descriptions of DNA
There seems to be some overlap between Physical definitions#Functional structure of a gene and Gene expression#Genetic code. I would suggest that the descriptions of nucleotide biochemistry could be reduced. I think it would be better to increase the focus on larger structure (promoters, ORFs, terminators, enhancers, introns etc).
Mutation
Perhaps this can be combined into either replication or evolution sections?
- Agreed with pretty much all of this. I made some grouchier and less useful notes
in my sandbox last week, but haven't had time to actually do anything about it yet.
Opabinia regalis (
talk)
03:22, 5 April 2015 (UTC)
- Good work on today's edits
Opabinia regalis. I'm still thinking of moving the evolutionary concept and functional concept sections up into History since they seem to fit with a continuing evolution of our understanding. However I don't want to make the history section balloon in size again.
T.Shafee(Evo﹠Evo)
talk
12:30, 16 April 2015 (UTC)
- Thanks! I did some overall restructuring without changing the text much, moving the bottom evolutionary gene section into history and reorganizing some of the middle. I think some of the power of those great gene structure diagrams was being lost by having such detailed information before basics like transcription.
- Still not sure what to do with the mutation section, but the article is really missing a discussion of sequencing and sequence analysis and homology, so maybe that (ie, the interpretation of variation) and mutation can go under a new top-level section on something like "sequence variation"? Opabinia regalis ( talk) 06:52, 18 April 2015 (UTC)
- Good work on today's edits
Opabinia regalis. I'm still thinking of moving the evolutionary concept and functional concept sections up into History since they seem to fit with a continuing evolution of our understanding. However I don't want to make the history section balloon in size again.
T.Shafee(Evo﹠Evo)
talk
12:30, 16 April 2015 (UTC)
In a family of four kids, one will be sick

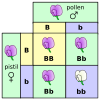
Well, no, but that seems to be what this graph implies. The alt text is equally problematic, and I've only just somewhat clarified this in the caption. I think the notion of probability needs to be more clearly communicated. Samsara 05:01, 19 April 2015 (UTC)
- @ Samsara: Not sure what you mean here. I don't think anybody's substantively edited that section of text yet, but the image seems clear enough. Is the issue that "affected" implies a disease but the description is just of two alleles, without specifying the trait? Representing the possible outcomes is pretty standard in this kind of diagram; I don't think it implies a probability error any more than a Punnett square does.
- The only criticism I have of the image/alt text is that "white" might not be the best color to use as an illustration, since "white" is in fact the common name of a trait some humans have.
Opabinia regalis (
talk)
05:48, 19 April 2015 (UTC)
- My concern is that there is no explicit statement that this deals with probabilities. A naive reader might think that if you have four kids, they turn out 1:2:1. The "alt" parameter in the article in fact reinforces this idea by explicitly talking about four children. But there are no four children in reality. There are four probabilities. Unfortunately, if you take a close look, this is a problem with most representations of this kind. Thinking outside the box, I wonder if it wouldn't make sense to have a graph that shows a number of different families, where the sum of the children ends up in a 1:2:1 ratio. Also, that would allow us to have green male with blue female as well as green female with blue male pairings, if that helps to avoid the impression that this refers to any real trait.
Samsara
06:13, 19 April 2015 (UTC)
- Hmm. The alt text, as I understand it, is intended to describe rather than interpret the image, and should not duplicate material in the caption. For that purpose the current text seems adequate. Your suggested expanded graph sounds a little too bulky for this article, but could fit well in Mendelian inheritance or genetics (which really needs a going-over to reduce redundancy with this article). Opabinia regalis ( talk) 21:17, 19 April 2015 (UTC)
- My concern is that there is no explicit statement that this deals with probabilities. A naive reader might think that if you have four kids, they turn out 1:2:1. The "alt" parameter in the article in fact reinforces this idea by explicitly talking about four children. But there are no four children in reality. There are four probabilities. Unfortunately, if you take a close look, this is a problem with most representations of this kind. Thinking outside the box, I wonder if it wouldn't make sense to have a graph that shows a number of different families, where the sum of the children ends up in a 1:2:1 ratio. Also, that would allow us to have green male with blue female as well as green female with blue male pairings, if that helps to avoid the impression that this refers to any real trait.
Samsara
06:13, 19 April 2015 (UTC)
At least the text IN the figure should use plural forms rather than singular. -- Ettrig ( talk) 12:24, 27 April 2015 (UTC)
Clarify the difference among gene, locus and allele (previously GA3)
- Note: comments below were initially added as the Good Article review in error. See here for details.
Well, allele seems to be clearly defined. What about gene and locus? In a classical sense, a locus can be a placeholder for the corresponding alleles. Is it also applicable to a gene? If so, what is the difference? If no difference, locus shall be a synonym of gene. By the way, what about a null allele, where a placeholder can be nowhere on the DNA? I am totally confused. ヒストリ案 ( talk) 13:31, 3 June 2015 (UTC)
- Good point, We only mentioned the term locus once! I'm adding a section on Locus/ Allele/ Gene clarification to the Gene#Chromosomes and Gene#Mendelian inheritance sections. I think that probably explaining null alleles is beyond the scope of this article, but I'll make sure that it comes up in both the allele and locus articles. T.Shafee(Evo﹠Evo) talk 12:08, 5 June 2015 (UTC)
- T.Shafee(Evo﹠Evo), thank you for improvement. BTW, what happen if "quantitative trait loci" taken into account? Is it a completely different concept, or share some? If the former is the case, why do we use the same term? If the latter, the idea of "a" gene is already confused... ヒストリ案 ( talk) 13:35, 8 June 2015 (UTC)
- @ ヒストリ案: The term Quantitative trait locus isn't quite the same as a gene. A QTL is a region of DNA on a chromosome known to be correlated with some phenotype change, but it can be a gene, or several genes, or part of a gene. Identifying the QTL of some phenotype can be a first step in finding a gene related to that trait. It's actually a bit of an old fashioned term which was used more extensively when we weren't able to exactly define what the DNA sequence of the gene was. It is probably more relevant to the genetics page but could certainly go in the 'see also' section for the moment. T.Shafee(Evo﹠Evo) talk 13:04, 10 June 2015 (UTC)
- ヒストリ案, I notice that this is your first-ever edit under this account, and perhaps ever on Wikipedia. Are you prepared to fully review this nomination according to the WP:WIAGA criteria, which is what is required here? If not, we should probably get a new reviewer to undertake the complete review beyond your initial questions. Thanks. BlueMoonset ( talk) 17:07, 5 June 2015 (UTC)
- BlueMoonset, I apologize for misplacement of my comment. Should I replace all the above somewhere appropriate, or delete them? Thank you for advice. ヒストリ案 ( talk) 13:27, 8 June 2015 (UTC)
- @ ヒストリ案: Your comments are still useful and definitely won't be deleted. The link that you clicked to create this section of the page are part of a process called Good article review in which an editor volunteers themself to do a thorough read-through and assessment of whether the page should be promoted to be Good article status (for example this previous review). If you were not intending to nominate yourself as the GA reviewer then we can just move your comments into the non-review part of the gene talk page. T.Shafee(Evo﹠Evo) talk 12:06, 9 June 2015 (UTC)
- Any progress so far? The current lede seems to define that a gene is a locus. Is that widely accepted, or not? -- Wordmasterexpress ( talk) 09:22, 5 October 2016 (UTC)
- A paper that clarifies the terminology (incl. gene, locus and allele among of the tens of other terms): Genetic Terminology (2012). Cheater no1 ( talk) 18:39, 9 March 2020 (UTC)
Image suggestions
Below are my opinions for changes that could be made to the article's images. T.Shafee(Evo﹠Evo) talk 00:40, 4 April 2015 (UTC)
| ===Lead image=== |
I don't think introns need to be introduced in the very first image. Certainly not in favour over other genetic elements like promoters.

| ||||||||||||||||
| DNA nucleotides |
This definitely doesn't need to be in the lead section and elements needs to relate to other images.

| ||||||||||||||||
| Gregor Mendel |
You're all good Gregor. You can stay as you are.
| ||||||||||||||||
| Punnett square |
Currently odd colour choices. Significance of the background colours not clear.
| ||||||||||||||||
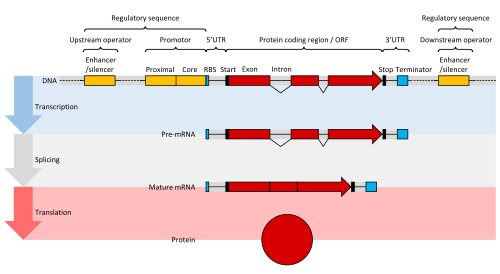
| Gene structure |
Currently no image in the article uses the traditional 'arrow' symbol of an ORF or shows all the elements of a gene together


| ||||||||||||||||
| RNA codons |
It's not immediately obvious that the letters relate to the nucleotides in the DNA image. Similarly, it's not obvious that the right hand side words are amino acids in a protein.
| ||||||||||||||||
| Function pie chart |
No fundamental problem with the image, but is there something more useful that we could show? It's no better or worse than the old gene number table that the article had 10 years ago.

| ||||||||||||||||
RNA-coding gene
no image yet |
I think it would be good to have some side-by-side comparison of a protein-coding gene vs an RNA gene (e.g tRNA).

| ||||||||||||||||
Inheritance
no image yet |
 There's currently no image actually showing genes being inherited from parent to offspring. I think this could be more useful that the Punnett square
|
Hey there, I'm a scientific illustrator and just by chance came across this article and saw you are pushing it to GA. Here's a selection of the illustrations I made for Wikipedia. As time permits I might be able to help out with improving the images of this article. Once you've decided which ones need to be improved and how, leave me a message on my talk page if you need any help (I'm not online here that often anymore nowadays). PS. The pie chart should definitely be replaced with a (horizontal) bar chart. See here why. -- SPLETTE :] How's my driving? 01:56, 4 April 2015 (UTC)
- I'm currently working on a few of the images.
- Lead image (sorry to accidentally compete User:splette, perhaps we can combine?)
- DNA structure
- RNA gene
- Hopefully should be done in next couple of days. T.Shafee(Evo﹠Evo) talk 09:06, 6 April 2015 (UTC)
Refs
- ^ Watson, JD, Baker TA, Bell SP, Gann A, Levine M, Losick R. (2004). “Ch9-10”, Molecular Biology of the Gene, 5th ed., Peason Benjamin Cummings; CSHL Press.
- ^ "Integr8 - A.thaliana Genome Statistics:".
- ^ "Understanding the Basics". The Human Genome Project. Retrieved 26 April 2015.
- ^ "WS227 Release Letter". WormBase. 10 August 2011. Retrieved 2013-11-19.
- ^ Yu, J. (5 April 2002). "A Draft Sequence of the Rice Genome (Oryza sativa L. ssp. indica)". Science. 296 (5565): 79–92. doi: 10.1126/science.1068037.
- ^ Anderson, S.; Bankier, A. T.; Barrell, B. G.; de Bruijn, M. H. L.; Coulson, A. R.; Drouin, J.; Eperon, I. C.; Nierlich, D. P.; Roe, B. A.; Sanger, F.; Schreier, P. H.; Smith, A. J. H.; Staden, R.; Young, I. G. (9 April 1981). "Sequence and organization of the human mitochondrial genome". Nature. 290 (5806): 457–465. doi: 10.1038/290457a0.
- ^ Adams, M. D. (24 March 2000). "The Genome Sequence of Drosophila melanogaster". Science. 287 (5461): 2185–2195. doi: 10.1126/science.287.5461.2185.
-
^ Pertea, Mihaela; Salzberg, Steven L (2010). "Between a chicken and a grape: estimating the number of human genes". Genome Biology. 11 (5): 206.
doi:
10.1186/gb-2010-11-5-206.
{{ cite journal}}: CS1 maint: unflagged free DOI ( link)
Revision of lede
The lede needs further work. For now, I've taken out some of the grammatical problems as well as corrected the following fallacies:
- that genes take effect on only one thing, which is later (correctly) contradicted in the article
- that genes must code for proteins (nope, ribosomal RNA and transfer RNA genes are obvious and essential counter-examples)
- that genes (or proteins) perform only cellular functions (nope, those would typically be housekeeping genes, one of many categories!)
I'm also wondering if it shouldn't be mentioned, for accuracy's sake, that RNA can also carry genes (NB: retrovirus).
Regards,
Samsara 10:12, 29 June 2015 (UTC)
- @
Samsara: Thanks for the lead section edits. You're probably right that we should mention ncRNA products early in the lead. Perhaps genes affecting multiple traits can be incorporate into the sentence "Most biological traits are under the influence of many genes as well as the environment"? RNA genomes are left to paragraph 3 of the lead currently, and I think this is reasonable since they're the exception, otherwise it makes the first sentence confusing with both RNA genomes and ncRNA products.
T.Shafee(Evo﹠Evo)
talk
12:01, 29 June 2015 (UTC)
- Yes, that is why I brought it here for discussion. Ideally imo, we would have a diagram that shows that RNA can transmit genes, but that the route via DNA is required for gene expression. Samsara 12:04, 29 June 2015 (UTC)
2015 GA Review (July)
| GA toolbox |
|---|
| Reviewing |
- This review is transcluded from Talk:Gene/GA4. The edit link for this section can be used to add comments to the review.
Reviewer: Cerebellum ( talk · contribs) 16:46, 25 July 2015 (UTC)
Sorry it took me so long to get to this, but I'm ready to do a second review now. --
Cerebellum (
talk)
16:46, 25 July 2015 (UTC)
- It is reasonably well written.
- a (prose, no copyvios, spelling and grammar):
 b (
MoS for
lead,
layout,
word choice,
fiction, and
lists):
b (
MoS for
lead,
layout,
word choice,
fiction, and
lists): 
- In the first sentence of the lead, would it be OK to replace A gene is a locus (or region) of DNA with A gene is a segment of DNA? It would be simpler, and it's what the caption of the first image says. However, if there's a big difference between the terms locus and segment then it can stay as it is. Other than that though, I made a few minor changes but the prose is generally good.
- a (prose, no copyvios, spelling and grammar):
- It is factually accurate and
verifiable.
- a (
reference section):
 b (citations to
reliable sources):
b (citations to
reliable sources):  c (
OR):
c (
OR): 
- Great job specifying the page numbers from Molecular Biology of the Cell! I added a couple of refs to the section on Mendel.
- a (
reference section):
- It is broad in its coverage.
- a (
major aspects):
 b (
focused):
b (
focused): 
- a (
major aspects):
- It follows the
neutral point of view policy.
- Fair representation without bias:

- Fair representation without bias:
- It is stable.
- No edit wars, etc.:

- No edit wars, etc.:
- It is illustrated by
images and other media, where possible and appropriate.
- a (images are tagged and non-free content have
fair use rationales):
 b (
appropriate use with
suitable captions):
b (
appropriate use with
suitable captions): 
- As before, the images are fantastic.
- a (images are tagged and non-free content have
fair use rationales):
- Overall:
- Pass/Fail:

- Thanks for all the work on this, and sorry again to keep you waiting so long for a review! Everything looks good to me though, so I'm closing this review as pass and promoting the article to GA. -- Cerebellum ( talk) 17:22, 25 July 2015 (UTC)
- Pass/Fail:
Thanks
Thank you to everyone who aided in getting this article up to GA level. It was definitely a job worth doing and hopefully sets a reasonable standard for the high-importance biology articles. T.Shafee(Evo﹠Evo) talk 00:02, 27 July 2015 (UTC)
More misc prose/additions
- in "Structure and Function" The term mRNA appears without any further explanation, for a non biologist reader (like me) this is irritating. — Preceding unsigned comment added by 141.42.200.71 ( talk) 07:01, 22 September 2015 (UTC)
- The Williams quote is widely cited elsewhere as p24 of the 1966 edition, but I don't have a copy of the book and it isn't in Google Books. Can anyone confirm?
I don't have a copy of Watson 2013 and can't preview that through Google Books either, so I'll leave going through that to those with a taste for lots of footnotes.Adding citations to the Alberts edition in NCBI is a good idea also, though in the absence of numbered chapters/sections the {{ rp}} solution to repetitive citations is less appealing. Opabinia regalis ( talk) 06:52, 18 April 2015 (UTC)Add paragraph on gene duplication, de novo genes, pseudogenes (after sequence homology?)Do we need something about genetic disorders? Currently there is the inheritance image, but that only refers to a 'trait', not a disorder, and there's nothing specifically about molecular mechanism.- Companion article genetics is highly redundant with this one.
I got distracted for a bit but have been meaning to get back to this. I think the above is the rest of my intended new content list. Any other suggestions? Opabinia regalis ( talk) 07:03, 5 May 2015 (UTC)
- Ok, the article has come a long way since we started.
I agree that we need something on genetic disorders.Possibly also a sentence or two on gene therapy in the engineering section since it's something many people will have heard about?Half of the sections have a {{ main}} template - do we really need them?
- I don't think there are any other major rearrangements of the sections left. It looks like mostly clarifying language, some small additions and adding the final references. T.Shafee(Evo﹠Evo) talk 12:57, 15 May 2015 (UTC)
- I did see this; I just haven't had time to actually write anything. Googling for refs is a lot easier :) Opabinia regalis ( talk) 07:03, 20 May 2015 (UTC)
Is gene a locus or a segment, or something more generic?
This topic contains two definitions at first paragraph and as figure legend:
A gene is a locus (or region) of DNA that encodes a functional RNA or protein product, and is the molecular unit of heredity.
A gene is a segment of DNA that encodes function. A chromosome consists of a long strand of DNA containing many genes. A human chromosome can have up to 500 million base pairs of DNA with thousands of genes.
Do they define the same thing? If so, why two different expressions? Well, those represent as if a locus is a segment. Is that so?
A gene can be just a segment with a fixed position and length on genomic DNA in very simple cases. However, the nature is often more complex.
If a gene is really a segment of DNA, is a multi-exonic gene a segment? Then what distinguishes multi-cistronic genes from exons? What about nested genes? What about overlapped genes? What about a trans-spliced gene which often have discontinued fragments on different chromosomes? They are not rare cases.
I think the stated definition is just a WORKING HYPOTHESIS with too much simplification.
Finally, this is my opinion: When Mendel proposed the concept of gene (though he did not use that term), it was a concept of atomic inheritable unit. In genetics, including molecular, it is still so, except modification for quantitative trait loci.
The definition on Wikipedia is biased too much toward molecular biology, and unexplainable phenomena are left by the definition.
Wordmasterexpress ( talk) 09:00, 8 April 2016 (UTC)
- @ Wordmasterexpress: Interesting points. Both "region" and "segment" are used as non-technical synonyms for locus. Although both are vague, I agree that "region" is slightly more appropriate since it is broad enough to encompass the various oddities like nested genes etc. I've therefore updated the lead image's caption. As for the definition overall: There has been a strong molecular biology focus on the definition of a gene since the modern evolutionary synthesis, so it has shifted away from the original Mendelian definition. We've preferenced the MBOC definition as representing the mainstream scientific consensus definition. As you say, there is always going to be difficulty in precisely defining a gene, since nature is complicated. A simple and broad definition is need in the lead, so we attempt to cover more complicated scenarios in the Gene#Functional_definitions section. T.Shafee(Evo﹠Evo) talk 05:36, 9 April 2016 (UTC)
- @ Evolution and evolvability: Thank you for edits. Now I have more questions. The lead seems to say a gene is a locus and a locus is equivalent to a region. Is that so? If yes, genes are a subset of loci. I doubt the second point that a locus is equivalent to a region, but it may depending on the definition. Either positive or negative, the lead is better clarifying those, I would suggest. -- Wordmasterexpress ( talk) 01:43, 11 April 2016 (UTC)
Proposed merge with Genomic organization
I propose that Genomic organization be merged into Gene. I think that the content in the Genomic organization can easily be explained in the context of Gene, and the resulting article will be of a reasonable size. Any input is welcome :) GiggsIsLegend ( talk) 01:47, 23 June 2016 (UTC)
- Comment - Genome could be a more appropriate merge destination. T.Shafee(Evo﹠Evo) talk 04:01, 23 June 2016 (UTC)
- Do not support - both topics are notable and distinct. A reader would want information on genes while another reader would want information on the organization of genomic organization. Best Regards, Barbara (WVS) ( talk) 10:52, 30 June 2016 (UTC)
- Do not support - to clarify, I don't think Genomic organisation should be merged into Gene. It could conceivably be merged into Genome though. T.Shafee(Evo﹠Evo) talk 12:24, 30 June 2016 (UTC)
- do not support(oppose) per T.Shafee(Evo﹠Evo)-- Ozzie10aaaa ( talk) 12:44, 30 June 2016 (UTC)
- Oppose As others have said above, it is a distinct topic from Gene. If it was to be merged anywhere it should be into Genome but I think it a notable enough topic to warrant its own article. It needs a bit of work but there is good potential for it to be expanded and improved. Sarahj2107 ( talk) 13:13, 30 June 2016 (UTC)
- Oppose I agree with all the arguments put forth. Genomic organisation is an up an coming research area - it would be great if it attracted some dedicated editors. McortNGHH ( talk) 15:29, 30 June 2016 (UTC)
- Oppose. Genomic organisation is a distinct topic, and warrants its own article. However its current lede starts poorly, and may give casual readers the impression that it covers the same material as Gene. Maproom ( talk) 08:13, 3 July 2016 (UTC)
Not only proteins
The first sentence says that a gene codes for a protein. The last paragraph says that this is under discussion, that a gene also can code for functional non-coding RNAs. But that discussion was concluded long ago. I checked several of my rather oldish biology books. For example, the glossary in Hartwell, Hood, Goldberg, Reynolds, Silver, Veres; Genetics, from Genes to Genomes; 2004: ... segment of DNA in a discrete region of a chromosome that serves as a unit of function by encoding a particular RNA or Protein. -- Ettrig ( talk) 09:18, 28 October 2017 (UTC)
- Different versions of the article have mentioned RNA genes in the lead e.g.
- "A gene is a locus (or region) of DNA that encodes a functional RNA or protein product" Special:Permalink/702868373
- "The word is used extensively by the scientific community for stretches of deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) that code for a polypeptide or for an RNA chain that has a function in the organism." Special:Permalink/666720556
- There is the later section on RNA genes too. After browsing through the history, it seems that mentions of RNA genes in the lead are frequently removed to "improve clarity". Personally I think these not mentioning them introduces inaccuracy. It may be best to involve other editors of the article in this discussion to prevent any rapid reversions e.g. @ Evolution and evolvability: & @ Headbomb: who mention RNAs in their edits. -- Paul ( talk) 08:59, 29 October 2017 (UTC)
- @ Ettrig: That's a great improvement, I'm not sure about this sentence "A gene is a subsequence of DNA which codes for a molecule that has a function". What about RNA viruses? Maybe add a "usually" in there somewhere. -- Paul ( talk) 21:52, 29 October 2017 (UTC)
- Agree, RNA should be included. This is an omission of the same kind as the one I fixed. I will add or RNA. This is a bit crude. But my feeling is that nucleotides is less well known. -- Ettrig ( talk) 07:15, 30 October 2017 (UTC)
- Added RNA, with some difficulty. The current text does not cover the retrovirus case. I now empathize with the clarity argument. We want the first sentences to be correct and also very succinct and accessible. -- Ettrig ( talk) 07:22, 30 October 2017 (UTC)
- Hello all. Sorry to be late to the thread! Great work so far. You're correct that it was high time for the lead to be brought into line with the rest of the article. My main outstanding issue is the term "subsequence" in the first sentence. I think that "sequence" would suffice and I don't think extra useful specificity is added to the definition by using "subsequence". Indeed it's the only time the term is used in the article.
T.Shafee(Evo&Evo)
talk
00:36, 31 October 2017 (UTC)
- Yes, indeed. And thanks to @ Chiswick Chap:. -- Ettrig ( talk) 12:43, 31 October 2017 (UTC)
- Hello all. Sorry to be late to the thread! Great work so far. You're correct that it was high time for the lead to be brought into line with the rest of the article. My main outstanding issue is the term "subsequence" in the first sentence. I think that "sequence" would suffice and I don't think extra useful specificity is added to the definition by using "subsequence". Indeed it's the only time the term is used in the article.
T.Shafee(Evo&Evo)
talk
00:36, 31 October 2017 (UTC)
Definitions
The definition of gene is probably one of the most important parts of this article. At the risk of opening Pandora's box on, it might be worth checking that we're using the best available definitions. Currently we list three definitions:
-
Gene#Lead - A gene is the molecular unit of
heredity of a living
organism.
[1]
- Clear and simple, but lacks any mention of function.
-
Gene#Lead - Therefore, a modern working definition is "a
locatable region of
genomic sequence, corresponding to a unit of inheritance, which is associated with regulatory regions, transcribed regions, and/or other functional sequence regions".
[2]
[3]
- The only definition that we are directly quoting from a reference and seems verbose to me.
- The use of "associated with " feels unnecessarily confusing.
- Also, close to tautology: "genomic sequence... which is associated with... other functional sequence regions".
-
Gene#Functional definitions - A broad operational definition is sometimes used to encompass the complexity of these diverse phenomena, where a gene is defined as a union of genomic sequences encoding a coherent set of potentially overlapping functional products.
[4]
- It think this is fine but omits heredity. Does it need to? What phenomena fit this definition but are not heritable?
- Exclusion of regulatory sequence doesn't seem hugely necessary.
Overall I think that we do need a simple layman's definition as well as a more nuanced, broader, technical definition. Both probably need to encompass:
- Molecular / sequence / locus / encoding
- Heritable
- Functional / expressed
T.Shafee(Evo﹠Evo) talk 01:10, 14 June 2015 (UTC)
[1] perhaps?..-- Ozzie10aaaa ( talk) 01:22, 14 June 2015 (UTC)
- You'd think that developing a really good single-sentence definition would be really important, but... I just can't get myself too worked up about it. It's a fuzzy concept; people use slightly different definitions for different purposes; the question of 'what do we annotate as a gene' is subtly but significantly different; etc. etc. I looked around a little specifically in the education literature and found this hilarious "simplified" model: DOI 10.1007/s11191-008-9161-7. (This thesis studying representations of genes in textbooks is pretty interesting: [2] Not so useful for our purposes, though.) Opabinia regalis ( talk) 05:38, 16 June 2015 (UTC)
- You're probably right. I might still have a crack at polishing the currently existing sentences over the next few days, but we do a pretty decent job of describing the key elements in the functional structure section. Also, one of the papers that comprises the thesis you mentioned was actually a pretty good history of the definition. T.Shafee(Evo﹠Evo) talk 12:08, 19 June 2015 (UTC)
I've now updated the definitions in the article:
- a region of DNA that controls a discrete, hereditary trait in an organism
- based on the the definition from the MBOC glossary since I think this more clearly includes the key elements [5]: Glossary
- any discrete region of heritable, genomic sequence which affect an organism's traits by being expressed as a functional product or by regulating expression
- still based on the same references, but reworded to simplify the language and sentence structure whilst still being thorough.
- a union of genomic sequences encoding a coherent set of potentially overlapping functional products
- left alone with only a minor change to the sentence before it.
I think that these should suffice as a simple summary for a brief visitor, as well as a more detailed definition for anyone more interested. T.Shafee(Evo﹠Evo) talk 12:16, 25 June 2015 (UTC)
- so, is gene a material, address, or information? If material, it should be DNA (or RNA for minor cases), which is very clear, but remains some unexplainable attribute. If address, even more unexplainability, which seems current definition. If information, I find no problem when I consider genetic materials are just media of genetic information, but it's just my impression. What is the formal, gold standard consensus of current definition? Current lede seems tautological, or pretty much redundant at least, and does not seem to provide clear idea.
- A gene is a locus (or region) of blah blah blha defines a gene is an address.
- Genes can acquire mutations in their sequence, leading to different variants, known as alleles defines a gene contains sequence a part of which is mutable in their address. It also defines the mutated variant is called allele. The question is that, if an allele is also defined as sequence, can it be a gene when sequence is contained in the address (gene). Also if a gene is defined as sequence, can it point an address when many experiments had suggested that genes can function regardless of, even out of, the chromosomal location.
- Therefore, a broad, modern working definition of a gene is any discrete locus of heritable, genomic sequence. might represent best, but it still states that a gene is a locus. Uh, could be tautology. Well, polygenes?
-- Wordmasterexpress ( talk) 09:55, 5 October 2016 (UTC)
- Dear @ Wordmasterexpress, Evolution and evolvability, Ozzie10aaaa, and Opabinia regalis: I appreciate your efforts to come up with a definition. When writing an intro paragraph, I think it is allowed to use a heuristic, where short and efficient may take the overhand over "complete". For the latter, one may expect the user to read the article. The first paragraph is what people see on other pages where the word is used and linked so that when readers hoover over the word, they get a pop-up of the first paragraph to remind them what the word is about or give a heuristic that makes them understand what it is about. Right? It's quite popular to refer to things as "the smallest unit of" e.g. a genetic codification of a protein that will eventually result in a trait. Isn't that good enough for most popular readings or when you come across the word in a wikipedia article? Look e.g. at how " engram" - the unit of cognitive information inside the brain - is defined, or atom, or molecule. Sincerely, SvenAERTS ( talk) 03:45, 24 January 2020 (UTC)
refs
References
- ^ Slack, J.M.W. Genes-A Very Short Introduction. Oxford University Press 2014
- ^ Pearson H (May 2006). "Genetics: what is a gene?". Nature. 441 (7092): 398–401. Bibcode: 2006Natur.441..398P. doi: 10.1038/441398a. PMID 16724031.
- ^ Pennisi E (June 2007). "Genomics. DNA study forces rethink of what it means to be a gene". Science. 316 (5831): 1556–1557. doi: 10.1126/science.316.5831.1556. PMID 17569836.
- ^ Gerstein MB, Bruce C, Rozowsky JS, Zheng D, Du J, Korbel JO, Emanuelsson O, Zhang ZD, Weissman S, Snyder M (June 2007). "What is a gene, post-ENCODE? History and updated definition". Genome Research. 17 (6): 669–681. doi: 10.1101/gr.6339607. PMID 17567988.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). Molecular Biology of the Cell (Fourth ed.). New York: Garland Science. ISBN 978-0-8153-3218-3.
Requested Edit: Structure and Function / Structure chapter referencing own research
| This edit request by an editor with a conflict of interest has now been answered. |
I request a volunteer to review a proposed reference to own research. Details below.
At the end of this paragraph:
The transcribed pre-mRNA contains untranslated regions at both ends which contain a ribosome binding site, terminator, and start and stop codons.[42] In addition, most eukaryotic open reading frames contain untranslated introns, which are removed and exons, which are connected together in a process known as RNA splicing. Finally, the ends of gene transcripts are defined by CPA sites, where newly produced pre-mRNA gets cleaved and a string of ~200 adenosine monophosphates is added at 3′ end. PolyA tail protects mature mRNA from degradation and has other functions, affecting translation, localization, and transport of the transcript from the nucleus. Splicing, followed by CPA generate the final mature mRNA which encodes the protein or RNA product.[43] Although the general mechanisms defining locations of human genes are known, identification of the exact factors regulating these cellular processes is an area of active research. For example, known sequence features in 3′-UTR can only explain half of all human gene ends. [CITATION PROPOSED HERE]
I am requesting a reference to a recent paper from our lab - https://pubmed.ncbi.nlm.nih.gov/34104882/
Wikipedia guidelines suggest citing review papers first, however, this is the first study that assesses whether known sequence elements are sufficient to predict human gene ends. Strikingly, we found that known elements can only explain half of all human gene ends. This project was aimed at answering fundamental questions of human gene definition. The results are significant and highlight the magnitude of missing information in the current understanding of processes human cells utilize to locate their genes. Therefore, this study is the most relevant and most recent source on the subject.
To justify expert knowledge - this paper comes from Hughes lab at the University of Toronto. Dr. Timothy R Hughes is a John W. Billes Chair of Medical Research and Canada Research Chair in Decoding Gene Regulation, one of the most cited Canadian researchers.
 Done with minor edits (missing articles, punctuation).
Heartmusic678 (
talk)
15:06, 22 July 2021 (UTC)
Done with minor edits (missing articles, punctuation).
Heartmusic678 (
talk)
15:06, 22 July 2021 (UTC)
Gwne
Hjaj 2409:4052:4D1F:E11E:8CDF:AA35:8201:7E15 ( talk) 19:03, 20 March 2022 (UTC)
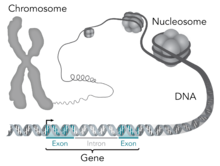
Difference between gene and chromosome
How does the gene differ from a chromosome? 41.116.8.55 ( talk) 16:08, 13 January 2022 (UTC)
- Genes control cell function and overlay the chromosome, which is the unit of transmission, all of which is a single molecule of DNA. Don't know why I answered that. Habit. 2600:8807:1C0A:9600:C53E:7E54:BE58:FCED ( talk) 12:08, 31 January 2023 (UTC)
Defining "gene"
- Continued from talk:Gene structure#Definition of a gene
Arguably, one of the most important purposes of this article is to come up with a good definition of "gene." I'm not happy with the current definition for many reasons so I'd like to start a discussion about how we can change it. Check out an old blog post of mine from 2007 where I define a gene as, "A gene is a DNA sequence that is transcribed to produce a functional product."
What Is a Gene? https://sandwalk.blogspot.com/2007/01/what-is-gene.html
This is the standard definition dating back (at least) to Watson's textbook "Molecular Biology of the Gene" (1965). The idea is that a gene is a transcription unit and there are two kinds of genes: protein-coding genes and noncoding genes. In addition, the product has to have a function - junk RNA transcripts do not define genes.
All of the best textbooks continue to use this standard definition and nothing has changed since the publication of the human genome sequence in spite of all the rhetoric that has been published. See my post on Gerald Fink for an example of the kind of misinformation that we have to correct in this article.
Gerald Fink promotes a new definition of a gene https://sandwalk.blogspot.com/2019/09/gerald-fink-promotes-new-definition-of.html
Genome42 ( talk) 17:07, 1 August 2022 (UTC)
- I use a carefully defined single sentence definition meant to encompass all cases for identified genes. " A gene is a chromosomal locus that governs the expression of a heritable trait or is homologous to a known gene." Genes have multiple aspects each with its own homologs, so in practice it still is complicated. My course also contains a longer molecular definition. "A gene is a nucleic acid, all or in part, that is composed of dispersed, modular units of molecular function that have an emergent property of immediate or secondary cellular activity. Gene function is not an inherent property of the nucleic acid sequence but depends on the cellular context and activity. The gene is the total of all the nucleic acid sequences that significantly influence the achievement of that cellular activity and share a dependency on the function of at least one single modular unit. While a certain number of correct modular units of function are required, there appears no limit to the number of such units or their dispersal over a single chromosome." The word choice is generic and the definition requires descriptions or example of molecular function (start, stop; +1, terminator;5', 3' splice sites; etc.. Dr. Rogers 2600:8807:1C0A:9600:C53E:7E54:BE58:FCED ( talk) 12:53, 31 January 2023 (UTC)
Genome, Molecular Evolution, Inheritance, Gene Expression
Do we really need extensive discussion of these topics here since there are separate articles for all of them? The problem is that when the main articles are updated and corrected the information here then conflicts with the main article.
For example, under "Molecular evolution: Mutation" we find the following statement "This means that each generation, each human genome accumulates 1–2 new mutations." This is wrong in two possible ways. If it refers to a cell generation then you can do a simple calculation based on the preceding statement that the error rate is "as low as 10−8 per nucleotide per replication." Since there are 6.1 x 109 nucleotides being synthesized that means at least 62 mutations or 31 in each daughter cell. (The actual mutation rate due to DNA replication errors is 10-10 per base pair or less than one per replication.)
If it refers to human generations then it's also wrong. Each newborn baby has about 100 new spontaneous mutations. (The Wikipedia mutation rate article says 64 new mutations per generation but that needs to be updated.)
This is just one example of the problems that arise when there are too many editors duplicating information in these articles. We need to concentrate our efforts on a few key articles that we link to whenever the topic comes up somewhere else. That way we don't risk spreading and perpetuating misinformation because the updates and corrections aren't propagated to the other articles.
I'm proposing a drastic change because it means deleting, or substantially reducing, a lot of stuff in this article.
What do you think? Genome42 ( talk) 17:37, 7 March 2023 (UTC)
- Hi Larry, I just googled for the WP policy on redundancy (of course there would be a policy). You can see it here
Wikipedia:Abundance and redundancy, I agree it's a pain having conflicting information between articles &/or maintaining redundant information. But it looks like you might be better off correcting the more obvious flaws you see here, but taking a lighter touch than wholesale removal/reduction of topics. --
Paul (
talk)
20:27, 8 March 2023 (UTC)
- Paul, I’m can’t imagine why we need an extensive discussion of molecular evolution (including mutation) in an article on “gene.” Are you saying that we should keep it just because deleting might cause trouble with Wikipedians or is it because you think it’s important to discuss molecular evolution here?
Genome42 (
talk)
13:23, 9 March 2023 (UTC)
- Larry, no that is very much not what I'm saying. If the text is clearly tangential to the main topic, then by all means move it somewhere else, or remove/reduce. I thought I was responding to your query about redundant information between general and specific articles. A case could be made to split some of this article out into other topics. -- Paul ( talk) 08:08, 10 March 2023 (UTC)
- Paul, I’m can’t imagine why we need an extensive discussion of molecular evolution (including mutation) in an article on “gene.” Are you saying that we should keep it just because deleting might cause trouble with Wikipedians or is it because you think it’s important to discuss molecular evolution here?
Genome42 (
talk)
13:23, 9 March 2023 (UTC)
- Both pages even cite the
same 1998 source for those different human mutation rate numbers! Definitely needs an in-depth read to work out which is more accurate (or if there have been new alternative calculations done since we've so many more sequenced genomes now than back then!). Some level of overlap on broad topics like this is inevitable, since almost all of the material is also covered in more detail in other narrower-scope pages, so we'll always have some amount of maintenance to do no matter how long the page. However it's currently ~8000 words long so certainly be pruned down to 6000 with judicious removal of material that is a) less core to an expected average reader and b) covered elsewhere easily navigated to.
T.Shafee(Evo&Evo)
talk
02:53, 9 March 2023 (UTC)
- @ Evolution and evolvability I’m quite knowledgeable on this topic so I don’t really need to do any more research. The main issue is finding the right citations and figuring out how to deal with editors who might want to keep the incorrect information just because it is supported by a “reliable source” (sensu Wikipedia). It would be a lot easier to fix the mutation article and then link to it from here if we think it’s really necessary to discuss mutation in this article. What do you think? Should I risk being accused of edit wars on both articles? Genome42 ( talk) 13:32, 9 March 2023 (UTC)
| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | Archive 2 | Archive 3 |
Deleted text?
Something is missing in this sentence: "Because they use RNA to store In 2006, French researchers came across a puzzling example of RNA-mediated inheritance in mice." Perhaps someone could fix? Jimjamjak ( talk) 15:38, 11 April 2012 (UTC)
- It was an edit in March that broke the text, and I have restored the original. Johnuniq ( talk) 02:05, 12 April 2012 (UTC)
2012 GA Review
| GA toolbox |
|---|
| Reviewing |
- This review is transcluded from Talk:Gene/GA1. The edit link for this section can be used to add comments to the review.
Reviewer: Sasata ( talk · contribs) 20:23, 19 August 2012 (UTC)
I'll review this. Sasata ( talk) 20:23, 19 August 2012 (UTC)
Before we start, could you please ensure that all paragraphs, and end-of-paragraph sentences have citations? This will help me (and other readers) verify the material. The material is pretty basic, and so a general text like Genes (Lewin) or perhaps a middle-level university genetics text would work nicely for this purpose. Sasata ( talk) 23:27, 19 August 2012 (UTC)
- There's been no editing activity on this article by the nominator, so I'm going to close this GAN now. Quite a bit a work needs to be with sourcing to meet the GA criteria. Sasata ( talk) 15:35, 4 September 2012 (UTC)
Gene exactly?
"though there still are controversies about what plays the role of the genetic material.[1]" Strikes me as a very dubious opening. DNA & RNA are all that need be mentioned (in the beginning) for genes. Prions and epigenetic factors can be left for later. I tried to find out about ref 1, without buying it, by reading articles by the editors. Plutynski is a normal biologist. Sarkar is an anti-reductionist, but his webpage fails to provide any links to his articles. How can a wiki throw doubt about DNA being a genetic material in the introduction? It is similar to starting an AIDS article with a discussion of HIV denialists. OK at the end, not at the beginning. Peggy hopper ( talk) 03:23, 13 April 2013 (UTC) "
- Yes, this is terrible in the opening. Perhaps something like "Although information is transmitted from parent to offspring in many ways, a gene is... " It has been a while since I have looked at this article, maybe I'll come back to this.
Abductive (
reasoning)
04:33, 13 April 2013 (UTC)
- I've deleted this considering there is no known controversy about DNA being the genetic material and considering there were no responses to this post giving any reason to keep it. I can see how it might be handy to have mention of other mechanisms of inheritance such as epigenetics somewhere else in this article but I'll leave that for other people if they so choose. 2403:7900:ADE1:A1DE:250:56FF:FEA6:3B1F ( talk) 09:09, 19 March 2014 (UTC)
"The genetic code is nearly the same for all known organisms." is false. Human mtDNA has a different genetic code than human nuclear DNA. This discovery was crucial in proving the symbiotic origin of eukaryotes. (ref. Lynn Margulis) (Comp Biochem Physiol B. 1993 Nov;106(3):489-94. Evolutionary changes in the genetic code. Jukes TH, Osawa S.) Peggy hopper ( talk) 04:03, 13 April 2013 (UTC)
- Here "nearly" encapsulates that fact. Readers can click on genetic code to find out more. Abductive ( reasoning) 04:33, 13 April 2013 (UTC)
discontinuing inheritance
The history section uses the expression discontinuing inheritance. I guess this means that a phenotypic trait can be observable in one generation, "disappear" i a following generation and then reappear in an even later generation. Regardless of whether this or something else is meant, it needs to be explained in a way that is more understandable for the general reader. — Preceding unsigned comment added by Ettrig ( talk • contribs)
- Should be 'discontinuous', and you're right. There's a lot of clunky prose in this article at the moment. Opabinia regalis ( talk) 09:12, 7 April 2015 (UTC)
2015 GA Review (April)
| GA toolbox |
|---|
| Reviewing |
- This review is transcluded from Talk:Gene/GA2. The edit link for this section can be used to add comments to the review.
Reviewer: Cerebellum ( talk · contribs) 01:25, 10 April 2015 (UTC)
Hello! I will be reviewing this article. --
Cerebellum (
talk)
01:25, 10 April 2015 (UTC)
- It is reasonably well written.
- a (prose, no copyvios, spelling and grammar):
 b (
MoS for
lead,
layout,
word choice,
fiction, and
lists):
b (
MoS for
lead,
layout,
word choice,
fiction, and
lists): 
- In the "Translation" section you used the word "ligates," which I do not know. Is there a more common word you could use there? Also, the first sentence of the "Gene targeting and implications" section left me confused. You say that gene targeting provides "mouse models for studying the roles of individual genes," but the sentence seems abrupt and I'm not really sure why we are talking about mice all of a sudden or what mouse models are. Maybe start off with a broad intro sentence to transition from the previous section on genes in evolution, and explain why we are now talking about mice whereas the rest of the article seemed to be about genes in all forms of life. As far as layout goes, I recommend putting the two sections on the concept of a gene ("Changing concept" and "evolutionary concept of a gene") in sequence, right now they are split up by the gene targeting paragraph.
- a (prose, no copyvios, spelling and grammar):
- It is factually accurate and
verifiable.
- a (
reference section):
 b (citations to
reliable sources):
b (citations to
reliable sources):  c (
OR):
c (
OR): 
- I think the 2012 review was too harsh on this but some of the same criticism applies. Per WP:SCG you don't need a ref for every paragraph if the information all comes from a basic textbook, but it's not clear to me where some information comes from. For example, the section on Mendelian inheritance has no references. I assume most of the information in later sections can be found in Molecular Biology of the Cell, but is that the case for Mendelian inheritance as well? Please add at least one reference to that section. Also, you always need to cite direct quotations, so please provide a reference for the Williams definition of a gene in the "Evolutionary concept of a gene" section.
- a (
reference section):
- It is broad in its coverage.
- a (
major aspects):
 b (
focused):
b (
focused): 
- a (
major aspects):
- It follows the
neutral point of view policy.
- Fair representation without bias:

- Fair representation without bias:
- It is stable.
- No edit wars, etc.:

- No edit wars, etc.:
- It is illustrated by
images and other media, where possible and appropriate.
- a (images are tagged and non-free content have
fair use rationales):
 b (
appropriate use with
suitable captions):
b (
appropriate use with
suitable captions): 
- Great use of diagrams!
- a (images are tagged and non-free content have
fair use rationales):
- Overall:
- Pass/Fail:

Good article, but I'm putting it on hold for now so you can make some tweaks. Cerebellum ( talk) 17:15, 11 April 2015 (UTC)
- Pass/Fail:
- Thanks everyone for your work on this article this week, in particular BlueMoonset for identifying the copyright issue, Opabinia regalis for correcting it, and Evolution and evolvability for fixes throughout. Unfortunately some of the issues from the review, in particular regarding sourcing, have not been fixed. Because of that I have to fail the article for now, but please let me know if you renominate it and I'll take another look. -- Cerebellum ( talk) 03:26, 19 April 2015 (UTC)
Comment
I thought I'd take a look after recent exchanges about the nomination on the GAN talk page, which ended by noting that the nomination was under review. It's an impressive article, but at the moment does not meet some of the criteria
It seems to me that the article has a classic violation of
WP:LEAD, a GA criterion: Apart from trivial basic facts, significant information should not appear in the lead if it is not covered in the remainder of the article.
The lead's third paragraph is solely about a topic, "big genes", that is only mentioned there; I could not find the phrase "big genes" anywhere in the body.
I'm completely at sea as to the sourcing, since this is a very long article where so much of the material is not sourced; I don't see how any assumption can be made about what came from the textbook (which is only cited twice) and what didn't. (Also, textbooks are big: these citations should be to a page or page range, not to an immense tome where it's impractical to find the information being referenced.) I think that every subsection should be sourced, not merely every section, and more than a source for a parenthetical comment at that (as in "Genetic code", which is the only citation in the entire section). Genes are complicated and involve technical explanations, as is plain in this article; there needs to be concomitant sourcing. As it says, Complex, current, or controversial subjects may require many citations
in the lead; this is even more true of the article body of a very complex scientific topic.
BlueMoonset (
talk)
23:46, 12 April 2015 (UTC)
- I just ran Earwig's Copyvio Detector on the article, and found significant copyvio in the article, enough to stop this nomination in its tracks until a thorough check and cleanup job has been completed. For example, a great deal of the "Gene targeting and implications" section is taken from the FN24 (www.biolsci.org) source; the second paragraph is copied almost verbatim in its entirety, as you can see in this report, and much of the first paragraph is very like FN25. BlueMoonset ( talk) 03:23, 13 April 2015 (UTC)
- Cleaned up the gene-targeting section (which was also rather undue). I had planned/promised to work on the text of this article awhile back but just haven't had the time. Images look fantastic though! Opabinia regalis ( talk) 19:41, 15 April 2015 (UTC)
Prose suggestions
Below are the main issues that I've spotted or now. Hopefully they're logically laid out. Let me know if you agree or disagree with any. T.Shafee(Evo﹠Evo) talk 00:02, 4 April 2015 (UTC)
RNA genes
Certainly an important topic but it is currently brought up in several places.
- History
- Physical definitions#RNA genes and genomes in the world (before description of protein-coding genes)
- Changing concept
This leads to it being massively over-weighted in the article when really it needs only to be a single paragraph total. RNA genes are also not referred to in the images at all. We also need to clearly distinguish between a gene encoded by RNA (e.g. genes in an RNA virus) versus a gene that encodes a functional RNA product (e.g. genes for tRNAs or siRNAs).
History and concepts
Perhaps the evolutionary concept and changing concept sections can be folded into the history section? That way the History section is structured:
- How genes were understood before we knew about DNA
- The understanding that DNA genes that encode proteins
- Minor expansion of the simple model of #2 (RNA genomes, functional RNAs, splice variants etc).
Descriptions of DNA
There seems to be some overlap between Physical definitions#Functional structure of a gene and Gene expression#Genetic code. I would suggest that the descriptions of nucleotide biochemistry could be reduced. I think it would be better to increase the focus on larger structure (promoters, ORFs, terminators, enhancers, introns etc).
Mutation
Perhaps this can be combined into either replication or evolution sections?
- Agreed with pretty much all of this. I made some grouchier and less useful notes
in my sandbox last week, but haven't had time to actually do anything about it yet.
Opabinia regalis (
talk)
03:22, 5 April 2015 (UTC)
- Good work on today's edits
Opabinia regalis. I'm still thinking of moving the evolutionary concept and functional concept sections up into History since they seem to fit with a continuing evolution of our understanding. However I don't want to make the history section balloon in size again.
T.Shafee(Evo﹠Evo)
talk
12:30, 16 April 2015 (UTC)
- Thanks! I did some overall restructuring without changing the text much, moving the bottom evolutionary gene section into history and reorganizing some of the middle. I think some of the power of those great gene structure diagrams was being lost by having such detailed information before basics like transcription.
- Still not sure what to do with the mutation section, but the article is really missing a discussion of sequencing and sequence analysis and homology, so maybe that (ie, the interpretation of variation) and mutation can go under a new top-level section on something like "sequence variation"? Opabinia regalis ( talk) 06:52, 18 April 2015 (UTC)
- Good work on today's edits
Opabinia regalis. I'm still thinking of moving the evolutionary concept and functional concept sections up into History since they seem to fit with a continuing evolution of our understanding. However I don't want to make the history section balloon in size again.
T.Shafee(Evo﹠Evo)
talk
12:30, 16 April 2015 (UTC)
In a family of four kids, one will be sick

Well, no, but that seems to be what this graph implies. The alt text is equally problematic, and I've only just somewhat clarified this in the caption. I think the notion of probability needs to be more clearly communicated. Samsara 05:01, 19 April 2015 (UTC)
- @ Samsara: Not sure what you mean here. I don't think anybody's substantively edited that section of text yet, but the image seems clear enough. Is the issue that "affected" implies a disease but the description is just of two alleles, without specifying the trait? Representing the possible outcomes is pretty standard in this kind of diagram; I don't think it implies a probability error any more than a Punnett square does.
- The only criticism I have of the image/alt text is that "white" might not be the best color to use as an illustration, since "white" is in fact the common name of a trait some humans have.
Opabinia regalis (
talk)
05:48, 19 April 2015 (UTC)
- My concern is that there is no explicit statement that this deals with probabilities. A naive reader might think that if you have four kids, they turn out 1:2:1. The "alt" parameter in the article in fact reinforces this idea by explicitly talking about four children. But there are no four children in reality. There are four probabilities. Unfortunately, if you take a close look, this is a problem with most representations of this kind. Thinking outside the box, I wonder if it wouldn't make sense to have a graph that shows a number of different families, where the sum of the children ends up in a 1:2:1 ratio. Also, that would allow us to have green male with blue female as well as green female with blue male pairings, if that helps to avoid the impression that this refers to any real trait.
Samsara
06:13, 19 April 2015 (UTC)
- Hmm. The alt text, as I understand it, is intended to describe rather than interpret the image, and should not duplicate material in the caption. For that purpose the current text seems adequate. Your suggested expanded graph sounds a little too bulky for this article, but could fit well in Mendelian inheritance or genetics (which really needs a going-over to reduce redundancy with this article). Opabinia regalis ( talk) 21:17, 19 April 2015 (UTC)
- My concern is that there is no explicit statement that this deals with probabilities. A naive reader might think that if you have four kids, they turn out 1:2:1. The "alt" parameter in the article in fact reinforces this idea by explicitly talking about four children. But there are no four children in reality. There are four probabilities. Unfortunately, if you take a close look, this is a problem with most representations of this kind. Thinking outside the box, I wonder if it wouldn't make sense to have a graph that shows a number of different families, where the sum of the children ends up in a 1:2:1 ratio. Also, that would allow us to have green male with blue female as well as green female with blue male pairings, if that helps to avoid the impression that this refers to any real trait.
Samsara
06:13, 19 April 2015 (UTC)
At least the text IN the figure should use plural forms rather than singular. -- Ettrig ( talk) 12:24, 27 April 2015 (UTC)
Clarify the difference among gene, locus and allele (previously GA3)
- Note: comments below were initially added as the Good Article review in error. See here for details.
Well, allele seems to be clearly defined. What about gene and locus? In a classical sense, a locus can be a placeholder for the corresponding alleles. Is it also applicable to a gene? If so, what is the difference? If no difference, locus shall be a synonym of gene. By the way, what about a null allele, where a placeholder can be nowhere on the DNA? I am totally confused. ヒストリ案 ( talk) 13:31, 3 June 2015 (UTC)
- Good point, We only mentioned the term locus once! I'm adding a section on Locus/ Allele/ Gene clarification to the Gene#Chromosomes and Gene#Mendelian inheritance sections. I think that probably explaining null alleles is beyond the scope of this article, but I'll make sure that it comes up in both the allele and locus articles. T.Shafee(Evo﹠Evo) talk 12:08, 5 June 2015 (UTC)
- T.Shafee(Evo﹠Evo), thank you for improvement. BTW, what happen if "quantitative trait loci" taken into account? Is it a completely different concept, or share some? If the former is the case, why do we use the same term? If the latter, the idea of "a" gene is already confused... ヒストリ案 ( talk) 13:35, 8 June 2015 (UTC)
- @ ヒストリ案: The term Quantitative trait locus isn't quite the same as a gene. A QTL is a region of DNA on a chromosome known to be correlated with some phenotype change, but it can be a gene, or several genes, or part of a gene. Identifying the QTL of some phenotype can be a first step in finding a gene related to that trait. It's actually a bit of an old fashioned term which was used more extensively when we weren't able to exactly define what the DNA sequence of the gene was. It is probably more relevant to the genetics page but could certainly go in the 'see also' section for the moment. T.Shafee(Evo﹠Evo) talk 13:04, 10 June 2015 (UTC)
- ヒストリ案, I notice that this is your first-ever edit under this account, and perhaps ever on Wikipedia. Are you prepared to fully review this nomination according to the WP:WIAGA criteria, which is what is required here? If not, we should probably get a new reviewer to undertake the complete review beyond your initial questions. Thanks. BlueMoonset ( talk) 17:07, 5 June 2015 (UTC)
- BlueMoonset, I apologize for misplacement of my comment. Should I replace all the above somewhere appropriate, or delete them? Thank you for advice. ヒストリ案 ( talk) 13:27, 8 June 2015 (UTC)
- @ ヒストリ案: Your comments are still useful and definitely won't be deleted. The link that you clicked to create this section of the page are part of a process called Good article review in which an editor volunteers themself to do a thorough read-through and assessment of whether the page should be promoted to be Good article status (for example this previous review). If you were not intending to nominate yourself as the GA reviewer then we can just move your comments into the non-review part of the gene talk page. T.Shafee(Evo﹠Evo) talk 12:06, 9 June 2015 (UTC)
- Any progress so far? The current lede seems to define that a gene is a locus. Is that widely accepted, or not? -- Wordmasterexpress ( talk) 09:22, 5 October 2016 (UTC)
- A paper that clarifies the terminology (incl. gene, locus and allele among of the tens of other terms): Genetic Terminology (2012). Cheater no1 ( talk) 18:39, 9 March 2020 (UTC)
Image suggestions
Below are my opinions for changes that could be made to the article's images. T.Shafee(Evo﹠Evo) talk 00:40, 4 April 2015 (UTC)
| ===Lead image=== |
I don't think introns need to be introduced in the very first image. Certainly not in favour over other genetic elements like promoters.


| ||||||||||||||||
| DNA nucleotides |
This definitely doesn't need to be in the lead section and elements needs to relate to other images.

| ||||||||||||||||
| Gregor Mendel |
You're all good Gregor. You can stay as you are.
| ||||||||||||||||
| Punnett square |
Currently odd colour choices. Significance of the background colours not clear.
| ||||||||||||||||
| Gene structure |
Currently no image in the article uses the traditional 'arrow' symbol of an ORF or shows all the elements of a gene together



| ||||||||||||||||
| RNA codons |
It's not immediately obvious that the letters relate to the nucleotides in the DNA image. Similarly, it's not obvious that the right hand side words are amino acids in a protein.
| ||||||||||||||||
| Function pie chart |
No fundamental problem with the image, but is there something more useful that we could show? It's no better or worse than the old gene number table that the article had 10 years ago.

| ||||||||||||||||
RNA-coding gene
no image yet |
I think it would be good to have some side-by-side comparison of a protein-coding gene vs an RNA gene (e.g tRNA).

| ||||||||||||||||
Inheritance
no image yet |
 There's currently no image actually showing genes being inherited from parent to offspring. I think this could be more useful that the Punnett square
|
Hey there, I'm a scientific illustrator and just by chance came across this article and saw you are pushing it to GA. Here's a selection of the illustrations I made for Wikipedia. As time permits I might be able to help out with improving the images of this article. Once you've decided which ones need to be improved and how, leave me a message on my talk page if you need any help (I'm not online here that often anymore nowadays). PS. The pie chart should definitely be replaced with a (horizontal) bar chart. See here why. -- SPLETTE :] How's my driving? 01:56, 4 April 2015 (UTC)
- I'm currently working on a few of the images.
- Lead image (sorry to accidentally compete User:splette, perhaps we can combine?)
- DNA structure
- RNA gene
- Hopefully should be done in next couple of days. T.Shafee(Evo﹠Evo) talk 09:06, 6 April 2015 (UTC)
Refs
- ^ Watson, JD, Baker TA, Bell SP, Gann A, Levine M, Losick R. (2004). “Ch9-10”, Molecular Biology of the Gene, 5th ed., Peason Benjamin Cummings; CSHL Press.
- ^ "Integr8 - A.thaliana Genome Statistics:".
- ^ "Understanding the Basics". The Human Genome Project. Retrieved 26 April 2015.
- ^ "WS227 Release Letter". WormBase. 10 August 2011. Retrieved 2013-11-19.
- ^ Yu, J. (5 April 2002). "A Draft Sequence of the Rice Genome (Oryza sativa L. ssp. indica)". Science. 296 (5565): 79–92. doi: 10.1126/science.1068037.
- ^ Anderson, S.; Bankier, A. T.; Barrell, B. G.; de Bruijn, M. H. L.; Coulson, A. R.; Drouin, J.; Eperon, I. C.; Nierlich, D. P.; Roe, B. A.; Sanger, F.; Schreier, P. H.; Smith, A. J. H.; Staden, R.; Young, I. G. (9 April 1981). "Sequence and organization of the human mitochondrial genome". Nature. 290 (5806): 457–465. doi: 10.1038/290457a0.
- ^ Adams, M. D. (24 March 2000). "The Genome Sequence of Drosophila melanogaster". Science. 287 (5461): 2185–2195. doi: 10.1126/science.287.5461.2185.
-
^ Pertea, Mihaela; Salzberg, Steven L (2010). "Between a chicken and a grape: estimating the number of human genes". Genome Biology. 11 (5): 206.
doi:
10.1186/gb-2010-11-5-206.
{{ cite journal}}: CS1 maint: unflagged free DOI ( link)
Revision of lede
The lede needs further work. For now, I've taken out some of the grammatical problems as well as corrected the following fallacies:
- that genes take effect on only one thing, which is later (correctly) contradicted in the article
- that genes must code for proteins (nope, ribosomal RNA and transfer RNA genes are obvious and essential counter-examples)
- that genes (or proteins) perform only cellular functions (nope, those would typically be housekeeping genes, one of many categories!)
I'm also wondering if it shouldn't be mentioned, for accuracy's sake, that RNA can also carry genes (NB: retrovirus).
Regards,
Samsara 10:12, 29 June 2015 (UTC)
- @
Samsara: Thanks for the lead section edits. You're probably right that we should mention ncRNA products early in the lead. Perhaps genes affecting multiple traits can be incorporate into the sentence "Most biological traits are under the influence of many genes as well as the environment"? RNA genomes are left to paragraph 3 of the lead currently, and I think this is reasonable since they're the exception, otherwise it makes the first sentence confusing with both RNA genomes and ncRNA products.
T.Shafee(Evo﹠Evo)
talk
12:01, 29 June 2015 (UTC)
- Yes, that is why I brought it here for discussion. Ideally imo, we would have a diagram that shows that RNA can transmit genes, but that the route via DNA is required for gene expression. Samsara 12:04, 29 June 2015 (UTC)
2015 GA Review (July)
| GA toolbox |
|---|
| Reviewing |
- This review is transcluded from Talk:Gene/GA4. The edit link for this section can be used to add comments to the review.
Reviewer: Cerebellum ( talk · contribs) 16:46, 25 July 2015 (UTC)
Sorry it took me so long to get to this, but I'm ready to do a second review now. --
Cerebellum (
talk)
16:46, 25 July 2015 (UTC)
- It is reasonably well written.
- a (prose, no copyvios, spelling and grammar):
 b (
MoS for
lead,
layout,
word choice,
fiction, and
lists):
b (
MoS for
lead,
layout,
word choice,
fiction, and
lists): 
- In the first sentence of the lead, would it be OK to replace A gene is a locus (or region) of DNA with A gene is a segment of DNA? It would be simpler, and it's what the caption of the first image says. However, if there's a big difference between the terms locus and segment then it can stay as it is. Other than that though, I made a few minor changes but the prose is generally good.
- a (prose, no copyvios, spelling and grammar):
- It is factually accurate and
verifiable.
- a (
reference section):
 b (citations to
reliable sources):
b (citations to
reliable sources):  c (
OR):
c (
OR): 
- Great job specifying the page numbers from Molecular Biology of the Cell! I added a couple of refs to the section on Mendel.
- a (
reference section):
- It is broad in its coverage.
- a (
major aspects):
 b (
focused):
b (
focused): 
- a (
major aspects):
- It follows the
neutral point of view policy.
- Fair representation without bias:

- Fair representation without bias:
- It is stable.
- No edit wars, etc.:

- No edit wars, etc.:
- It is illustrated by
images and other media, where possible and appropriate.
- a (images are tagged and non-free content have
fair use rationales):
 b (
appropriate use with
suitable captions):
b (
appropriate use with
suitable captions): 
- As before, the images are fantastic.
- a (images are tagged and non-free content have
fair use rationales):
- Overall:
- Pass/Fail:

- Thanks for all the work on this, and sorry again to keep you waiting so long for a review! Everything looks good to me though, so I'm closing this review as pass and promoting the article to GA. -- Cerebellum ( talk) 17:22, 25 July 2015 (UTC)
- Pass/Fail:
Thanks
Thank you to everyone who aided in getting this article up to GA level. It was definitely a job worth doing and hopefully sets a reasonable standard for the high-importance biology articles. T.Shafee(Evo﹠Evo) talk 00:02, 27 July 2015 (UTC)
More misc prose/additions
- in "Structure and Function" The term mRNA appears without any further explanation, for a non biologist reader (like me) this is irritating. — Preceding unsigned comment added by 141.42.200.71 ( talk) 07:01, 22 September 2015 (UTC)
- The Williams quote is widely cited elsewhere as p24 of the 1966 edition, but I don't have a copy of the book and it isn't in Google Books. Can anyone confirm?
I don't have a copy of Watson 2013 and can't preview that through Google Books either, so I'll leave going through that to those with a taste for lots of footnotes.Adding citations to the Alberts edition in NCBI is a good idea also, though in the absence of numbered chapters/sections the {{ rp}} solution to repetitive citations is less appealing. Opabinia regalis ( talk) 06:52, 18 April 2015 (UTC)Add paragraph on gene duplication, de novo genes, pseudogenes (after sequence homology?)Do we need something about genetic disorders? Currently there is the inheritance image, but that only refers to a 'trait', not a disorder, and there's nothing specifically about molecular mechanism.- Companion article genetics is highly redundant with this one.
I got distracted for a bit but have been meaning to get back to this. I think the above is the rest of my intended new content list. Any other suggestions? Opabinia regalis ( talk) 07:03, 5 May 2015 (UTC)
- Ok, the article has come a long way since we started.
I agree that we need something on genetic disorders.Possibly also a sentence or two on gene therapy in the engineering section since it's something many people will have heard about?Half of the sections have a {{ main}} template - do we really need them?
- I don't think there are any other major rearrangements of the sections left. It looks like mostly clarifying language, some small additions and adding the final references. T.Shafee(Evo﹠Evo) talk 12:57, 15 May 2015 (UTC)
- I did see this; I just haven't had time to actually write anything. Googling for refs is a lot easier :) Opabinia regalis ( talk) 07:03, 20 May 2015 (UTC)
Is gene a locus or a segment, or something more generic?
This topic contains two definitions at first paragraph and as figure legend:
A gene is a locus (or region) of DNA that encodes a functional RNA or protein product, and is the molecular unit of heredity.
A gene is a segment of DNA that encodes function. A chromosome consists of a long strand of DNA containing many genes. A human chromosome can have up to 500 million base pairs of DNA with thousands of genes.
Do they define the same thing? If so, why two different expressions? Well, those represent as if a locus is a segment. Is that so?
A gene can be just a segment with a fixed position and length on genomic DNA in very simple cases. However, the nature is often more complex.
If a gene is really a segment of DNA, is a multi-exonic gene a segment? Then what distinguishes multi-cistronic genes from exons? What about nested genes? What about overlapped genes? What about a trans-spliced gene which often have discontinued fragments on different chromosomes? They are not rare cases.
I think the stated definition is just a WORKING HYPOTHESIS with too much simplification.
Finally, this is my opinion: When Mendel proposed the concept of gene (though he did not use that term), it was a concept of atomic inheritable unit. In genetics, including molecular, it is still so, except modification for quantitative trait loci.
The definition on Wikipedia is biased too much toward molecular biology, and unexplainable phenomena are left by the definition.
Wordmasterexpress ( talk) 09:00, 8 April 2016 (UTC)
- @ Wordmasterexpress: Interesting points. Both "region" and "segment" are used as non-technical synonyms for locus. Although both are vague, I agree that "region" is slightly more appropriate since it is broad enough to encompass the various oddities like nested genes etc. I've therefore updated the lead image's caption. As for the definition overall: There has been a strong molecular biology focus on the definition of a gene since the modern evolutionary synthesis, so it has shifted away from the original Mendelian definition. We've preferenced the MBOC definition as representing the mainstream scientific consensus definition. As you say, there is always going to be difficulty in precisely defining a gene, since nature is complicated. A simple and broad definition is need in the lead, so we attempt to cover more complicated scenarios in the Gene#Functional_definitions section. T.Shafee(Evo﹠Evo) talk 05:36, 9 April 2016 (UTC)
- @ Evolution and evolvability: Thank you for edits. Now I have more questions. The lead seems to say a gene is a locus and a locus is equivalent to a region. Is that so? If yes, genes are a subset of loci. I doubt the second point that a locus is equivalent to a region, but it may depending on the definition. Either positive or negative, the lead is better clarifying those, I would suggest. -- Wordmasterexpress ( talk) 01:43, 11 April 2016 (UTC)
Proposed merge with Genomic organization
I propose that Genomic organization be merged into Gene. I think that the content in the Genomic organization can easily be explained in the context of Gene, and the resulting article will be of a reasonable size. Any input is welcome :) GiggsIsLegend ( talk) 01:47, 23 June 2016 (UTC)
- Comment - Genome could be a more appropriate merge destination. T.Shafee(Evo﹠Evo) talk 04:01, 23 June 2016 (UTC)
- Do not support - both topics are notable and distinct. A reader would want information on genes while another reader would want information on the organization of genomic organization. Best Regards, Barbara (WVS) ( talk) 10:52, 30 June 2016 (UTC)
- Do not support - to clarify, I don't think Genomic organisation should be merged into Gene. It could conceivably be merged into Genome though. T.Shafee(Evo﹠Evo) talk 12:24, 30 June 2016 (UTC)
- do not support(oppose) per T.Shafee(Evo﹠Evo)-- Ozzie10aaaa ( talk) 12:44, 30 June 2016 (UTC)
- Oppose As others have said above, it is a distinct topic from Gene. If it was to be merged anywhere it should be into Genome but I think it a notable enough topic to warrant its own article. It needs a bit of work but there is good potential for it to be expanded and improved. Sarahj2107 ( talk) 13:13, 30 June 2016 (UTC)
- Oppose I agree with all the arguments put forth. Genomic organisation is an up an coming research area - it would be great if it attracted some dedicated editors. McortNGHH ( talk) 15:29, 30 June 2016 (UTC)
- Oppose. Genomic organisation is a distinct topic, and warrants its own article. However its current lede starts poorly, and may give casual readers the impression that it covers the same material as Gene. Maproom ( talk) 08:13, 3 July 2016 (UTC)
Not only proteins
The first sentence says that a gene codes for a protein. The last paragraph says that this is under discussion, that a gene also can code for functional non-coding RNAs. But that discussion was concluded long ago. I checked several of my rather oldish biology books. For example, the glossary in Hartwell, Hood, Goldberg, Reynolds, Silver, Veres; Genetics, from Genes to Genomes; 2004: ... segment of DNA in a discrete region of a chromosome that serves as a unit of function by encoding a particular RNA or Protein. -- Ettrig ( talk) 09:18, 28 October 2017 (UTC)
- Different versions of the article have mentioned RNA genes in the lead e.g.
- "A gene is a locus (or region) of DNA that encodes a functional RNA or protein product" Special:Permalink/702868373
- "The word is used extensively by the scientific community for stretches of deoxyribonucleic acid (DNA) and ribonucleic acid (RNA) that code for a polypeptide or for an RNA chain that has a function in the organism." Special:Permalink/666720556
- There is the later section on RNA genes too. After browsing through the history, it seems that mentions of RNA genes in the lead are frequently removed to "improve clarity". Personally I think these not mentioning them introduces inaccuracy. It may be best to involve other editors of the article in this discussion to prevent any rapid reversions e.g. @ Evolution and evolvability: & @ Headbomb: who mention RNAs in their edits. -- Paul ( talk) 08:59, 29 October 2017 (UTC)
- @ Ettrig: That's a great improvement, I'm not sure about this sentence "A gene is a subsequence of DNA which codes for a molecule that has a function". What about RNA viruses? Maybe add a "usually" in there somewhere. -- Paul ( talk) 21:52, 29 October 2017 (UTC)
- Agree, RNA should be included. This is an omission of the same kind as the one I fixed. I will add or RNA. This is a bit crude. But my feeling is that nucleotides is less well known. -- Ettrig ( talk) 07:15, 30 October 2017 (UTC)
- Added RNA, with some difficulty. The current text does not cover the retrovirus case. I now empathize with the clarity argument. We want the first sentences to be correct and also very succinct and accessible. -- Ettrig ( talk) 07:22, 30 October 2017 (UTC)
- Hello all. Sorry to be late to the thread! Great work so far. You're correct that it was high time for the lead to be brought into line with the rest of the article. My main outstanding issue is the term "subsequence" in the first sentence. I think that "sequence" would suffice and I don't think extra useful specificity is added to the definition by using "subsequence". Indeed it's the only time the term is used in the article.
T.Shafee(Evo&Evo)
talk
00:36, 31 October 2017 (UTC)
- Yes, indeed. And thanks to @ Chiswick Chap:. -- Ettrig ( talk) 12:43, 31 October 2017 (UTC)
- Hello all. Sorry to be late to the thread! Great work so far. You're correct that it was high time for the lead to be brought into line with the rest of the article. My main outstanding issue is the term "subsequence" in the first sentence. I think that "sequence" would suffice and I don't think extra useful specificity is added to the definition by using "subsequence". Indeed it's the only time the term is used in the article.
T.Shafee(Evo&Evo)
talk
00:36, 31 October 2017 (UTC)
Definitions
The definition of gene is probably one of the most important parts of this article. At the risk of opening Pandora's box on, it might be worth checking that we're using the best available definitions. Currently we list three definitions:
-
Gene#Lead - A gene is the molecular unit of
heredity of a living
organism.
[1]
- Clear and simple, but lacks any mention of function.
-
Gene#Lead - Therefore, a modern working definition is "a
locatable region of
genomic sequence, corresponding to a unit of inheritance, which is associated with regulatory regions, transcribed regions, and/or other functional sequence regions".
[2]
[3]
- The only definition that we are directly quoting from a reference and seems verbose to me.
- The use of "associated with " feels unnecessarily confusing.
- Also, close to tautology: "genomic sequence... which is associated with... other functional sequence regions".
-
Gene#Functional definitions - A broad operational definition is sometimes used to encompass the complexity of these diverse phenomena, where a gene is defined as a union of genomic sequences encoding a coherent set of potentially overlapping functional products.
[4]
- It think this is fine but omits heredity. Does it need to? What phenomena fit this definition but are not heritable?
- Exclusion of regulatory sequence doesn't seem hugely necessary.
Overall I think that we do need a simple layman's definition as well as a more nuanced, broader, technical definition. Both probably need to encompass:
- Molecular / sequence / locus / encoding
- Heritable
- Functional / expressed
T.Shafee(Evo﹠Evo) talk 01:10, 14 June 2015 (UTC)
[1] perhaps?..-- Ozzie10aaaa ( talk) 01:22, 14 June 2015 (UTC)
- You'd think that developing a really good single-sentence definition would be really important, but... I just can't get myself too worked up about it. It's a fuzzy concept; people use slightly different definitions for different purposes; the question of 'what do we annotate as a gene' is subtly but significantly different; etc. etc. I looked around a little specifically in the education literature and found this hilarious "simplified" model: DOI 10.1007/s11191-008-9161-7. (This thesis studying representations of genes in textbooks is pretty interesting: [2] Not so useful for our purposes, though.) Opabinia regalis ( talk) 05:38, 16 June 2015 (UTC)
- You're probably right. I might still have a crack at polishing the currently existing sentences over the next few days, but we do a pretty decent job of describing the key elements in the functional structure section. Also, one of the papers that comprises the thesis you mentioned was actually a pretty good history of the definition. T.Shafee(Evo﹠Evo) talk 12:08, 19 June 2015 (UTC)
I've now updated the definitions in the article:
- a region of DNA that controls a discrete, hereditary trait in an organism
- based on the the definition from the MBOC glossary since I think this more clearly includes the key elements [5]: Glossary
- any discrete region of heritable, genomic sequence which affect an organism's traits by being expressed as a functional product or by regulating expression
- still based on the same references, but reworded to simplify the language and sentence structure whilst still being thorough.
- a union of genomic sequences encoding a coherent set of potentially overlapping functional products
- left alone with only a minor change to the sentence before it.
I think that these should suffice as a simple summary for a brief visitor, as well as a more detailed definition for anyone more interested. T.Shafee(Evo﹠Evo) talk 12:16, 25 June 2015 (UTC)
- so, is gene a material, address, or information? If material, it should be DNA (or RNA for minor cases), which is very clear, but remains some unexplainable attribute. If address, even more unexplainability, which seems current definition. If information, I find no problem when I consider genetic materials are just media of genetic information, but it's just my impression. What is the formal, gold standard consensus of current definition? Current lede seems tautological, or pretty much redundant at least, and does not seem to provide clear idea.
- A gene is a locus (or region) of blah blah blha defines a gene is an address.
- Genes can acquire mutations in their sequence, leading to different variants, known as alleles defines a gene contains sequence a part of which is mutable in their address. It also defines the mutated variant is called allele. The question is that, if an allele is also defined as sequence, can it be a gene when sequence is contained in the address (gene). Also if a gene is defined as sequence, can it point an address when many experiments had suggested that genes can function regardless of, even out of, the chromosomal location.
- Therefore, a broad, modern working definition of a gene is any discrete locus of heritable, genomic sequence. might represent best, but it still states that a gene is a locus. Uh, could be tautology. Well, polygenes?
-- Wordmasterexpress ( talk) 09:55, 5 October 2016 (UTC)
- Dear @ Wordmasterexpress, Evolution and evolvability, Ozzie10aaaa, and Opabinia regalis: I appreciate your efforts to come up with a definition. When writing an intro paragraph, I think it is allowed to use a heuristic, where short and efficient may take the overhand over "complete". For the latter, one may expect the user to read the article. The first paragraph is what people see on other pages where the word is used and linked so that when readers hoover over the word, they get a pop-up of the first paragraph to remind them what the word is about or give a heuristic that makes them understand what it is about. Right? It's quite popular to refer to things as "the smallest unit of" e.g. a genetic codification of a protein that will eventually result in a trait. Isn't that good enough for most popular readings or when you come across the word in a wikipedia article? Look e.g. at how " engram" - the unit of cognitive information inside the brain - is defined, or atom, or molecule. Sincerely, SvenAERTS ( talk) 03:45, 24 January 2020 (UTC)
refs
References
- ^ Slack, J.M.W. Genes-A Very Short Introduction. Oxford University Press 2014
- ^ Pearson H (May 2006). "Genetics: what is a gene?". Nature. 441 (7092): 398–401. Bibcode: 2006Natur.441..398P. doi: 10.1038/441398a. PMID 16724031.
- ^ Pennisi E (June 2007). "Genomics. DNA study forces rethink of what it means to be a gene". Science. 316 (5831): 1556–1557. doi: 10.1126/science.316.5831.1556. PMID 17569836.
- ^ Gerstein MB, Bruce C, Rozowsky JS, Zheng D, Du J, Korbel JO, Emanuelsson O, Zhang ZD, Weissman S, Snyder M (June 2007). "What is a gene, post-ENCODE? History and updated definition". Genome Research. 17 (6): 669–681. doi: 10.1101/gr.6339607. PMID 17567988.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). Molecular Biology of the Cell (Fourth ed.). New York: Garland Science. ISBN 978-0-8153-3218-3.
Requested Edit: Structure and Function / Structure chapter referencing own research
| This edit request by an editor with a conflict of interest has now been answered. |
I request a volunteer to review a proposed reference to own research. Details below.
At the end of this paragraph:
The transcribed pre-mRNA contains untranslated regions at both ends which contain a ribosome binding site, terminator, and start and stop codons.[42] In addition, most eukaryotic open reading frames contain untranslated introns, which are removed and exons, which are connected together in a process known as RNA splicing. Finally, the ends of gene transcripts are defined by CPA sites, where newly produced pre-mRNA gets cleaved and a string of ~200 adenosine monophosphates is added at 3′ end. PolyA tail protects mature mRNA from degradation and has other functions, affecting translation, localization, and transport of the transcript from the nucleus. Splicing, followed by CPA generate the final mature mRNA which encodes the protein or RNA product.[43] Although the general mechanisms defining locations of human genes are known, identification of the exact factors regulating these cellular processes is an area of active research. For example, known sequence features in 3′-UTR can only explain half of all human gene ends. [CITATION PROPOSED HERE]
I am requesting a reference to a recent paper from our lab - https://pubmed.ncbi.nlm.nih.gov/34104882/
Wikipedia guidelines suggest citing review papers first, however, this is the first study that assesses whether known sequence elements are sufficient to predict human gene ends. Strikingly, we found that known elements can only explain half of all human gene ends. This project was aimed at answering fundamental questions of human gene definition. The results are significant and highlight the magnitude of missing information in the current understanding of processes human cells utilize to locate their genes. Therefore, this study is the most relevant and most recent source on the subject.
To justify expert knowledge - this paper comes from Hughes lab at the University of Toronto. Dr. Timothy R Hughes is a John W. Billes Chair of Medical Research and Canada Research Chair in Decoding Gene Regulation, one of the most cited Canadian researchers.
 Done with minor edits (missing articles, punctuation).
Heartmusic678 (
talk)
15:06, 22 July 2021 (UTC)
Done with minor edits (missing articles, punctuation).
Heartmusic678 (
talk)
15:06, 22 July 2021 (UTC)
Gwne
Hjaj 2409:4052:4D1F:E11E:8CDF:AA35:8201:7E15 ( talk) 19:03, 20 March 2022 (UTC)
Difference between gene and chromosome
How does the gene differ from a chromosome? 41.116.8.55 ( talk) 16:08, 13 January 2022 (UTC)
- Genes control cell function and overlay the chromosome, which is the unit of transmission, all of which is a single molecule of DNA. Don't know why I answered that. Habit. 2600:8807:1C0A:9600:C53E:7E54:BE58:FCED ( talk) 12:08, 31 January 2023 (UTC)
Defining "gene"
- Continued from talk:Gene structure#Definition of a gene
Arguably, one of the most important purposes of this article is to come up with a good definition of "gene." I'm not happy with the current definition for many reasons so I'd like to start a discussion about how we can change it. Check out an old blog post of mine from 2007 where I define a gene as, "A gene is a DNA sequence that is transcribed to produce a functional product."
What Is a Gene? https://sandwalk.blogspot.com/2007/01/what-is-gene.html
This is the standard definition dating back (at least) to Watson's textbook "Molecular Biology of the Gene" (1965). The idea is that a gene is a transcription unit and there are two kinds of genes: protein-coding genes and noncoding genes. In addition, the product has to have a function - junk RNA transcripts do not define genes.
All of the best textbooks continue to use this standard definition and nothing has changed since the publication of the human genome sequence in spite of all the rhetoric that has been published. See my post on Gerald Fink for an example of the kind of misinformation that we have to correct in this article.
Gerald Fink promotes a new definition of a gene https://sandwalk.blogspot.com/2019/09/gerald-fink-promotes-new-definition-of.html
Genome42 ( talk) 17:07, 1 August 2022 (UTC)
- I use a carefully defined single sentence definition meant to encompass all cases for identified genes. " A gene is a chromosomal locus that governs the expression of a heritable trait or is homologous to a known gene." Genes have multiple aspects each with its own homologs, so in practice it still is complicated. My course also contains a longer molecular definition. "A gene is a nucleic acid, all or in part, that is composed of dispersed, modular units of molecular function that have an emergent property of immediate or secondary cellular activity. Gene function is not an inherent property of the nucleic acid sequence but depends on the cellular context and activity. The gene is the total of all the nucleic acid sequences that significantly influence the achievement of that cellular activity and share a dependency on the function of at least one single modular unit. While a certain number of correct modular units of function are required, there appears no limit to the number of such units or their dispersal over a single chromosome." The word choice is generic and the definition requires descriptions or example of molecular function (start, stop; +1, terminator;5', 3' splice sites; etc.. Dr. Rogers 2600:8807:1C0A:9600:C53E:7E54:BE58:FCED ( talk) 12:53, 31 January 2023 (UTC)
Genome, Molecular Evolution, Inheritance, Gene Expression
Do we really need extensive discussion of these topics here since there are separate articles for all of them? The problem is that when the main articles are updated and corrected the information here then conflicts with the main article.
For example, under "Molecular evolution: Mutation" we find the following statement "This means that each generation, each human genome accumulates 1–2 new mutations." This is wrong in two possible ways. If it refers to a cell generation then you can do a simple calculation based on the preceding statement that the error rate is "as low as 10−8 per nucleotide per replication." Since there are 6.1 x 109 nucleotides being synthesized that means at least 62 mutations or 31 in each daughter cell. (The actual mutation rate due to DNA replication errors is 10-10 per base pair or less than one per replication.)
If it refers to human generations then it's also wrong. Each newborn baby has about 100 new spontaneous mutations. (The Wikipedia mutation rate article says 64 new mutations per generation but that needs to be updated.)
This is just one example of the problems that arise when there are too many editors duplicating information in these articles. We need to concentrate our efforts on a few key articles that we link to whenever the topic comes up somewhere else. That way we don't risk spreading and perpetuating misinformation because the updates and corrections aren't propagated to the other articles.
I'm proposing a drastic change because it means deleting, or substantially reducing, a lot of stuff in this article.
What do you think? Genome42 ( talk) 17:37, 7 March 2023 (UTC)
- Hi Larry, I just googled for the WP policy on redundancy (of course there would be a policy). You can see it here
Wikipedia:Abundance and redundancy, I agree it's a pain having conflicting information between articles &/or maintaining redundant information. But it looks like you might be better off correcting the more obvious flaws you see here, but taking a lighter touch than wholesale removal/reduction of topics. --
Paul (
talk)
20:27, 8 March 2023 (UTC)
- Paul, I’m can’t imagine why we need an extensive discussion of molecular evolution (including mutation) in an article on “gene.” Are you saying that we should keep it just because deleting might cause trouble with Wikipedians or is it because you think it’s important to discuss molecular evolution here?
Genome42 (
talk)
13:23, 9 March 2023 (UTC)
- Larry, no that is very much not what I'm saying. If the text is clearly tangential to the main topic, then by all means move it somewhere else, or remove/reduce. I thought I was responding to your query about redundant information between general and specific articles. A case could be made to split some of this article out into other topics. -- Paul ( talk) 08:08, 10 March 2023 (UTC)
- Paul, I’m can’t imagine why we need an extensive discussion of molecular evolution (including mutation) in an article on “gene.” Are you saying that we should keep it just because deleting might cause trouble with Wikipedians or is it because you think it’s important to discuss molecular evolution here?
Genome42 (
talk)
13:23, 9 March 2023 (UTC)
- Both pages even cite the
same 1998 source for those different human mutation rate numbers! Definitely needs an in-depth read to work out which is more accurate (or if there have been new alternative calculations done since we've so many more sequenced genomes now than back then!). Some level of overlap on broad topics like this is inevitable, since almost all of the material is also covered in more detail in other narrower-scope pages, so we'll always have some amount of maintenance to do no matter how long the page. However it's currently ~8000 words long so certainly be pruned down to 6000 with judicious removal of material that is a) less core to an expected average reader and b) covered elsewhere easily navigated to.
T.Shafee(Evo&Evo)
talk
02:53, 9 March 2023 (UTC)
- @ Evolution and evolvability I’m quite knowledgeable on this topic so I don’t really need to do any more research. The main issue is finding the right citations and figuring out how to deal with editors who might want to keep the incorrect information just because it is supported by a “reliable source” (sensu Wikipedia). It would be a lot easier to fix the mutation article and then link to it from here if we think it’s really necessary to discuss mutation in this article. What do you think? Should I risk being accused of edit wars on both articles? Genome42 ( talk) 13:32, 9 March 2023 (UTC)