| Haplogroup S-M230 (S1a1b) | |
|---|---|
 | |
| Possible time of origin | 28,000-41,000 years before present ( Scheinfeldt 2006) |
| Possible place of origin | New Guinea/ Indonesia |
| Ancestor | S1a1 (Z42413) [1] [2] |
| Descendants | S1a1b1 (M254) |
| Defining mutations | M230, P202 |
| Highest frequencies | Ekari 74% ( Mona 2007) |
Haplogroup S-M230, also known as S1a1b (and previously as S* or K2b1a4), is a Y-chromosome DNA haplogroup. It is by far the most numerically significant subclade of Haplogroup S1a (and its sole primary subclade, Haplogroup S-P405).
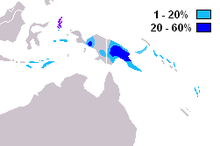
S-M230 is commonly found among populations of the highlands of Papua New Guinea ( Kayser 2003). It is also found at lower frequencies in adjacent parts of Indonesia and Melanesia. [1] ( Kayser 2003 Cox 2006)
Origin
|
| This section is empty. You can help by
adding to it. (January 2013) |
Distribution
One study has reported finding haplogroup S-M230 in: 52% (16/31) of a sample from the Papua New Guinea (PNG) Highlands; 21% (7/34) of a sample from the Moluccas; 16% (5/31) of a sample from the Papua New Guinea coast; 12.5% (2/16) of a sample of Tolai from New Britain; 10% (3/31) of a sample from Nusa Tenggara, and; 2% (2/89) of a sample from the West New Guinea lowlands/coast.( Kayser 2003 Cox 2006)
One subclade, Haplogroup S-M226.1 (S1a1b1d1a; previously S1d) has been found at low frequencies in the Admiralty Islands and along the coast of mainland PNG. ( Kayser 2008).
Phylogenetics
Structure & position within Haplogroup S
- S1a
- S1a1 Z42413
- S1a1a
- S1a1a1 P60, P304, P308
- S1a1a2
- S1a1b M230, P202, P204 – "demoted" in 2016 from its previous position as the basal Haplogroup S* (and known before that as Haplogroup K5)
- S1a1b1 (M254) (previously known as K2b1a4a)
- S1a1b1 (M254)
- S1a1b1a (P57)
- S1a1b1b (P61)
- S1a1b1c (P83)
- S1a1b1d (SK1891)
- S1a1a
- S1a2 P79, P307
-
S1a3 P315
- S1a3a Z41763
- S1a3b~ P401
- (Based on the 2017 ISOGG tree, [1] the 2015 ISOGG tree, [3]) the 2008 YCC tree ( Karafet 2008) and other published research.)
History
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Later, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| K-M9 | 26 | VIII | 1U | 25 | Eu16 | H5 | F | K* | K | K | K | - | - | - | - | - | - | - |
Research publications
The following research teams per their publications were represented in the creation of the YCC tree.
2005 YCC tree
|
| This section is empty. You can help by
adding to it. (January 2013) |
2008 YCC tree
From 2002 to 2008, it was known as Haplogroup K5.[ citation needed]
See also
Genetics
Y-DNA S subclades
- S-M230
Y-DNA backbone tree
References
- ^ a b c International Society of Genetic Genealogy (ISOGG; 2017), Y-DNA Haplogroup Tree 2017 (24 March 2017).
- ^ "Browse Articles | European Journal of Human Genetics". www.nature.com. Retrieved 25 March 2023.
- ^ International Society of Genetic Genealogy (ISOGG; 2015), Y-DNA Haplogroup Tree 2015. (Access date: 1 February 2015.)
Footnotes
Works cited
- Cox, Murray P.; Mirazón Lahr, Marta (2006). "Y-chromosome diversity is inversely associated with language affiliation in paired Austronesian- and Papuan-speaking communities from Solomon Islands". American Journal of Human Biology. 18 (1): 35–50. doi: 10.1002/ajhb.20459. PMID 16378340. S2CID 4824401.
- Karafet, T. M.; Mendez, F. L.; Meilerman, M. B.; Underhill, P. A.; Zegura, S. L.; Hammer, M. F. (2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research. 18 (5): 830–8. doi: 10.1101/gr.7172008. PMC 2336805. PMID 18385274.
- Kayser, Manfred; Brauer, Silke; Weiss, Gunter; Schiefenhövel, Wulf; Underhill, Peter; Shen, Peidong; Oefner, Peter; Tommaseo-Ponzetta, Mila; Stoneking, Mark (2003). "Reduced Y-Chromosome, but Not Mitochondrial DNA, Diversity in Human Populations from West New Guinea". The American Journal of Human Genetics. 72 (2): 281–302. doi: 10.1086/346065. PMC 379223. PMID 12532283.
- Kayser, M.; Choi, Y.; Van Oven, M.; Mona, S.; Brauer, S.; Trent, R. J.; Suarkia, D.; Schiefenhovel, W.; Stoneking, M. (2008). "The Impact of the Austronesian Expansion: Evidence from mtDNA and Y Chromosome Diversity in the Admiralty Islands of Melanesia". Molecular Biology and Evolution. 25 (7): 1362–74. doi: 10.1093/molbev/msn078. PMID 18390477.
- Mona, S.; Tommaseo-Ponzetta, M.; Brauer, S.; Sudoyo, H.; Marzuki, S.; Kayser, M. (2007). "Patterns of Y-chromosome diversity intersect with the Trans-New Guinea hypothesis". Mol. Biol. Evol. 24 (11): 2546–55. doi: 10.1093/molbev/msm187. PMID 17846104.
- Scheinfeldt, L.; Friedlaender, F; Friedlaender, J; Latham, K; Koki, G; Karafet, T; Hammer, M; Lorenz, J (2006). "Unexpected NRY Chromosome Variation in Northern Island Melanesia". Molecular Biology and Evolution. 23 (8): 1628–41. doi: 10.1093/molbev/msl028. PMID 16754639.
Sources for conversion tables
- Capelli, Cristian; Wilson, James F.; Richards, Martin; Stumpf, Michael P.H.; et al. (February 2001). "A Predominantly Indigenous Paternal Heritage for the Austronesian-Speaking Peoples of Insular Southeast Asia and Oceania". The American Journal of Human Genetics. 68 (2): 432–443. doi: 10.1086/318205. PMC 1235276. PMID 11170891.
- Hammer, Michael F.; Karafet, Tatiana M.; Redd, Alan J.; Jarjanazi, Hamdi; et al. (1 July 2001). "Hierarchical Patterns of Global Human Y-Chromosome Diversity". Molecular Biology and Evolution. 18 (7): 1189–1203. doi: 10.1093/oxfordjournals.molbev.a003906. PMID 11420360.
- Jobling, Mark A.; Tyler-Smith, Chris (2000), "New uses for new haplotypes", Trends in Genetics, 16 (8): 356–62, doi: 10.1016/S0168-9525(00)02057-6, PMID 10904265
- Kaladjieva, Luba; Calafell, Francesc; Jobling, Mark A; Angelicheva, Dora; et al. (February 2001). "Patterns of inter- and intra-group genetic diversity in the Vlax Roma as revealed by Y chromosome and mitochondrial DNA lineages". European Journal of Human Genetics. 9 (2): 97–104. doi: 10.1038/sj.ejhg.5200597. PMID 11313742. S2CID 21432405.
- Karafet, Tatiana; Xu, Liping; Du, Ruofu; Wang, William; et al. (September 2001). "Paternal Population History of East Asia: Sources, Patterns, and Microevolutionary Processes". The American Journal of Human Genetics. 69 (3): 615–628. doi: 10.1086/323299. PMC 1235490. PMID 11481588.
- Semino, O.; Passarino, G; Oefner, PJ; Lin, AA; et al. (2000), "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective", Science, 290 (5494): 1155–9, Bibcode: 2000Sci...290.1155S, doi: 10.1126/science.290.5494.1155, PMID 11073453
- Su, Bing; Xiao, Junhua; Underhill, Peter; Deka, Ranjan; et al. (December 1999). "Y-Chromosome Evidence for a Northward Migration of Modern Humans into Eastern Asia during the Last Ice Age". The American Journal of Human Genetics. 65 (6): 1718–1724. doi: 10.1086/302680. PMC 1288383. PMID 10577926.
- Underhill, Peter A.; Shen, Peidong; Lin, Alice A.; Jin, Li; et al. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics. 26 (3): 358–361. doi: 10.1038/81685. PMID 11062480. S2CID 12893406.
Further reading
- Haber, Marc; Platt, Daniel E.; Ashrafian Bonab, Maziar; Youhanna, Sonia C.; Soria-Hernanz, David F.; Martínez-Cruz, Begoña; Douaihy, Bouchra; Ghassibe-Sabbagh, Michella; et al. (2012). Kayser, Manfred (ed.). "Afghanistan's Ethnic Groups Share a Y-Chromosomal Heritage Structured by Historical Events". PLOS ONE. 7 (3): e34288. Bibcode: 2012PLoSO...734288H. doi: 10.1371/journal.pone.0034288. PMC 3314501. PMID 22470552.
- Karafet, Tatiana M.; Lansing, J. S.; Redd, Alan J.; Watkins, Joseph C.; Surata, S. P. K.; Arthawiguna, W. A.; Mayer, Laura; Bamshad, Michael; et al. (2005). "Balinese Y-Chromosome Perspective on the Peopling of Indonesia: Genetic Contributions from Pre-Neolithic Hunter-Gatherers, Austronesian Farmers, and Indian Traders". Human Biology. 77 (1): 93–114. doi: 10.1353/hub.2005.0030. hdl: 1808/13586. PMID 16114819. S2CID 7953854.
- Kayser, M.; Brauer, S; Cordaux, R; Casto, A; Lao, O; Zhivotovsky, LA; Moyse-Faurie, C; Rutledge, RB; et al. (2006). "Melanesian and Asian Origins of Polynesians: MtDNA and Y Chromosome Gradients Across the Pacific". Molecular Biology and Evolution. 23 (11): 2234–44. doi: 10.1093/molbev/msl093. hdl: 11858/00-001M-0000-0010-0145-0. PMID 16923821.
Phylogenetics
| Haplogroup S-M230 (S1a1b) | |
|---|---|
 | |
| Possible time of origin | 28,000-41,000 years before present ( Scheinfeldt 2006) |
| Possible place of origin | New Guinea/ Indonesia |
| Ancestor | S1a1 (Z42413) [1] [2] |
| Descendants | S1a1b1 (M254) |
| Defining mutations | M230, P202 |
| Highest frequencies | Ekari 74% ( Mona 2007) |
Haplogroup S-M230, also known as S1a1b (and previously as S* or K2b1a4), is a Y-chromosome DNA haplogroup. It is by far the most numerically significant subclade of Haplogroup S1a (and its sole primary subclade, Haplogroup S-P405).
S-M230 is commonly found among populations of the highlands of Papua New Guinea ( Kayser 2003). It is also found at lower frequencies in adjacent parts of Indonesia and Melanesia. [1] ( Kayser 2003 Cox 2006)
Origin
|
| This section is empty. You can help by
adding to it. (January 2013) |
Distribution
One study has reported finding haplogroup S-M230 in: 52% (16/31) of a sample from the Papua New Guinea (PNG) Highlands; 21% (7/34) of a sample from the Moluccas; 16% (5/31) of a sample from the Papua New Guinea coast; 12.5% (2/16) of a sample of Tolai from New Britain; 10% (3/31) of a sample from Nusa Tenggara, and; 2% (2/89) of a sample from the West New Guinea lowlands/coast.( Kayser 2003 Cox 2006)
One subclade, Haplogroup S-M226.1 (S1a1b1d1a; previously S1d) has been found at low frequencies in the Admiralty Islands and along the coast of mainland PNG. ( Kayser 2008).
Phylogenetics
Structure & position within Haplogroup S
- S1a
- S1a1 Z42413
- S1a1a
- S1a1a1 P60, P304, P308
- S1a1a2
- S1a1b M230, P202, P204 – "demoted" in 2016 from its previous position as the basal Haplogroup S* (and known before that as Haplogroup K5)
- S1a1b1 (M254) (previously known as K2b1a4a)
- S1a1b1 (M254)
- S1a1b1a (P57)
- S1a1b1b (P61)
- S1a1b1c (P83)
- S1a1b1d (SK1891)
- S1a1a
- S1a2 P79, P307
-
S1a3 P315
- S1a3a Z41763
- S1a3b~ P401
- (Based on the 2017 ISOGG tree, [1] the 2015 ISOGG tree, [3]) the 2008 YCC tree ( Karafet 2008) and other published research.)
History
Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome Phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Later, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| K-M9 | 26 | VIII | 1U | 25 | Eu16 | H5 | F | K* | K | K | K | - | - | - | - | - | - | - |
Research publications
The following research teams per their publications were represented in the creation of the YCC tree.
2005 YCC tree
|
| This section is empty. You can help by
adding to it. (January 2013) |
2008 YCC tree
From 2002 to 2008, it was known as Haplogroup K5.[ citation needed]
See also
Genetics
Y-DNA S subclades
- S-M230
Y-DNA backbone tree
References
- ^ a b c International Society of Genetic Genealogy (ISOGG; 2017), Y-DNA Haplogroup Tree 2017 (24 March 2017).
- ^ "Browse Articles | European Journal of Human Genetics". www.nature.com. Retrieved 25 March 2023.
- ^ International Society of Genetic Genealogy (ISOGG; 2015), Y-DNA Haplogroup Tree 2015. (Access date: 1 February 2015.)
Footnotes
Works cited
- Cox, Murray P.; Mirazón Lahr, Marta (2006). "Y-chromosome diversity is inversely associated with language affiliation in paired Austronesian- and Papuan-speaking communities from Solomon Islands". American Journal of Human Biology. 18 (1): 35–50. doi: 10.1002/ajhb.20459. PMID 16378340. S2CID 4824401.
- Karafet, T. M.; Mendez, F. L.; Meilerman, M. B.; Underhill, P. A.; Zegura, S. L.; Hammer, M. F. (2008). "New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree". Genome Research. 18 (5): 830–8. doi: 10.1101/gr.7172008. PMC 2336805. PMID 18385274.
- Kayser, Manfred; Brauer, Silke; Weiss, Gunter; Schiefenhövel, Wulf; Underhill, Peter; Shen, Peidong; Oefner, Peter; Tommaseo-Ponzetta, Mila; Stoneking, Mark (2003). "Reduced Y-Chromosome, but Not Mitochondrial DNA, Diversity in Human Populations from West New Guinea". The American Journal of Human Genetics. 72 (2): 281–302. doi: 10.1086/346065. PMC 379223. PMID 12532283.
- Kayser, M.; Choi, Y.; Van Oven, M.; Mona, S.; Brauer, S.; Trent, R. J.; Suarkia, D.; Schiefenhovel, W.; Stoneking, M. (2008). "The Impact of the Austronesian Expansion: Evidence from mtDNA and Y Chromosome Diversity in the Admiralty Islands of Melanesia". Molecular Biology and Evolution. 25 (7): 1362–74. doi: 10.1093/molbev/msn078. PMID 18390477.
- Mona, S.; Tommaseo-Ponzetta, M.; Brauer, S.; Sudoyo, H.; Marzuki, S.; Kayser, M. (2007). "Patterns of Y-chromosome diversity intersect with the Trans-New Guinea hypothesis". Mol. Biol. Evol. 24 (11): 2546–55. doi: 10.1093/molbev/msm187. PMID 17846104.
- Scheinfeldt, L.; Friedlaender, F; Friedlaender, J; Latham, K; Koki, G; Karafet, T; Hammer, M; Lorenz, J (2006). "Unexpected NRY Chromosome Variation in Northern Island Melanesia". Molecular Biology and Evolution. 23 (8): 1628–41. doi: 10.1093/molbev/msl028. PMID 16754639.
Sources for conversion tables
- Capelli, Cristian; Wilson, James F.; Richards, Martin; Stumpf, Michael P.H.; et al. (February 2001). "A Predominantly Indigenous Paternal Heritage for the Austronesian-Speaking Peoples of Insular Southeast Asia and Oceania". The American Journal of Human Genetics. 68 (2): 432–443. doi: 10.1086/318205. PMC 1235276. PMID 11170891.
- Hammer, Michael F.; Karafet, Tatiana M.; Redd, Alan J.; Jarjanazi, Hamdi; et al. (1 July 2001). "Hierarchical Patterns of Global Human Y-Chromosome Diversity". Molecular Biology and Evolution. 18 (7): 1189–1203. doi: 10.1093/oxfordjournals.molbev.a003906. PMID 11420360.
- Jobling, Mark A.; Tyler-Smith, Chris (2000), "New uses for new haplotypes", Trends in Genetics, 16 (8): 356–62, doi: 10.1016/S0168-9525(00)02057-6, PMID 10904265
- Kaladjieva, Luba; Calafell, Francesc; Jobling, Mark A; Angelicheva, Dora; et al. (February 2001). "Patterns of inter- and intra-group genetic diversity in the Vlax Roma as revealed by Y chromosome and mitochondrial DNA lineages". European Journal of Human Genetics. 9 (2): 97–104. doi: 10.1038/sj.ejhg.5200597. PMID 11313742. S2CID 21432405.
- Karafet, Tatiana; Xu, Liping; Du, Ruofu; Wang, William; et al. (September 2001). "Paternal Population History of East Asia: Sources, Patterns, and Microevolutionary Processes". The American Journal of Human Genetics. 69 (3): 615–628. doi: 10.1086/323299. PMC 1235490. PMID 11481588.
- Semino, O.; Passarino, G; Oefner, PJ; Lin, AA; et al. (2000), "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective", Science, 290 (5494): 1155–9, Bibcode: 2000Sci...290.1155S, doi: 10.1126/science.290.5494.1155, PMID 11073453
- Su, Bing; Xiao, Junhua; Underhill, Peter; Deka, Ranjan; et al. (December 1999). "Y-Chromosome Evidence for a Northward Migration of Modern Humans into Eastern Asia during the Last Ice Age". The American Journal of Human Genetics. 65 (6): 1718–1724. doi: 10.1086/302680. PMC 1288383. PMID 10577926.
- Underhill, Peter A.; Shen, Peidong; Lin, Alice A.; Jin, Li; et al. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics. 26 (3): 358–361. doi: 10.1038/81685. PMID 11062480. S2CID 12893406.
Further reading
- Haber, Marc; Platt, Daniel E.; Ashrafian Bonab, Maziar; Youhanna, Sonia C.; Soria-Hernanz, David F.; Martínez-Cruz, Begoña; Douaihy, Bouchra; Ghassibe-Sabbagh, Michella; et al. (2012). Kayser, Manfred (ed.). "Afghanistan's Ethnic Groups Share a Y-Chromosomal Heritage Structured by Historical Events". PLOS ONE. 7 (3): e34288. Bibcode: 2012PLoSO...734288H. doi: 10.1371/journal.pone.0034288. PMC 3314501. PMID 22470552.
- Karafet, Tatiana M.; Lansing, J. S.; Redd, Alan J.; Watkins, Joseph C.; Surata, S. P. K.; Arthawiguna, W. A.; Mayer, Laura; Bamshad, Michael; et al. (2005). "Balinese Y-Chromosome Perspective on the Peopling of Indonesia: Genetic Contributions from Pre-Neolithic Hunter-Gatherers, Austronesian Farmers, and Indian Traders". Human Biology. 77 (1): 93–114. doi: 10.1353/hub.2005.0030. hdl: 1808/13586. PMID 16114819. S2CID 7953854.
- Kayser, M.; Brauer, S; Cordaux, R; Casto, A; Lao, O; Zhivotovsky, LA; Moyse-Faurie, C; Rutledge, RB; et al. (2006). "Melanesian and Asian Origins of Polynesians: MtDNA and Y Chromosome Gradients Across the Pacific". Molecular Biology and Evolution. 23 (11): 2234–44. doi: 10.1093/molbev/msl093. hdl: 11858/00-001M-0000-0010-0145-0. PMID 16923821.