| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | Archive 2 | Archive 3 | Archive 4 |
Mutabox: a bot-generated box in mammalian gene pages, referring to targeted mutant alleles in mice.
Dear MCB group,
I am involved in the informatics of two high-throughput gene-knockout projects conducted in part by Team87 at the Wellcome Trust Sanger Institute. These are the eucomm project ( http://www.eucomm.org) and the KOMP project ( http://www.knockoutmouse.org). These projects have been running for approximately three years, and between them aim to create a library of mutant mouse ES cells with targeted knockouts for about half the genes in the mouse genome. Currently the mutant ES cell archive has about 2000 mouse genes knocked out with targeted, conditional mutations. The distribution of vectors and ES cell lines arising from these efforts is done on a cost-recovery basis to serve the international research community.
The information for these resources has been publicly visible for over two years (see www.knockoutmouse.org or www.eucomm.org). The experimental techniques and resources have some supporting published material (see Brief Funct Genomic Proteomic. 2007 Sep;6(3):180-5. Epub 2007 Oct 29 for EUCOMM or Nat Genet. 2004 Sep;36(9):921-4 for KOMP) but the main publication for the experimental technique is to be submitted to Nature Genetics within a couple of weeks. In addition, there are a large number of existing 'random' knockouts for about 10,000 mouse genes, best catalogued at http://www.genetrap.org, which have been in the public eye for a while (Nucleic Acids Res. 2006 Jan 1;34). The 'knockout mouse' wiki page ( http://en.wikipedia.org/wiki/Knockout_mouse) does point to some of these resources already.
I would like to propose that where a mouse gene has an available mutant ES cell generated by these programs, that the corresponding mammalian gene entry is appended with a 'box' similar in vision to the protein box on the mammalian gene pages.
An example of a mammalian gene page is this: http://en.wikipedia.org/wiki/ART4. The 'mutabox' box would give a pointer to further information on all existing mutant alleles for the gene, as well as providing a picture of the molecular structure of a single mutant. Here is a simple straw-man for the Art4 mutabox:
External links to GeneGo Pathways
Would you support the addition of external links on the relevant individual Gene pages to the GeneGo pathways? There are many pathways available in the free portion of the GeneGo site. This would be a parallel resource to the existing GeneCards and OMIM links. -- Kariohlsen ( talk) 01:54, 17 January 2010 (UTC)
- Do you have any examples? What license is the free portion released under? --
Paul (
talk) 21:14, 17 January 2010 (UTC)
- Here is an example for the AKT pathway
link AKT There are relevant pathways for 1000-1500 genes on the free content portion. You can browse from the above link to check them out.--
Kariohlsen (
talk) 19:07, 19 January 2010 (UTC)
- The graphic is pretty an presumably accurate. The site itself is extremely irritating, almost every link I clicked it requested a registration for a 2 week free trial. I suspect this come under point 6 at Wikipedia:External_links#Links_normally_to_be_avoided. What to the other MCBers think? -- Paul ( talk) 20:56, 19 January 2010 (UTC)
- Here is an example for the AKT pathway
link AKT There are relevant pathways for 1000-1500 genes on the free content portion. You can browse from the above link to check them out.--
Kariohlsen (
talk) 19:07, 19 January 2010 (UTC)
Adding interactive pathway maps
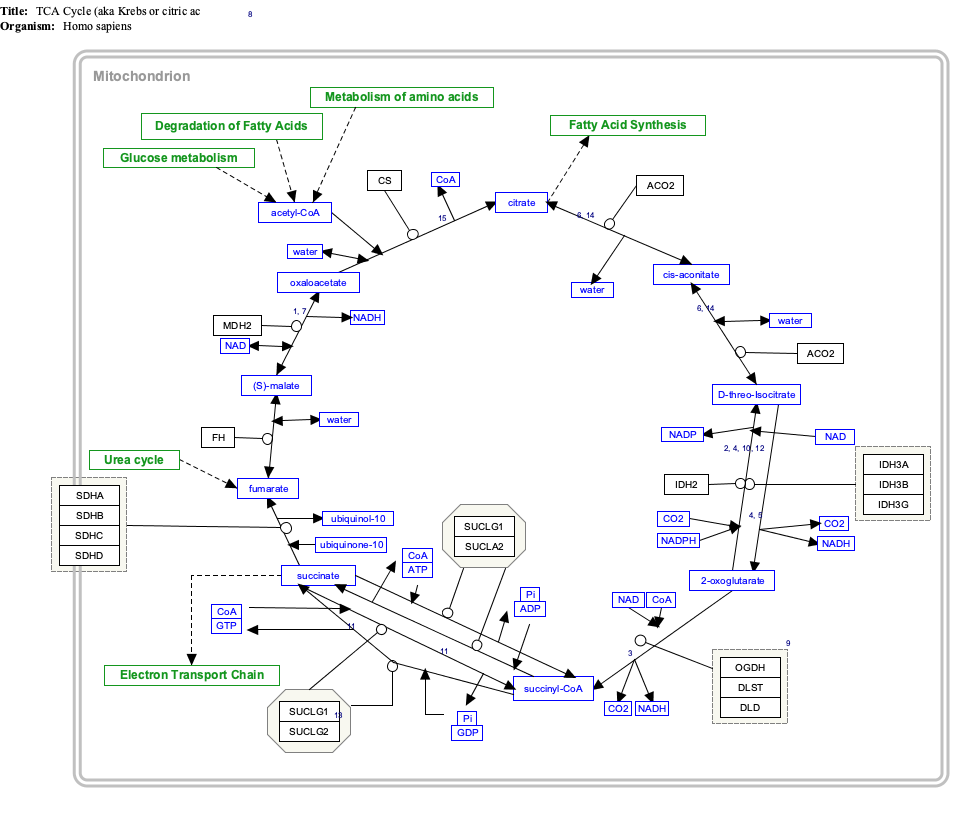
As part of an independent effort called WikiPathways, we are managing the curation of hundreds of biological pathways. This is a 100% free, open source and open access effort. We recently produced imagemaps of these pathway diagrams that include links to wikipedia articles for every gene, protein and metabolite in the pathway, or a link to "start a new article" in cases where one does not exist. We believe these pathway diagrams will not only enrich existing protein and pathways articles, but also enhance the connections across related articles. We've prepared a demonstration using a TCA Cycle pathway template on the following pages:
Notice how the imagemap links to protein and metabolite articles. There are template variables to highlight a particular protein and to control the map size per usage.
I've done my homework and coordinated with Gene Wiki folks in preparing this demo and I've read up on the Metabolic Pathways Task Force, Talk pages and External_links#Links_normally_to_be_avoided, etc. I'm eager for feedback on this proposal. Thanks! AlexanderPico ( talk) 01:29, 1 August 2010 (UTC)
- I think these graphics look great. In addition, they may help reduce the number of orphan Gene Wiki articles. One concern I have however is the size of some of these graphics which tends to overwhelm the rest of the article. I know many of these pathways are complex and therefore the large size may be unavoidable. In addition, the relevant part of the pathway (for example DLAT in the dihydrolipoyl_transacetylase article) may be difficult to locate in the graphic. A possible solution which may alleviate both problems is to highlight the relevant part of the path in much larger font and display the graphic as a smaller thumbnail diagram. That way the relevant part of the pathway is still readable in the thumbnail and the displayed digram is much smaller. If a reader wants to look at the details of the rest of the pathway, then (s)he can clinic on the thumbnail to reveal a larger image. Finally the requirement to edit the diagrams in WikiPathways is a concern. Currently this requires a download of a Java applet that has a certificate that isn't trusted.
- I think these diagrams would be valuable addition to Wikipedia, but I think they may need to tweaked before large scale integration. Cheers.
Boghog (
talk) 07:13, 2 August 2010 (UTC)
- I agree with
Boghog. These have great potential for systematically linking (& de-orphaning) GeneWiki pages. However the are a bit too large. A thumbnail or a collapsible template would be useful. I could envisage similar diagrams using Eg. GO, SO, domains, regulatory networks, ... --
Paul (
talk) 12:47, 2 August 2010 (UTC)
- I think the zoomed-view/thumbnail idea is intriguing. For simplicity, I think Alex is shooting to have one image/template per pathway that is transcluded on multiple gene pages. And he's put in a lot of work to add highlighting of the relevant gene using a passed parameter to the template. I can't think of a way to increase the font size of that highlighted gene, but I wonder if the zoomed/thumbnail diagram could be achieved using
CSS sprites? That might allow there to be only one template per pathway, but still allow a small image to be dynamically generated that shows the local pathway around the highlighted gene. Perhaps someone with more Mediawiki/CSS experience can speak to the feasibility? Cheers,
AndrewGNF (
talk) 15:35, 2 August 2010 (UTC)
- I agree cropping the image around the relevant part of the pathway might be a good solution. Perhaps something like {{
Css Image Crop}} could be used?
Boghog (
talk) 16:44, 2 August 2010 (UTC)
- I now see that the relevant box (DLAT) in the figure is highlighted in yellow, but I initially missed it, since I wasn't aware of the color scheme. Below is an example of a cropped diagram. Ideally clicking on the image should lead to the full sized graphic in Wikipedia which in turn would take you to the WikiPathways page, but it is not immediately clear to me how to do this. Boghog ( talk) 20:42, 2 August 2010 (UTC)
- I agree cropping the image around the relevant part of the pathway might be a good solution. Perhaps something like {{
Css Image Crop}} could be used?
Boghog (
talk) 16:44, 2 August 2010 (UTC)
- I think the zoomed-view/thumbnail idea is intriguing. For simplicity, I think Alex is shooting to have one image/template per pathway that is transcluded on multiple gene pages. And he's put in a lot of work to add highlighting of the relevant gene using a passed parameter to the template. I can't think of a way to increase the font size of that highlighted gene, but I wonder if the zoomed/thumbnail diagram could be achieved using
CSS sprites? That might allow there to be only one template per pathway, but still allow a small image to be dynamically generated that shows the local pathway around the highlighted gene. Perhaps someone with more Mediawiki/CSS experience can speak to the feasibility? Cheers,
AndrewGNF (
talk) 15:35, 2 August 2010 (UTC)
- I agree with
Boghog. These have great potential for systematically linking (& de-orphaning) GeneWiki pages. However the are a bit too large. A thumbnail or a collapsible template would be useful. I could envisage similar diagrams using Eg. GO, SO, domains, regulatory networks, ... --
Paul (
talk) 12:47, 2 August 2010 (UTC)
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "TCACycle_WP78".
Thanks for all the feedback! We were also concerned about the size of the maps. Here are possible solutions:
- Template variables for width and height. The current template supports these variables in units of px. So, we can adopt the convention of a maximum width and heigth when transcluding the imagemap templates. This example is 600x350:
User:AlexanderPico/Test3b
- Disadvantage: you never see the full picture and your highlighted protein may be outside of the initial view. I don't know how to initialize a scroll pane without invoking a hyperlink to it.
- Collapsible navbox. I could enclose the current code in a collapsible navbox (a la metabolic pathways task force), like so:
- User:AlexanderPico/Test4b Click hide/show to control viewing size
- Disadvantage: many users will not click show and completely miss the pathway information and links to related GeneWiki pages. I've tried adding a thumbnail into the header of the navbox, but it's ugly.
- Thumbnail links. Alternatively, there could just be thumbnails on GeneWiki pages that link to the full imagemap at the pathway article.
- Disadvantage: you lose the context of the gene of interest (i.e., the highlighting), thus some users may get disoriented. Unfortunately, we can't have subpages for content like this (I think that's a strict rule, no?). And I don't know how to expand a thumbnail into a template within an article page.
I also looked into CSS Sprites, but again, the problem is how to link from the sprited (or cropped) view to the full view containing useful links to all the related GeneWiki articles. Overall, #1 is the easiest and most flexible. #2 could probably be improved upon if encouraged. I'm concerned that #3 would defeat the purpose. Your continued feedback and ideas would be appreciated! AlexanderPico ( talk) 18:46, 4 August 2010 (UTC)
- I actually quite like Boghog's prototype of {{ TCACycle_WP78 cropped}}, which I think corresponds to your #3. As I see it, isn't the only issue the fact that there is no way to get to the full pathway diagram on Wikipedia (the equivalent of {{ TCACycle_WP78}}, I think)? If my understanding is correct, then I naively assume that there must be a way to change the default link in the cropped template (such that any click not on an explicit entity then goes to the full view). I tried unsuccessfully to modify {{ TCACycle_WP78 cropped}}, but then again there are much more skilled coders here. My two cents... Cheers, AndrewGNF ( talk) 14:36, 5 August 2010 (UTC)
- I like it as well. The challenge seems to be exactly what to link to. Perhaps linking to the full view provided by the corresponding pathway article is sufficient. I've adapted Boghog's "cropped" template accordingly (revisit the example above). The only "bug" in this solution is the confused enlarge-icon which goes to its own image: Magnify-clip.png. Not a big deal? I will update the existing template and instances, and wait for further feedback. Thanks! AlexanderPico ( talk) 01:09, 6 August 2010 (UTC)
- Update: I am pleasantly surprised by how enjoyable the user experience is when going from gene article to pathway article to gene article, etc. Test it out here with PDHA1, Citric Acid Cycle and DLAT. They all interconnect nicely. And by linking specifically to the #Interactive_pathway_map anchor, you could actually navigate the entire pathway from one gene article to the next slightly shifting the crop window each step along the way. Fun! AlexanderPico ( talk) 04:36, 6 August 2010 (UTC)
- Nice! The only thing I'm not sure about is linking to the #Interactive_pathway_map anchor when clicking on defined genes/metabolites. Should it be assumed that the user wants to see the pathway context for that entity, or should it just link to the top of that page? I think I have a slight preference for the latter, but I'm not overly passionate about it either. (Of course, I think the link to #Interactive_pathway_map is perfect when clicking on the map background, since there I think the user's intent is to "enlarge". And agreed about the magnify-clip.png -- I think people would click this icon also thinking "enlarge"...) Cheers, AndrewGNF ( talk) 15:43, 6 August 2010 (UTC)
- All the crop variables are now part of the automatically generated template and can be specified by the single "highlight" variable that also controls the yellow highlighting. This make integration into GeneWiki articles trivial and sustainable, i.e., the template can be updated over the years and the gene articles will automagically adapt to the right crop simply by referencing its gene. Regarding the anchor-links, I'm also not sure. Fortunately, this is trivial to change (adding /removing the anchor from hyperlinks in the template). So, I'd like to roll out 1 or 2 pathways and add the cropped views to all the relevant GeneWiki articles, and then after getting the full experience of how the cross-linking looks and feels we can make a call. Sound good? AlexanderPico ( talk) 19:33, 6 August 2010 (UTC)
- Sounds great to me! Cheers,
AndrewGNF (
talk) 21:15, 6 August 2010 (UTC)
- Looks great to me too! One minor request. The usual Wikipedia convention is that self links, for example in {{
Navbox}} templates, should be bolded. So in addition to highlighting the self link as yellow, would it be possible to also bold the text?
Boghog (
talk) 21:38, 6 August 2010 (UTC)
- Bolding the text is difficult since we have a single png running the template behind every cropped instance. However, we can accommodate the other key aspect of the self-link convention, which is to kill the link. I've adapted the template so that it now removes the blue link line and removes the hyperlink action from the highlighted gene (revisit your cropped example above). I think this gets us pretty close to the convention. And it actually simplies the template quite a bit. Previously, I had to draw 4 separate yellow lines so as to not obscure the self link :)
AlexanderPico (
talk) 04:17, 7 August 2010 (UTC)
- That looks much better. I don't know how it looked in other browsers, but at least in Safari, the yellow lines around the self-link previously didn't completely line the four sides of the box. In the new version, the yellow lines now form a continuous rectangle so that the self-link stands out better.
Boghog (
talk) 05:49, 7 August 2010 (UTC)
- Agreed. Looks better in Firefox and Chrome as well. In keeping with the self-link convention, perhaps a bold black border is more effective than yellow? (see updated example above). I also added template documentation that will be transcluded into each pathway template, e.g.,
Template:TCACycle_WP78.
AlexanderPico (
talk) 20:30, 7 August 2010 (UTC)
- I think the black bold border is slightly preferable to the yellow one since the black border is closer to the self-link convention. I also appreciate your careful documentation of the template. I think these templates will become very popular and good documentation of course will make both maintenance and transcluding these templates into additional articles easier. Finally the templates look quite flexible and easy to use. Nice work! Boghog ( talk) 22:04, 7 August 2010 (UTC)
- Agreed. Looks better in Firefox and Chrome as well. In keeping with the self-link convention, perhaps a bold black border is more effective than yellow? (see updated example above). I also added template documentation that will be transcluded into each pathway template, e.g.,
Template:TCACycle_WP78.
AlexanderPico (
talk) 20:30, 7 August 2010 (UTC)
- That looks much better. I don't know how it looked in other browsers, but at least in Safari, the yellow lines around the self-link previously didn't completely line the four sides of the box. In the new version, the yellow lines now form a continuous rectangle so that the self-link stands out better.
Boghog (
talk) 05:49, 7 August 2010 (UTC)
- Bolding the text is difficult since we have a single png running the template behind every cropped instance. However, we can accommodate the other key aspect of the self-link convention, which is to kill the link. I've adapted the template so that it now removes the blue link line and removes the hyperlink action from the highlighted gene (revisit your cropped example above). I think this gets us pretty close to the convention. And it actually simplies the template quite a bit. Previously, I had to draw 4 separate yellow lines so as to not obscure the self link :)
AlexanderPico (
talk) 04:17, 7 August 2010 (UTC)
- Looks great to me too! One minor request. The usual Wikipedia convention is that self links, for example in {{
Navbox}} templates, should be bolded. So in addition to highlighting the self link as yellow, would it be possible to also bold the text?
Boghog (
talk) 21:38, 6 August 2010 (UTC)
- Sounds great to me! Cheers,
AndrewGNF (
talk) 21:15, 6 August 2010 (UTC)
Interactive pathway maps (part 2)
I was unaware of this thread until I just got a message about it at my talk. Being unaware of it, I had (sorry!) reverted some of the additions of these pathways to pages that I watch (mostly dopamine-related). But I feel flexible about it. I will repeat here part of what I said about it at my talk:
- "To some extent, I was uncomfortable with the sort of promotional nature of the way the text around the pathways image seemed to be promoting an external Wiki, per WP:EL. I'm not sure whether it is within policy here to have links to other Wikis within pages here. More significantly, I was reacting to the size of the images, which I see was also a matter of discussion at the link you gave me. The ones I saw are simply way too big, and should be configured as thumbnails if they are brought back to the main text. In thumbnails, the reader can click on it to see it full size. Finally, some of the text around the images was factually incorrect. For instance, there is no nicotine expressed in dopamine neurons. And there are acetylcholine pathways, but not nicotine ones."
That's what I said by way of concerns at my talk. But I see that this is a work in progress being discussed here, and as I said, I feel flexible about it. I'll watch this discussion now. -- Tryptofish ( talk) 22:13, 3 September 2010 (UTC)
- A specific problem with interactive maps that were added to the dopamine and nicotine articles is that the highlight parameters for dopamine and nicotine are not working in the pathway {{ NicotineDopaminergicActivity WP1602}} template. This brings up another issue that I had not thought of. I think we need to be somewhat selective in deciding which articles that interactive maps should be added to. There are some proteins and metabolites that are involved in many pathways, and some articles could quickly become overwhelmed.
- Concerning the external links, these could be moved to the reference section by replacing the initial sentence with something like:
- Click on genes, proteins and metabolites below to link to the respective Wikipedia articles.<ref name="urlTCA Cycle (Homo sapiens) - WikiPathways">The pathway can be edited at {{cite web | url = http://www.wikipathways.org/index.php/Pathway:WP78 | title = WikiPathways | author = Alexander Pico, Thomas Kelder, Martijn van Iersel, Kristina Hanspers, Kdahlquist, Nick Fidelman | date = | work = | publisher = | pages = | accessdate = }}</ref>
- Based on
this discussion, I have removed the link to
Gene Wiki.
Boghog (
talk) 15:30, 4 September 2010 (UTC)
- I've thought about this some more since my comment here yesterday, and it seems to me that the two paramount issues, for me, are (1) don't mess up the page layout, and (2) don't add anything that is factually incorrect. (Obviously, I'm reacting in terms of the addition to the dopamine page, which is what brought this to my watchlist.) It occurs to me that one part of the problem is that the templates were added as new sections within the page, under their own header. That's the wrong way to go about it. If pathways are to be added to existing pages (it occurs to me that a case could, instead, be made for making new pages about individual pathways), they should be treated as other images are, as visual accompaniments to the text. They shouldn't just be plopped down somewhere on a page. They should appear, as thumbnails, at a reasonable size and with an informative image caption. That caption should relate to the page content, not to the other wiki or to the original topic of the template.
- To expand on these concerns, please look at where the template was added, also, to tyrosine and L-DOPA. In both cases, the template is given its own section header, with text at the top that is all about the project and not at all about the subject matter of the page. The caption at the bottom says "Nicotine in Dopaminergic Neurons (edit)". Well, there is no nicotine in dopaminergic neurons. There's no explanation for the reader of what this information has to do with the rest of those pages. Why is it on L-DOPA? That's a page about a molecule that, although it is a precursor to dopamine, is primarily of interest to readers because it is a pharmaceutical used clinically for Parkinson's disease. Why would a reader interested in Parkinson's medicines want to interact with a biochemical pathway, and what does that have to do with nicotine? And on tyrosine, it treats tyrosine as a precursor to dopamine which is a mediator of nicotine effects, which, by itself, violates WP:UNDUE. Tyrosine is a dopamine precursor, but what about other catecholamines? What about other effects of dopamine besides those related to nicotine? Most importantly, what about all the other pathways in which tyrosine plays important roles? If these kinds of things hadn't set off red flags for me, it would be just a matter of time before other editors would have objected. -- Tryptofish ( talk) 20:37, 4 September 2010 (UTC)
- Tryptofish makes some excellent points. I think the interactive pathways are appropriate to include in Gene Wiki articles where the primary function of the protein is directly linked to the pathway (e.g., PDHA1 in {{ TCACycle_WP78}}) but would be much less appropriate to include this graphic in articles like acetyl-CoA since this molecule is a part of so many biochemical pathways. In addition, it is appropriate to display the entire {{ TCACycle_WP78}} template in an article like citric acid cycle where there is a a close 1:1 relationship between the contents of the interactive pathway and the subject matter of the article. On the other hand, there does not currently appear to be appropriate "parent" articles for templates like {{ NicotineDopaminergicActivity WP1602}}. Hence I agree with Tryptofish that new Wikipedia articles need to be written specifically for these pathways. Boghog ( talk) 11:36, 5 September 2010 (UTC)
- Another issue that I am now reminded of is
WP:RS. The template and/or parent article needs to adequately document the source(s) that were used to create the interactive pathway.
Boghog (
talk) 15:41, 5 September 2010 (UTC)
- Yes, thank you, those are all very good ways to address my concerns. -- Tryptofish ( talk) 16:31, 5 September 2010 (UTC)
- This sounds perfectly reasonable to me. Thank you for the feedback. I will remove the maps where they are non-specific or adding undue weight. I will also update the template to document the sources used to create the content. AlexanderPico ( talk) 19:31, 5 September 2010 (UTC)
Could this be used to create an interactive claddogram navigation tool for taxoboxes? I mean, could there be in a taxobox a picture of the position of the branch of the tree of life of the refferent? And then could the user click on the claddogram and zoom in and out and click on taxa and be sent to articles? Have you seen how they do it at the hall of vertibrates at the American Museum of Natural History in Manhattan? There's a big claddogram at the front and then each hall is laid out to follow the clades, and a cladogram is there at each exhibit so you can see where the animal fits onto the tree of life? Could we do something like that using this tool? I hope I've said enough for you to understand the kind of thing I'm driving at, but if not but if I've piqued your interest and want me to describe this idea in more detail, please ask and I'll try to flesh it out more, but for the moment I don't want to go on too long. Chrisrus ( talk) 04:50, 22 October 2010 (UTC)
- I take it the particle "this" refers to the interactive pathway maps? yes, a
dendrogram/
cladogram is a particular type of
node graph with large limitations to the possible connections and in Matlab one can actually jury-rig (badly) graph theory for NJ nets and such when the standard (polytomy-hating) dendrogram in the Bioinf. toolbox is no good, but overall the conventional representation of dendrograms is very different from node maps and even more so pathway maps, also a modified node graph, which means that it may be best making a separate tool for the tree of life, but, I am sorry to say, I don't think it would be too successful though: currently there is a
Template:Cladogram in wikipedia, which is straightforward and is barely used as most phylogenies are debated (e.g.
malacostraca or the
root of the tree of life), bar for stuff closer to humans that is, there is taxonomy information which can be used uncurated (if you are happy to see a polytomy of 16+ bacterial divisions that is). --
Squidonius (
talk) 08:26, 22 October 2010 (UTC)
- If you visit the fouth floor of the American Museum of Natural History, you are greeted by a huge screen tree of life, which basically explains how it works and then focuses on the branch that is dealt with on the floor, that of the vertibrates. Then, each exhibit reproduces the same branch in more detail. It's really fabulous in terms of presenting thngs, as opposed to the traditional way, as on the other floors, by geographic area "Animals of Africa", or by era "dinosaurs of the Jerassic" or by a taxon or word like "Birds". While it may not be perfect, according to the article cladistics, it's more accurate then taxonomy, which (while indispensable) also features many areas that are debated or complicated, and is included in every article nevertheless. You will agree that the situation about every type of living thing should be presented in as complicated a way as it needs to be and no more. And we presently often settle on such tree diagrams in articles attempting to present such complicated situations, and tend to omit them more when things are relatively straightforward. So that makes me think that the fact that things are complicated or debated would argue for, rather than against, such an inclusion. Chrisrus ( talk) 13:03, 22 October 2010 (UTC)
Possible relation of gap-43 with autism disorders
I would like to know if we could include a possible relation of protein gap-43 with autism, as stated on this scientific article:
http://www.jneurosci.org/cgi/content/abstract/30/44/14595
And this:
http://blog.autismspeaks.org/2010/11/11/what-lies-beneath-brain-connections/#comment-9286
The page I think should be modified is this:
http://en.wikipedia.org/wiki/Gap-43_protein
- I don't see any problem with adding the first citation to the Gap-43_protein#Clinical_significance section since it is to a peer reviewed article and very relevant to the subject matter of the Wikipedia article. However the second link is a blog and blogs are often not considered a reliable sources. But this particular blog link since it sumarizes the peer reviewed article could be added as a "lay summary" combined with the first citation:
- * Zikopoulos B, Barbas H (November 2010). "Changes in prefrontal axons may disrupt the network in autism". J. Neurosci. 30 (44): 14595–609.
doi:
10.1523/JNEUROSCI.2257-10.2010.
PMID
21048117.
{{ cite journal}}: Unknown parameter|laysource=ignored ( help); Unknown parameter|laysummary=ignored ( help) - Finally please note that it is not necessary to ask the community about adding material to a Wikipedia article, unless you think it might be controversial (if so, it is appropriate to bring up the issue directly on the article's talk page). I don't see anything that is particularly controversial about the material that you propose to add, so be bold! Boghog ( talk) 20:46, 12 November 2010 (UTC)
Infobox biodatabase
Hi all, the latest Database Issue of NAR has just been published and I wonder if it would be worth to create an infobox biodatabase. For each database I would write an article with this infobox. The fields would be:
- center
- author(s)
- date ...
- specific organism(s)
- URL
- Web Services ( SOAP, WSDL URLs etc...)
- public Mysql access
- etc...
I plan to use a XSLT stylesheet to quiclky write a WP article from a pubmed abstract. -- Plindenbaum ( talk) 14:59, 5 January 2011 (UTC)
- i think that's a great idea, maybe even link it to Biological_databases. I'd be happy to go through them afterwards to organize them into some kind of grouped table to people can quickly see what is available. It might also be worthwhile to ping an email to some of the corresponding authors to let them know, and to invite them to keep their database page up to date. Davebridges ( talk)
- Good idea. You can find many existing DB articles in Category:Biological databases. -- Paul ( talk) 16:34, 5 January 2011 (UTC)
OK, I started here: Template:Infobox_biodatabase-- Plindenbaum ( talk) 18:17, 5 January 2011 (UTC)
- Thanks for staring this. Perhaps I am biased, but I think the current template places too much emphasis on computer science (i.e., the format) and not enough on biology (i.e., the content). Also I question the importance of including information on the release version. This will be difficult to maintain and is of questionable value to most readers. If someone is really interested about the version number of the latest release, they will check the website directly. Boghog ( talk) 18:33, 5 January 2011 (UTC)
- Suggested additions to template:
- Name
- Description
- Category (Nucleotide Sequence, RNA sequence, Protein sequence, Structure, Genomics (non-vertebrate), Metabolic and Signaling Pathways, Human and other Vertebrate Genomes, Human Genes and Diseases, Microarray Data and other Gene Expression, Proteomics Resources, Other Molecular Biology, Organelle, Plant, Immunological)
- Boghog ( talk) 18:53, 5 January 2011 (UTC)
- Copying the software template was just for a start, I modified the template, added a 'doc page' and a sandbox page. But feel free to edit this infobox :-)-- Plindenbaum ( talk) 19:52, 5 January 2011 (UTC)
- Suggested additions to template:
This is a timely suggestion. Recently a paper has been published about something called BioDBCore. This initiative aims to define a set of minimal information requirements about a database. In fact it is very much like what we would want for an infobox. BioDBCore currently define 17 different fields of information, including the resources Wikipedia entry! It would make sense that as far as possible we try to mirror that set up. If it does become a widely adopted standard it will make it easier for us to keep these boxes up to date in the future. The paper includes an excel file with some examples from some well known databases.
The paper is here: http://www.ncbi.nlm.nih.gov/pubmed/21097465 The field description is in the excel file in the supp. materials: http://nar.oxfordjournals.org/content/early/2010/11/17/nar.gkq1173/suppl/DC1
Here is the list of their fields:
- Database name
- Main resource URL
- Contact information
- Date resource established (year)
- Conditions of use
- Scope:
- Data types captured
- Curation policy
- Standards: MIs, Data formats, Terminologies
- Data formats
- Taxonomic coverage (use NCBI Taxid)
- Data accessibility/output options
- Data release frequency
- Versioning policy/ access to historical files
- Documentation available
- User support options
- Data submission policy
- Relevant publications
- Resource’s Wikipedia URL
- Tools available
Thanks for starting this! Alexbateman ( talk) 08:28, 6 January 2011 (UTC)
- Thanks for your comment Alex, I've added a few fields according to your suggestions and I've updated the documentation.--
Plindenbaum (
talk) 09:44, 6 January 2011 (UTC)
-
Looks good!
Boghog (
talk) 11:51, 6 January 2011 (UTC)
- In my opinion the description field will be redundant with the lead of the Wikipedia article and should be deleted. I don't see a description field being used for other infoboxes such as taxoboxes, protein family boxes etc. Although sometimes there is a figure caption used.
193.62.202.241 (
talk) 12:41, 6 January 2011 (UTC)
- I imagine, 'description' should be used for as a title, not for a complete description: eg:
- In my opinion the description field will be redundant with the lead of the Wikipedia article and should be deleted. I don't see a description field being used for other infoboxes such as taxoboxes, protein family boxes etc. Although sometimes there is a figure caption used.
193.62.202.241 (
talk) 12:41, 6 January 2011 (UTC)
-
Looks good!
Boghog (
talk) 11:51, 6 January 2011 (UTC)
| Content | |
|---|---|
| Description | Comparative genomics of Fusarium strains, |
| Contact | |
| Primary citation | PMID 21087991 |
| Access | |
| Website | http://www.fusarium.org |
-- Plindenbaum ( talk) 13:10, 6 January 2011 (UTC)
- I don't see the description as being necessarily redundant with the lead, but rather a very brief summary of what is contained in the database (see for example the GenBank article). The lead often runs several paragraphs and is not consistent between articles. The description in the infobox is more structured and allow the reader at a glance to figure out what is contained in the database. Boghog ( talk) 13:37, 6 January 2011 (UTC)
I wrote a Xslt stylesheet to transform a Pubmed article to wikipedia. See Template:Infobox_biodatabase/doc for a demo. And here are a two pages I created with this stylesheet : PlasmoDB and Rebase. I also created the template Template:Biodatabase-stub -- Plindenbaum ( talk) 21:19, 6 January 2011 (UTC)
- A while back I created a page for
cancer research databases. But if you now create a WP page for each database with an infobox it will be kind of redundant. So is it better to maintain both or move all to the same schema? I personally think that this kind of organization gives a better overview at one glance and is easier to maintain.
Michipanero (
talk) 09:12, 8 April 2011 (UTC)
- @ Michipanero : my goal is to be able to search for a databases using a SPARQL query (once the database will be digested by DBPedia). That's why I need this kind of infobox. -- Plindenbaum ( talk) 11:49, 13 April 2011 (UTC)
- A while back I created a page for
cancer research databases. But if you now create a WP page for each database with an infobox it will be kind of redundant. So is it better to maintain both or move all to the same schema? I personally think that this kind of organization gives a better overview at one glance and is easier to maintain.
Michipanero (
talk) 09:12, 8 April 2011 (UTC)
FYI the data from Template:Infobox_biodatabase have now been integrated into DBPedia 3.7. See:
- http://en.wikipedia.org/wiki/Wikipedia:WikiProject_Molecular_and_Cellular_Biology/Announcements#Infobox_biodatabase (for reference)
- http://dbpedia.org/page/Template:Infobox_biodatabase
- http://dbpedia.org/ontology/BiologicalDatabase
- http://dbpedia.org/page/L1Base
those data can now be searched through a SPARQL query -- Plindenbaum ( talk) 09:56, 11 September 2011 (UTC)
Ribosyl group
I believe it would be informative as well as useful to create a stub on the ribosyl group. I submitted an article for review, albeit, more of a definition, and it was promptly rejected. So I concluded that joining the project and hearing some opinions on whether it is important enough to have its own article, or if the ribose article could be edited to include its radical and its subsequent presence in proteins and enzymes. Let me know what you think. Dissonase 02:12, 17 February 2011 (UTC)
- Unless the proposed ribosyl article were significantly expanded, I think it would be better to combine it with the existing ribose article and create redirects for ribosyl, ribosyl group, ribosyl radical, etc. pointing to ribose. The ribose article already mentions that ribose is a component of RNA. This could be expanded to mention that the ribosyl functional group can also be attached to other macromolecules. Please be sure to include reliable sources to document that ribose is a component of each of the macromolecules mentioned. Cheers. Boghog ( talk) 06:44, 17 February 2011 (UTC)
Comments on Merging Invariant Chain with CD74
Discuss. Blahdenoma ( talk) 05:59, 30 April 2011 (UTC)
 Done I agree with the merger proposal. I therefore boldly went ahead and completed the merger.
Boghog (
talk) 08:07, 30 April 2011 (UTC)
Done I agree with the merger proposal. I therefore boldly went ahead and completed the merger.
Boghog (
talk) 08:07, 30 April 2011 (UTC)
| Mouse Mutant Alleles for Art4 | ||
|---|---|---|
| Marker Symbol for Mouse Gene. This symbol is assigned to the genomic locus by the MGI | Art4 | |
| Mutant Mouse Embryonic Stem Cell Clones. These are the known targeted mutations for this gene in a mouse. | Art4tm1aWTSI(KOMP) | |
| Example structure of targeted conditional mutant allele for this gene | ||
 | ||
| These Mutant ES Cells can be studied directly or used to generate mice with this gene knocked out. Study of these mice can shed light on the function of Art4 see Knockout mouse | ||
I envision the mutabox updates to the gene pages will be made by a robot ('mutabot', still to be proposed & written) which scans the known high-throughput mutations at www.knockoutmouse.org and applies the changes to a template in each mammalian gene entry. Also note that all mutabox entries would point back to the existing wiki page Knockout mouse], which explains further why these mutants are scientifically interesting (they help the study of gene function), and gives more links.
I would like to get opinions (and hopefully consensus) from the MCB group for this proposal.
Regards,
Vivek Iyer
High Throughput Gene Targeting
Wellcome Trust Sanger Institute
- Hi Vivek, On the surface this looks like it could be a useful resource for experimental biologists. But a concern could be that this effort could be construed as advertising for a commercial entity - the links to 'order vector' and 'order ES cells' set off warning bells. I've had a quick look around the Wikipedia policy pages and as usual I haven't found too much of use. Possibly WP:LINKS and conflict-of-interest should be checked against what you want to do.-- Paul ( talk) 09:52, 24 February 2009 (UTC)
- Hi Paul, Thanks for the observation and the references. Regarding the concern with advertising a commercial enterprise: I should have emphasised that these consortia are NOT commercial enterprises - they are funded by the NIH and the EU (and the Wellcome Trust) to serve the international research community. The 'order' buttons you mentioned put end-users quickly in touch with the vector & ES Cell distribution units, which are also run as part of academic institutes - UC Davis, CHORI in Oakland and the HMGU in Munich. Those units do function on a cost-recovery basis, so there is a charge for products, but that mode of working has been established for a long time. Anyway, I've modified the text of the proposal to say this explicitly. As for the conflict of interest, I hope I've made my affiliation to the program informatics clear, but let me know if I've missed something. Does this allay your concerns? -- Vivek 11.35, 24 February 2009 (UTC)
- I've now made the proposed edit directly onto http://en.wikipedia.org/wiki/ART4 . An proposal for a BOT is in progress. Vivek ( talk) 11.35, 19 March 2009 (UTC)
- Vivek, the prototypes look good to me... You may get more specific feedback from the bot approval group, but you may want to convert the mutabox to wikitext tables (see Help:Tables) instead of HTML. Cheers, AndrewGNF ( talk) 04:42, 14 April 2009 (UTC)
- Hi Vivek, apologies I have come rather late to this discussion. I have some concerns about your proposed boxes, mainly to do with their size and whether they give undue prominence to information that is not so important to the likely users of wikipedia. Some points:
- whilst some gene pages are probably only of interest to those of us who are researchers, some e.g. perhaps huntingtin, p53 etc. are probably of interest to undergraduates and non-scientists. I think a large box on a research tool that will dominate many short pages is inappropriate in these circumstances.
- the schematic showing your construct. Forgive me, but once you have seen one, you have seen them all essentially - they all follow the same strategy? I don't really think that they are useful. I would prefer them to be removed or initially hidden with a 'show' control.
- the boxes are very big. Would it not be preferable to add the information instead to the gene box in a short form e.g. "Mouse knockout available" "ES cell mutant available" with a link to your database here?
- (minor practical concern) in Safari 4 on my computer the mutabox display overlaps with the gene box breaking the page display.
- Best wishes, Celefin ( talk) 19:34, 17 April 2009 (UTC)
- Hi Vivek, apologies I have come rather late to this discussion. I have some concerns about your proposed boxes, mainly to do with their size and whether they give undue prominence to information that is not so important to the likely users of wikipedia. Some points:
- Hi Celefin, thankyou for the feedback, this is just the kind of thing I need (difficult to pick up from the 'inside'). I think the best response is a reworking of the mutabox examples to address both your and AndrewGNF's comments, but here are some quick thoughts
- The box is probably too big, and can be shrunk down somewhat, and changed from an html table into a wiki table. I haven't experimented to see if it will help the page display problem you mention, but I noticed the layout on my browser (also safari) was not happy when the browser window got too small - I am sure there is a good way of dealing with that without stuffing up the page or dominating it.
- Those pictures are all different, reflecting both gene structure and the exact mutagenic cassette, which can follow quite different strategies. The pipelines have been rejigging themselves a lot, and the changes matter. I think the existence of a picture does actually draw a readers attention better than plain text. A molecular biologist will interpret them, but I had in mind a non-specialist could at least get drawn by that into the text, which I have made deliberately very simple and general.
- I disagree that information about mutations shouldn't swamp a short page. First, I can & will rework the mutabox so it doesn't swamp anything, but If there are pages which are short (containing only a brief description and references), then shouldn't they be fleshed out with other relevant information? If that information is the existence of a mutant allele, isn't that a good thing? I was thinking that pages which are essentially stubs right now would / grow, right?
- Thanks again for the comments! Vivek ( talk) 21:45, 20 April 2009 (UTC)
- Hi Celefin, thankyou for the feedback, this is just the kind of thing I need (difficult to pick up from the 'inside'). I think the best response is a reworking of the mutabox examples to address both your and AndrewGNF's comments, but here are some quick thoughts
IRC channel to recruit people and as a place to hang out...
I'd propose a channel on freenode.net (just like #wikipedia has)... my suggestion would be the following channel irc://irc.freenode.net/#wikipedia-mcb
There are a bunch of people in irc://irc.freenode.net/#bioinformatics for example... I'm sure you could recruit some of them if you had an irc presence... also I'd just quite like to chat to some of you guys some time ;-)
-- Dan| (talk) 18:35, 14 April 2009 (UTC)
Merge Wikipedia:WikiProject Cell Signaling with this project as a task force?
I made a suggestion over at the cell signaling wikiproject (see here [1]) to consider merging into this project as a task force. I noticed above there has been suggestions to make a few task forces but not sure whether they went ahead as planned? Anyone here happen to have any suggestions/comments they would like to make re: merging cell signalling wikiproject as a task force here? Cheers. Calaka ( talk) 11:13, 5 May 2009 (UTC)
Photosynthesis Taskforce
I think a photosynthesis taskforce should be a great idea. A couple of months ago I tried to make a wikiproject about this subject, but a taskforce should be better. At the moment the most important articles about this subject (Photosynthesis, Light Reactions, Dark reactions, Chloroplast) are not anything near FA-status. (Altough, I have put a lot time in the light reactions article). And all the other articles of this subject are mostly stubs.
I think the photosynthesis article itself is one of the examples of what happens if you have to many authors. One of the three guys that have ever posted something on the wikiproject photosynthesis wrote this about it:
"In my opinion, this article is one of the worst on WikiPedia. There is a heck of a lot of good information on this page, but it is obtuse, chaotic and so badly written that is absolutely useless. The only people who can understand it are the people who already know it; a solid case of preaching to the choir, if I've ever seen one."
And I think he is totally right...
So what do you guys think? Kasper90 ( talk) 16:42, 26 May 2009 (UTC)
- I don't see why not. Will you be willing to set up the task force page up though and add in the required info/introduction/guidelines? Might be a bit of work. In my case (for the eventual merger of the sister projects onto here), I pretty much got the template, just need to do a few tweaks here and there. Test out the look/feel of it on a sandbox perhaps? Calaka ( talk) 10:54, 30 May 2009 (UTC)
| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | Archive 2 | Archive 3 | Archive 4 |
Mutabox: a bot-generated box in mammalian gene pages, referring to targeted mutant alleles in mice.
Dear MCB group,
I am involved in the informatics of two high-throughput gene-knockout projects conducted in part by Team87 at the Wellcome Trust Sanger Institute. These are the eucomm project ( http://www.eucomm.org) and the KOMP project ( http://www.knockoutmouse.org). These projects have been running for approximately three years, and between them aim to create a library of mutant mouse ES cells with targeted knockouts for about half the genes in the mouse genome. Currently the mutant ES cell archive has about 2000 mouse genes knocked out with targeted, conditional mutations. The distribution of vectors and ES cell lines arising from these efforts is done on a cost-recovery basis to serve the international research community.
The information for these resources has been publicly visible for over two years (see www.knockoutmouse.org or www.eucomm.org). The experimental techniques and resources have some supporting published material (see Brief Funct Genomic Proteomic. 2007 Sep;6(3):180-5. Epub 2007 Oct 29 for EUCOMM or Nat Genet. 2004 Sep;36(9):921-4 for KOMP) but the main publication for the experimental technique is to be submitted to Nature Genetics within a couple of weeks. In addition, there are a large number of existing 'random' knockouts for about 10,000 mouse genes, best catalogued at http://www.genetrap.org, which have been in the public eye for a while (Nucleic Acids Res. 2006 Jan 1;34). The 'knockout mouse' wiki page ( http://en.wikipedia.org/wiki/Knockout_mouse) does point to some of these resources already.
I would like to propose that where a mouse gene has an available mutant ES cell generated by these programs, that the corresponding mammalian gene entry is appended with a 'box' similar in vision to the protein box on the mammalian gene pages.
An example of a mammalian gene page is this: http://en.wikipedia.org/wiki/ART4. The 'mutabox' box would give a pointer to further information on all existing mutant alleles for the gene, as well as providing a picture of the molecular structure of a single mutant. Here is a simple straw-man for the Art4 mutabox:
External links to GeneGo Pathways
Would you support the addition of external links on the relevant individual Gene pages to the GeneGo pathways? There are many pathways available in the free portion of the GeneGo site. This would be a parallel resource to the existing GeneCards and OMIM links. -- Kariohlsen ( talk) 01:54, 17 January 2010 (UTC)
- Do you have any examples? What license is the free portion released under? --
Paul (
talk) 21:14, 17 January 2010 (UTC)
- Here is an example for the AKT pathway
link AKT There are relevant pathways for 1000-1500 genes on the free content portion. You can browse from the above link to check them out.--
Kariohlsen (
talk) 19:07, 19 January 2010 (UTC)
- The graphic is pretty an presumably accurate. The site itself is extremely irritating, almost every link I clicked it requested a registration for a 2 week free trial. I suspect this come under point 6 at Wikipedia:External_links#Links_normally_to_be_avoided. What to the other MCBers think? -- Paul ( talk) 20:56, 19 January 2010 (UTC)
- Here is an example for the AKT pathway
link AKT There are relevant pathways for 1000-1500 genes on the free content portion. You can browse from the above link to check them out.--
Kariohlsen (
talk) 19:07, 19 January 2010 (UTC)
Adding interactive pathway maps
As part of an independent effort called WikiPathways, we are managing the curation of hundreds of biological pathways. This is a 100% free, open source and open access effort. We recently produced imagemaps of these pathway diagrams that include links to wikipedia articles for every gene, protein and metabolite in the pathway, or a link to "start a new article" in cases where one does not exist. We believe these pathway diagrams will not only enrich existing protein and pathways articles, but also enhance the connections across related articles. We've prepared a demonstration using a TCA Cycle pathway template on the following pages:
Notice how the imagemap links to protein and metabolite articles. There are template variables to highlight a particular protein and to control the map size per usage.
I've done my homework and coordinated with Gene Wiki folks in preparing this demo and I've read up on the Metabolic Pathways Task Force, Talk pages and External_links#Links_normally_to_be_avoided, etc. I'm eager for feedback on this proposal. Thanks! AlexanderPico ( talk) 01:29, 1 August 2010 (UTC)
- I think these graphics look great. In addition, they may help reduce the number of orphan Gene Wiki articles. One concern I have however is the size of some of these graphics which tends to overwhelm the rest of the article. I know many of these pathways are complex and therefore the large size may be unavoidable. In addition, the relevant part of the pathway (for example DLAT in the dihydrolipoyl_transacetylase article) may be difficult to locate in the graphic. A possible solution which may alleviate both problems is to highlight the relevant part of the path in much larger font and display the graphic as a smaller thumbnail diagram. That way the relevant part of the pathway is still readable in the thumbnail and the displayed digram is much smaller. If a reader wants to look at the details of the rest of the pathway, then (s)he can clinic on the thumbnail to reveal a larger image. Finally the requirement to edit the diagrams in WikiPathways is a concern. Currently this requires a download of a Java applet that has a certificate that isn't trusted.
- I think these diagrams would be valuable addition to Wikipedia, but I think they may need to tweaked before large scale integration. Cheers.
Boghog (
talk) 07:13, 2 August 2010 (UTC)
- I agree with
Boghog. These have great potential for systematically linking (& de-orphaning) GeneWiki pages. However the are a bit too large. A thumbnail or a collapsible template would be useful. I could envisage similar diagrams using Eg. GO, SO, domains, regulatory networks, ... --
Paul (
talk) 12:47, 2 August 2010 (UTC)
- I think the zoomed-view/thumbnail idea is intriguing. For simplicity, I think Alex is shooting to have one image/template per pathway that is transcluded on multiple gene pages. And he's put in a lot of work to add highlighting of the relevant gene using a passed parameter to the template. I can't think of a way to increase the font size of that highlighted gene, but I wonder if the zoomed/thumbnail diagram could be achieved using
CSS sprites? That might allow there to be only one template per pathway, but still allow a small image to be dynamically generated that shows the local pathway around the highlighted gene. Perhaps someone with more Mediawiki/CSS experience can speak to the feasibility? Cheers,
AndrewGNF (
talk) 15:35, 2 August 2010 (UTC)
- I agree cropping the image around the relevant part of the pathway might be a good solution. Perhaps something like {{
Css Image Crop}} could be used?
Boghog (
talk) 16:44, 2 August 2010 (UTC)
- I now see that the relevant box (DLAT) in the figure is highlighted in yellow, but I initially missed it, since I wasn't aware of the color scheme. Below is an example of a cropped diagram. Ideally clicking on the image should lead to the full sized graphic in Wikipedia which in turn would take you to the WikiPathways page, but it is not immediately clear to me how to do this. Boghog ( talk) 20:42, 2 August 2010 (UTC)
- I agree cropping the image around the relevant part of the pathway might be a good solution. Perhaps something like {{
Css Image Crop}} could be used?
Boghog (
talk) 16:44, 2 August 2010 (UTC)
- I think the zoomed-view/thumbnail idea is intriguing. For simplicity, I think Alex is shooting to have one image/template per pathway that is transcluded on multiple gene pages. And he's put in a lot of work to add highlighting of the relevant gene using a passed parameter to the template. I can't think of a way to increase the font size of that highlighted gene, but I wonder if the zoomed/thumbnail diagram could be achieved using
CSS sprites? That might allow there to be only one template per pathway, but still allow a small image to be dynamically generated that shows the local pathway around the highlighted gene. Perhaps someone with more Mediawiki/CSS experience can speak to the feasibility? Cheers,
AndrewGNF (
talk) 15:35, 2 August 2010 (UTC)
- I agree with
Boghog. These have great potential for systematically linking (& de-orphaning) GeneWiki pages. However the are a bit too large. A thumbnail or a collapsible template would be useful. I could envisage similar diagrams using Eg. GO, SO, domains, regulatory networks, ... --
Paul (
talk) 12:47, 2 August 2010 (UTC)
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "TCACycle_WP78".
Thanks for all the feedback! We were also concerned about the size of the maps. Here are possible solutions:
- Template variables for width and height. The current template supports these variables in units of px. So, we can adopt the convention of a maximum width and heigth when transcluding the imagemap templates. This example is 600x350:
User:AlexanderPico/Test3b
- Disadvantage: you never see the full picture and your highlighted protein may be outside of the initial view. I don't know how to initialize a scroll pane without invoking a hyperlink to it.
- Collapsible navbox. I could enclose the current code in a collapsible navbox (a la metabolic pathways task force), like so:
- User:AlexanderPico/Test4b Click hide/show to control viewing size
- Disadvantage: many users will not click show and completely miss the pathway information and links to related GeneWiki pages. I've tried adding a thumbnail into the header of the navbox, but it's ugly.
- Thumbnail links. Alternatively, there could just be thumbnails on GeneWiki pages that link to the full imagemap at the pathway article.
- Disadvantage: you lose the context of the gene of interest (i.e., the highlighting), thus some users may get disoriented. Unfortunately, we can't have subpages for content like this (I think that's a strict rule, no?). And I don't know how to expand a thumbnail into a template within an article page.
I also looked into CSS Sprites, but again, the problem is how to link from the sprited (or cropped) view to the full view containing useful links to all the related GeneWiki articles. Overall, #1 is the easiest and most flexible. #2 could probably be improved upon if encouraged. I'm concerned that #3 would defeat the purpose. Your continued feedback and ideas would be appreciated! AlexanderPico ( talk) 18:46, 4 August 2010 (UTC)
- I actually quite like Boghog's prototype of {{ TCACycle_WP78 cropped}}, which I think corresponds to your #3. As I see it, isn't the only issue the fact that there is no way to get to the full pathway diagram on Wikipedia (the equivalent of {{ TCACycle_WP78}}, I think)? If my understanding is correct, then I naively assume that there must be a way to change the default link in the cropped template (such that any click not on an explicit entity then goes to the full view). I tried unsuccessfully to modify {{ TCACycle_WP78 cropped}}, but then again there are much more skilled coders here. My two cents... Cheers, AndrewGNF ( talk) 14:36, 5 August 2010 (UTC)
- I like it as well. The challenge seems to be exactly what to link to. Perhaps linking to the full view provided by the corresponding pathway article is sufficient. I've adapted Boghog's "cropped" template accordingly (revisit the example above). The only "bug" in this solution is the confused enlarge-icon which goes to its own image: Magnify-clip.png. Not a big deal? I will update the existing template and instances, and wait for further feedback. Thanks! AlexanderPico ( talk) 01:09, 6 August 2010 (UTC)
- Update: I am pleasantly surprised by how enjoyable the user experience is when going from gene article to pathway article to gene article, etc. Test it out here with PDHA1, Citric Acid Cycle and DLAT. They all interconnect nicely. And by linking specifically to the #Interactive_pathway_map anchor, you could actually navigate the entire pathway from one gene article to the next slightly shifting the crop window each step along the way. Fun! AlexanderPico ( talk) 04:36, 6 August 2010 (UTC)
- Nice! The only thing I'm not sure about is linking to the #Interactive_pathway_map anchor when clicking on defined genes/metabolites. Should it be assumed that the user wants to see the pathway context for that entity, or should it just link to the top of that page? I think I have a slight preference for the latter, but I'm not overly passionate about it either. (Of course, I think the link to #Interactive_pathway_map is perfect when clicking on the map background, since there I think the user's intent is to "enlarge". And agreed about the magnify-clip.png -- I think people would click this icon also thinking "enlarge"...) Cheers, AndrewGNF ( talk) 15:43, 6 August 2010 (UTC)
- All the crop variables are now part of the automatically generated template and can be specified by the single "highlight" variable that also controls the yellow highlighting. This make integration into GeneWiki articles trivial and sustainable, i.e., the template can be updated over the years and the gene articles will automagically adapt to the right crop simply by referencing its gene. Regarding the anchor-links, I'm also not sure. Fortunately, this is trivial to change (adding /removing the anchor from hyperlinks in the template). So, I'd like to roll out 1 or 2 pathways and add the cropped views to all the relevant GeneWiki articles, and then after getting the full experience of how the cross-linking looks and feels we can make a call. Sound good? AlexanderPico ( talk) 19:33, 6 August 2010 (UTC)
- Sounds great to me! Cheers,
AndrewGNF (
talk) 21:15, 6 August 2010 (UTC)
- Looks great to me too! One minor request. The usual Wikipedia convention is that self links, for example in {{
Navbox}} templates, should be bolded. So in addition to highlighting the self link as yellow, would it be possible to also bold the text?
Boghog (
talk) 21:38, 6 August 2010 (UTC)
- Bolding the text is difficult since we have a single png running the template behind every cropped instance. However, we can accommodate the other key aspect of the self-link convention, which is to kill the link. I've adapted the template so that it now removes the blue link line and removes the hyperlink action from the highlighted gene (revisit your cropped example above). I think this gets us pretty close to the convention. And it actually simplies the template quite a bit. Previously, I had to draw 4 separate yellow lines so as to not obscure the self link :)
AlexanderPico (
talk) 04:17, 7 August 2010 (UTC)
- That looks much better. I don't know how it looked in other browsers, but at least in Safari, the yellow lines around the self-link previously didn't completely line the four sides of the box. In the new version, the yellow lines now form a continuous rectangle so that the self-link stands out better.
Boghog (
talk) 05:49, 7 August 2010 (UTC)
- Agreed. Looks better in Firefox and Chrome as well. In keeping with the self-link convention, perhaps a bold black border is more effective than yellow? (see updated example above). I also added template documentation that will be transcluded into each pathway template, e.g.,
Template:TCACycle_WP78.
AlexanderPico (
talk) 20:30, 7 August 2010 (UTC)
- I think the black bold border is slightly preferable to the yellow one since the black border is closer to the self-link convention. I also appreciate your careful documentation of the template. I think these templates will become very popular and good documentation of course will make both maintenance and transcluding these templates into additional articles easier. Finally the templates look quite flexible and easy to use. Nice work! Boghog ( talk) 22:04, 7 August 2010 (UTC)
- Agreed. Looks better in Firefox and Chrome as well. In keeping with the self-link convention, perhaps a bold black border is more effective than yellow? (see updated example above). I also added template documentation that will be transcluded into each pathway template, e.g.,
Template:TCACycle_WP78.
AlexanderPico (
talk) 20:30, 7 August 2010 (UTC)
- That looks much better. I don't know how it looked in other browsers, but at least in Safari, the yellow lines around the self-link previously didn't completely line the four sides of the box. In the new version, the yellow lines now form a continuous rectangle so that the self-link stands out better.
Boghog (
talk) 05:49, 7 August 2010 (UTC)
- Bolding the text is difficult since we have a single png running the template behind every cropped instance. However, we can accommodate the other key aspect of the self-link convention, which is to kill the link. I've adapted the template so that it now removes the blue link line and removes the hyperlink action from the highlighted gene (revisit your cropped example above). I think this gets us pretty close to the convention. And it actually simplies the template quite a bit. Previously, I had to draw 4 separate yellow lines so as to not obscure the self link :)
AlexanderPico (
talk) 04:17, 7 August 2010 (UTC)
- Looks great to me too! One minor request. The usual Wikipedia convention is that self links, for example in {{
Navbox}} templates, should be bolded. So in addition to highlighting the self link as yellow, would it be possible to also bold the text?
Boghog (
talk) 21:38, 6 August 2010 (UTC)
- Sounds great to me! Cheers,
AndrewGNF (
talk) 21:15, 6 August 2010 (UTC)
Interactive pathway maps (part 2)
I was unaware of this thread until I just got a message about it at my talk. Being unaware of it, I had (sorry!) reverted some of the additions of these pathways to pages that I watch (mostly dopamine-related). But I feel flexible about it. I will repeat here part of what I said about it at my talk:
- "To some extent, I was uncomfortable with the sort of promotional nature of the way the text around the pathways image seemed to be promoting an external Wiki, per WP:EL. I'm not sure whether it is within policy here to have links to other Wikis within pages here. More significantly, I was reacting to the size of the images, which I see was also a matter of discussion at the link you gave me. The ones I saw are simply way too big, and should be configured as thumbnails if they are brought back to the main text. In thumbnails, the reader can click on it to see it full size. Finally, some of the text around the images was factually incorrect. For instance, there is no nicotine expressed in dopamine neurons. And there are acetylcholine pathways, but not nicotine ones."
That's what I said by way of concerns at my talk. But I see that this is a work in progress being discussed here, and as I said, I feel flexible about it. I'll watch this discussion now. -- Tryptofish ( talk) 22:13, 3 September 2010 (UTC)
- A specific problem with interactive maps that were added to the dopamine and nicotine articles is that the highlight parameters for dopamine and nicotine are not working in the pathway {{ NicotineDopaminergicActivity WP1602}} template. This brings up another issue that I had not thought of. I think we need to be somewhat selective in deciding which articles that interactive maps should be added to. There are some proteins and metabolites that are involved in many pathways, and some articles could quickly become overwhelmed.
- Concerning the external links, these could be moved to the reference section by replacing the initial sentence with something like:
- Click on genes, proteins and metabolites below to link to the respective Wikipedia articles.<ref name="urlTCA Cycle (Homo sapiens) - WikiPathways">The pathway can be edited at {{cite web | url = http://www.wikipathways.org/index.php/Pathway:WP78 | title = WikiPathways | author = Alexander Pico, Thomas Kelder, Martijn van Iersel, Kristina Hanspers, Kdahlquist, Nick Fidelman | date = | work = | publisher = | pages = | accessdate = }}</ref>
- Based on
this discussion, I have removed the link to
Gene Wiki.
Boghog (
talk) 15:30, 4 September 2010 (UTC)
- I've thought about this some more since my comment here yesterday, and it seems to me that the two paramount issues, for me, are (1) don't mess up the page layout, and (2) don't add anything that is factually incorrect. (Obviously, I'm reacting in terms of the addition to the dopamine page, which is what brought this to my watchlist.) It occurs to me that one part of the problem is that the templates were added as new sections within the page, under their own header. That's the wrong way to go about it. If pathways are to be added to existing pages (it occurs to me that a case could, instead, be made for making new pages about individual pathways), they should be treated as other images are, as visual accompaniments to the text. They shouldn't just be plopped down somewhere on a page. They should appear, as thumbnails, at a reasonable size and with an informative image caption. That caption should relate to the page content, not to the other wiki or to the original topic of the template.
- To expand on these concerns, please look at where the template was added, also, to tyrosine and L-DOPA. In both cases, the template is given its own section header, with text at the top that is all about the project and not at all about the subject matter of the page. The caption at the bottom says "Nicotine in Dopaminergic Neurons (edit)". Well, there is no nicotine in dopaminergic neurons. There's no explanation for the reader of what this information has to do with the rest of those pages. Why is it on L-DOPA? That's a page about a molecule that, although it is a precursor to dopamine, is primarily of interest to readers because it is a pharmaceutical used clinically for Parkinson's disease. Why would a reader interested in Parkinson's medicines want to interact with a biochemical pathway, and what does that have to do with nicotine? And on tyrosine, it treats tyrosine as a precursor to dopamine which is a mediator of nicotine effects, which, by itself, violates WP:UNDUE. Tyrosine is a dopamine precursor, but what about other catecholamines? What about other effects of dopamine besides those related to nicotine? Most importantly, what about all the other pathways in which tyrosine plays important roles? If these kinds of things hadn't set off red flags for me, it would be just a matter of time before other editors would have objected. -- Tryptofish ( talk) 20:37, 4 September 2010 (UTC)
- Tryptofish makes some excellent points. I think the interactive pathways are appropriate to include in Gene Wiki articles where the primary function of the protein is directly linked to the pathway (e.g., PDHA1 in {{ TCACycle_WP78}}) but would be much less appropriate to include this graphic in articles like acetyl-CoA since this molecule is a part of so many biochemical pathways. In addition, it is appropriate to display the entire {{ TCACycle_WP78}} template in an article like citric acid cycle where there is a a close 1:1 relationship between the contents of the interactive pathway and the subject matter of the article. On the other hand, there does not currently appear to be appropriate "parent" articles for templates like {{ NicotineDopaminergicActivity WP1602}}. Hence I agree with Tryptofish that new Wikipedia articles need to be written specifically for these pathways. Boghog ( talk) 11:36, 5 September 2010 (UTC)
- Another issue that I am now reminded of is
WP:RS. The template and/or parent article needs to adequately document the source(s) that were used to create the interactive pathway.
Boghog (
talk) 15:41, 5 September 2010 (UTC)
- Yes, thank you, those are all very good ways to address my concerns. -- Tryptofish ( talk) 16:31, 5 September 2010 (UTC)
- This sounds perfectly reasonable to me. Thank you for the feedback. I will remove the maps where they are non-specific or adding undue weight. I will also update the template to document the sources used to create the content. AlexanderPico ( talk) 19:31, 5 September 2010 (UTC)
Could this be used to create an interactive claddogram navigation tool for taxoboxes? I mean, could there be in a taxobox a picture of the position of the branch of the tree of life of the refferent? And then could the user click on the claddogram and zoom in and out and click on taxa and be sent to articles? Have you seen how they do it at the hall of vertibrates at the American Museum of Natural History in Manhattan? There's a big claddogram at the front and then each hall is laid out to follow the clades, and a cladogram is there at each exhibit so you can see where the animal fits onto the tree of life? Could we do something like that using this tool? I hope I've said enough for you to understand the kind of thing I'm driving at, but if not but if I've piqued your interest and want me to describe this idea in more detail, please ask and I'll try to flesh it out more, but for the moment I don't want to go on too long. Chrisrus ( talk) 04:50, 22 October 2010 (UTC)
- I take it the particle "this" refers to the interactive pathway maps? yes, a
dendrogram/
cladogram is a particular type of
node graph with large limitations to the possible connections and in Matlab one can actually jury-rig (badly) graph theory for NJ nets and such when the standard (polytomy-hating) dendrogram in the Bioinf. toolbox is no good, but overall the conventional representation of dendrograms is very different from node maps and even more so pathway maps, also a modified node graph, which means that it may be best making a separate tool for the tree of life, but, I am sorry to say, I don't think it would be too successful though: currently there is a
Template:Cladogram in wikipedia, which is straightforward and is barely used as most phylogenies are debated (e.g.
malacostraca or the
root of the tree of life), bar for stuff closer to humans that is, there is taxonomy information which can be used uncurated (if you are happy to see a polytomy of 16+ bacterial divisions that is). --
Squidonius (
talk) 08:26, 22 October 2010 (UTC)
- If you visit the fouth floor of the American Museum of Natural History, you are greeted by a huge screen tree of life, which basically explains how it works and then focuses on the branch that is dealt with on the floor, that of the vertibrates. Then, each exhibit reproduces the same branch in more detail. It's really fabulous in terms of presenting thngs, as opposed to the traditional way, as on the other floors, by geographic area "Animals of Africa", or by era "dinosaurs of the Jerassic" or by a taxon or word like "Birds". While it may not be perfect, according to the article cladistics, it's more accurate then taxonomy, which (while indispensable) also features many areas that are debated or complicated, and is included in every article nevertheless. You will agree that the situation about every type of living thing should be presented in as complicated a way as it needs to be and no more. And we presently often settle on such tree diagrams in articles attempting to present such complicated situations, and tend to omit them more when things are relatively straightforward. So that makes me think that the fact that things are complicated or debated would argue for, rather than against, such an inclusion. Chrisrus ( talk) 13:03, 22 October 2010 (UTC)
Possible relation of gap-43 with autism disorders
I would like to know if we could include a possible relation of protein gap-43 with autism, as stated on this scientific article:
http://www.jneurosci.org/cgi/content/abstract/30/44/14595
And this:
http://blog.autismspeaks.org/2010/11/11/what-lies-beneath-brain-connections/#comment-9286
The page I think should be modified is this:
http://en.wikipedia.org/wiki/Gap-43_protein
- I don't see any problem with adding the first citation to the Gap-43_protein#Clinical_significance section since it is to a peer reviewed article and very relevant to the subject matter of the Wikipedia article. However the second link is a blog and blogs are often not considered a reliable sources. But this particular blog link since it sumarizes the peer reviewed article could be added as a "lay summary" combined with the first citation:
- * Zikopoulos B, Barbas H (November 2010). "Changes in prefrontal axons may disrupt the network in autism". J. Neurosci. 30 (44): 14595–609.
doi:
10.1523/JNEUROSCI.2257-10.2010.
PMID
21048117.
{{ cite journal}}: Unknown parameter|laysource=ignored ( help); Unknown parameter|laysummary=ignored ( help) - Finally please note that it is not necessary to ask the community about adding material to a Wikipedia article, unless you think it might be controversial (if so, it is appropriate to bring up the issue directly on the article's talk page). I don't see anything that is particularly controversial about the material that you propose to add, so be bold! Boghog ( talk) 20:46, 12 November 2010 (UTC)
Infobox biodatabase
Hi all, the latest Database Issue of NAR has just been published and I wonder if it would be worth to create an infobox biodatabase. For each database I would write an article with this infobox. The fields would be:
- center
- author(s)
- date ...
- specific organism(s)
- URL
- Web Services ( SOAP, WSDL URLs etc...)
- public Mysql access
- etc...
I plan to use a XSLT stylesheet to quiclky write a WP article from a pubmed abstract. -- Plindenbaum ( talk) 14:59, 5 January 2011 (UTC)
- i think that's a great idea, maybe even link it to Biological_databases. I'd be happy to go through them afterwards to organize them into some kind of grouped table to people can quickly see what is available. It might also be worthwhile to ping an email to some of the corresponding authors to let them know, and to invite them to keep their database page up to date. Davebridges ( talk)
- Good idea. You can find many existing DB articles in Category:Biological databases. -- Paul ( talk) 16:34, 5 January 2011 (UTC)
OK, I started here: Template:Infobox_biodatabase-- Plindenbaum ( talk) 18:17, 5 January 2011 (UTC)
- Thanks for staring this. Perhaps I am biased, but I think the current template places too much emphasis on computer science (i.e., the format) and not enough on biology (i.e., the content). Also I question the importance of including information on the release version. This will be difficult to maintain and is of questionable value to most readers. If someone is really interested about the version number of the latest release, they will check the website directly. Boghog ( talk) 18:33, 5 January 2011 (UTC)
- Suggested additions to template:
- Name
- Description
- Category (Nucleotide Sequence, RNA sequence, Protein sequence, Structure, Genomics (non-vertebrate), Metabolic and Signaling Pathways, Human and other Vertebrate Genomes, Human Genes and Diseases, Microarray Data and other Gene Expression, Proteomics Resources, Other Molecular Biology, Organelle, Plant, Immunological)
- Boghog ( talk) 18:53, 5 January 2011 (UTC)
- Copying the software template was just for a start, I modified the template, added a 'doc page' and a sandbox page. But feel free to edit this infobox :-)-- Plindenbaum ( talk) 19:52, 5 January 2011 (UTC)
- Suggested additions to template:
This is a timely suggestion. Recently a paper has been published about something called BioDBCore. This initiative aims to define a set of minimal information requirements about a database. In fact it is very much like what we would want for an infobox. BioDBCore currently define 17 different fields of information, including the resources Wikipedia entry! It would make sense that as far as possible we try to mirror that set up. If it does become a widely adopted standard it will make it easier for us to keep these boxes up to date in the future. The paper includes an excel file with some examples from some well known databases.
The paper is here: http://www.ncbi.nlm.nih.gov/pubmed/21097465 The field description is in the excel file in the supp. materials: http://nar.oxfordjournals.org/content/early/2010/11/17/nar.gkq1173/suppl/DC1
Here is the list of their fields:
- Database name
- Main resource URL
- Contact information
- Date resource established (year)
- Conditions of use
- Scope:
- Data types captured
- Curation policy
- Standards: MIs, Data formats, Terminologies
- Data formats
- Taxonomic coverage (use NCBI Taxid)
- Data accessibility/output options
- Data release frequency
- Versioning policy/ access to historical files
- Documentation available
- User support options
- Data submission policy
- Relevant publications
- Resource’s Wikipedia URL
- Tools available
Thanks for starting this! Alexbateman ( talk) 08:28, 6 January 2011 (UTC)
- Thanks for your comment Alex, I've added a few fields according to your suggestions and I've updated the documentation.--
Plindenbaum (
talk) 09:44, 6 January 2011 (UTC)
-
Looks good!
Boghog (
talk) 11:51, 6 January 2011 (UTC)
- In my opinion the description field will be redundant with the lead of the Wikipedia article and should be deleted. I don't see a description field being used for other infoboxes such as taxoboxes, protein family boxes etc. Although sometimes there is a figure caption used.
193.62.202.241 (
talk) 12:41, 6 January 2011 (UTC)
- I imagine, 'description' should be used for as a title, not for a complete description: eg:
- In my opinion the description field will be redundant with the lead of the Wikipedia article and should be deleted. I don't see a description field being used for other infoboxes such as taxoboxes, protein family boxes etc. Although sometimes there is a figure caption used.
193.62.202.241 (
talk) 12:41, 6 January 2011 (UTC)
-
Looks good!
Boghog (
talk) 11:51, 6 January 2011 (UTC)
| Content | |
|---|---|
| Description | Comparative genomics of Fusarium strains, |
| Contact | |
| Primary citation | PMID 21087991 |
| Access | |
| Website | http://www.fusarium.org |
-- Plindenbaum ( talk) 13:10, 6 January 2011 (UTC)
- I don't see the description as being necessarily redundant with the lead, but rather a very brief summary of what is contained in the database (see for example the GenBank article). The lead often runs several paragraphs and is not consistent between articles. The description in the infobox is more structured and allow the reader at a glance to figure out what is contained in the database. Boghog ( talk) 13:37, 6 January 2011 (UTC)
I wrote a Xslt stylesheet to transform a Pubmed article to wikipedia. See Template:Infobox_biodatabase/doc for a demo. And here are a two pages I created with this stylesheet : PlasmoDB and Rebase. I also created the template Template:Biodatabase-stub -- Plindenbaum ( talk) 21:19, 6 January 2011 (UTC)
- A while back I created a page for
cancer research databases. But if you now create a WP page for each database with an infobox it will be kind of redundant. So is it better to maintain both or move all to the same schema? I personally think that this kind of organization gives a better overview at one glance and is easier to maintain.
Michipanero (
talk) 09:12, 8 April 2011 (UTC)
- @ Michipanero : my goal is to be able to search for a databases using a SPARQL query (once the database will be digested by DBPedia). That's why I need this kind of infobox. -- Plindenbaum ( talk) 11:49, 13 April 2011 (UTC)
- A while back I created a page for
cancer research databases. But if you now create a WP page for each database with an infobox it will be kind of redundant. So is it better to maintain both or move all to the same schema? I personally think that this kind of organization gives a better overview at one glance and is easier to maintain.
Michipanero (
talk) 09:12, 8 April 2011 (UTC)
FYI the data from Template:Infobox_biodatabase have now been integrated into DBPedia 3.7. See:
- http://en.wikipedia.org/wiki/Wikipedia:WikiProject_Molecular_and_Cellular_Biology/Announcements#Infobox_biodatabase (for reference)
- http://dbpedia.org/page/Template:Infobox_biodatabase
- http://dbpedia.org/ontology/BiologicalDatabase
- http://dbpedia.org/page/L1Base
those data can now be searched through a SPARQL query -- Plindenbaum ( talk) 09:56, 11 September 2011 (UTC)
Ribosyl group
I believe it would be informative as well as useful to create a stub on the ribosyl group. I submitted an article for review, albeit, more of a definition, and it was promptly rejected. So I concluded that joining the project and hearing some opinions on whether it is important enough to have its own article, or if the ribose article could be edited to include its radical and its subsequent presence in proteins and enzymes. Let me know what you think. Dissonase 02:12, 17 February 2011 (UTC)
- Unless the proposed ribosyl article were significantly expanded, I think it would be better to combine it with the existing ribose article and create redirects for ribosyl, ribosyl group, ribosyl radical, etc. pointing to ribose. The ribose article already mentions that ribose is a component of RNA. This could be expanded to mention that the ribosyl functional group can also be attached to other macromolecules. Please be sure to include reliable sources to document that ribose is a component of each of the macromolecules mentioned. Cheers. Boghog ( talk) 06:44, 17 February 2011 (UTC)
Comments on Merging Invariant Chain with CD74
Discuss. Blahdenoma ( talk) 05:59, 30 April 2011 (UTC)
 Done I agree with the merger proposal. I therefore boldly went ahead and completed the merger.
Boghog (
talk) 08:07, 30 April 2011 (UTC)
Done I agree with the merger proposal. I therefore boldly went ahead and completed the merger.
Boghog (
talk) 08:07, 30 April 2011 (UTC)
| Mouse Mutant Alleles for Art4 | ||
|---|---|---|
| Marker Symbol for Mouse Gene. This symbol is assigned to the genomic locus by the MGI | Art4 | |
| Mutant Mouse Embryonic Stem Cell Clones. These are the known targeted mutations for this gene in a mouse. | Art4tm1aWTSI(KOMP) | |
| Example structure of targeted conditional mutant allele for this gene | ||
 | ||
| These Mutant ES Cells can be studied directly or used to generate mice with this gene knocked out. Study of these mice can shed light on the function of Art4 see Knockout mouse | ||
I envision the mutabox updates to the gene pages will be made by a robot ('mutabot', still to be proposed & written) which scans the known high-throughput mutations at www.knockoutmouse.org and applies the changes to a template in each mammalian gene entry. Also note that all mutabox entries would point back to the existing wiki page Knockout mouse], which explains further why these mutants are scientifically interesting (they help the study of gene function), and gives more links.
I would like to get opinions (and hopefully consensus) from the MCB group for this proposal.
Regards,
Vivek Iyer
High Throughput Gene Targeting
Wellcome Trust Sanger Institute
- Hi Vivek, On the surface this looks like it could be a useful resource for experimental biologists. But a concern could be that this effort could be construed as advertising for a commercial entity - the links to 'order vector' and 'order ES cells' set off warning bells. I've had a quick look around the Wikipedia policy pages and as usual I haven't found too much of use. Possibly WP:LINKS and conflict-of-interest should be checked against what you want to do.-- Paul ( talk) 09:52, 24 February 2009 (UTC)
- Hi Paul, Thanks for the observation and the references. Regarding the concern with advertising a commercial enterprise: I should have emphasised that these consortia are NOT commercial enterprises - they are funded by the NIH and the EU (and the Wellcome Trust) to serve the international research community. The 'order' buttons you mentioned put end-users quickly in touch with the vector & ES Cell distribution units, which are also run as part of academic institutes - UC Davis, CHORI in Oakland and the HMGU in Munich. Those units do function on a cost-recovery basis, so there is a charge for products, but that mode of working has been established for a long time. Anyway, I've modified the text of the proposal to say this explicitly. As for the conflict of interest, I hope I've made my affiliation to the program informatics clear, but let me know if I've missed something. Does this allay your concerns? -- Vivek 11.35, 24 February 2009 (UTC)
- I've now made the proposed edit directly onto http://en.wikipedia.org/wiki/ART4 . An proposal for a BOT is in progress. Vivek ( talk) 11.35, 19 March 2009 (UTC)
- Vivek, the prototypes look good to me... You may get more specific feedback from the bot approval group, but you may want to convert the mutabox to wikitext tables (see Help:Tables) instead of HTML. Cheers, AndrewGNF ( talk) 04:42, 14 April 2009 (UTC)
- Hi Vivek, apologies I have come rather late to this discussion. I have some concerns about your proposed boxes, mainly to do with their size and whether they give undue prominence to information that is not so important to the likely users of wikipedia. Some points:
- whilst some gene pages are probably only of interest to those of us who are researchers, some e.g. perhaps huntingtin, p53 etc. are probably of interest to undergraduates and non-scientists. I think a large box on a research tool that will dominate many short pages is inappropriate in these circumstances.
- the schematic showing your construct. Forgive me, but once you have seen one, you have seen them all essentially - they all follow the same strategy? I don't really think that they are useful. I would prefer them to be removed or initially hidden with a 'show' control.
- the boxes are very big. Would it not be preferable to add the information instead to the gene box in a short form e.g. "Mouse knockout available" "ES cell mutant available" with a link to your database here?
- (minor practical concern) in Safari 4 on my computer the mutabox display overlaps with the gene box breaking the page display.
- Best wishes, Celefin ( talk) 19:34, 17 April 2009 (UTC)
- Hi Vivek, apologies I have come rather late to this discussion. I have some concerns about your proposed boxes, mainly to do with their size and whether they give undue prominence to information that is not so important to the likely users of wikipedia. Some points:
- Hi Celefin, thankyou for the feedback, this is just the kind of thing I need (difficult to pick up from the 'inside'). I think the best response is a reworking of the mutabox examples to address both your and AndrewGNF's comments, but here are some quick thoughts
- The box is probably too big, and can be shrunk down somewhat, and changed from an html table into a wiki table. I haven't experimented to see if it will help the page display problem you mention, but I noticed the layout on my browser (also safari) was not happy when the browser window got too small - I am sure there is a good way of dealing with that without stuffing up the page or dominating it.
- Those pictures are all different, reflecting both gene structure and the exact mutagenic cassette, which can follow quite different strategies. The pipelines have been rejigging themselves a lot, and the changes matter. I think the existence of a picture does actually draw a readers attention better than plain text. A molecular biologist will interpret them, but I had in mind a non-specialist could at least get drawn by that into the text, which I have made deliberately very simple and general.
- I disagree that information about mutations shouldn't swamp a short page. First, I can & will rework the mutabox so it doesn't swamp anything, but If there are pages which are short (containing only a brief description and references), then shouldn't they be fleshed out with other relevant information? If that information is the existence of a mutant allele, isn't that a good thing? I was thinking that pages which are essentially stubs right now would / grow, right?
- Thanks again for the comments! Vivek ( talk) 21:45, 20 April 2009 (UTC)
- Hi Celefin, thankyou for the feedback, this is just the kind of thing I need (difficult to pick up from the 'inside'). I think the best response is a reworking of the mutabox examples to address both your and AndrewGNF's comments, but here are some quick thoughts
IRC channel to recruit people and as a place to hang out...
I'd propose a channel on freenode.net (just like #wikipedia has)... my suggestion would be the following channel irc://irc.freenode.net/#wikipedia-mcb
There are a bunch of people in irc://irc.freenode.net/#bioinformatics for example... I'm sure you could recruit some of them if you had an irc presence... also I'd just quite like to chat to some of you guys some time ;-)
-- Dan| (talk) 18:35, 14 April 2009 (UTC)
Merge Wikipedia:WikiProject Cell Signaling with this project as a task force?
I made a suggestion over at the cell signaling wikiproject (see here [1]) to consider merging into this project as a task force. I noticed above there has been suggestions to make a few task forces but not sure whether they went ahead as planned? Anyone here happen to have any suggestions/comments they would like to make re: merging cell signalling wikiproject as a task force here? Cheers. Calaka ( talk) 11:13, 5 May 2009 (UTC)
Photosynthesis Taskforce
I think a photosynthesis taskforce should be a great idea. A couple of months ago I tried to make a wikiproject about this subject, but a taskforce should be better. At the moment the most important articles about this subject (Photosynthesis, Light Reactions, Dark reactions, Chloroplast) are not anything near FA-status. (Altough, I have put a lot time in the light reactions article). And all the other articles of this subject are mostly stubs.
I think the photosynthesis article itself is one of the examples of what happens if you have to many authors. One of the three guys that have ever posted something on the wikiproject photosynthesis wrote this about it:
"In my opinion, this article is one of the worst on WikiPedia. There is a heck of a lot of good information on this page, but it is obtuse, chaotic and so badly written that is absolutely useless. The only people who can understand it are the people who already know it; a solid case of preaching to the choir, if I've ever seen one."
And I think he is totally right...
So what do you guys think? Kasper90 ( talk) 16:42, 26 May 2009 (UTC)
- I don't see why not. Will you be willing to set up the task force page up though and add in the required info/introduction/guidelines? Might be a bit of work. In my case (for the eventual merger of the sister projects onto here), I pretty much got the template, just need to do a few tweaks here and there. Test out the look/feel of it on a sandbox perhaps? Calaka ( talk) 10:54, 30 May 2009 (UTC)