| Mathematics desk | ||
|---|---|---|
| < December 2 | << Nov | December | Jan >> | December 4 > |
| Welcome to the Wikipedia Mathematics Reference Desk Archives |
|---|
| The page you are currently viewing is an archive page. While you can leave answers for any questions shown below, please ask new questions on one of the current reference desk pages. |
December 3 Information
Kind of an esoteric question; I'm modeling something rare, about 1000 cases in about a million population. (SAS, proc genmod, poisson distribution). Intending to do the more or less routine thing; model sample of 1/3, run that model on second third to check for overfitting, use final third to validate/score. Works fine, but.... i sampled cases and controls separately then combined each (i.e. 1/3 of cases + 1/3 of controls), rather than taking 1/3 sample of total controls+cases together, because I was worried that 1/3 of the whole thing might get a small number of cases. Anyway, practicality aside, I'd like to hear any arguments re the theoretical validity or lack of same of the aforementioned separate sampling of controls and cases and combining? Thanks. Gzuckier ( talk) 04:46, 3 December 2012 (UTC)
- This is basically stratified sampling, and is valid (and efficient) for the same reasons that it is. A full theoretical justification would be tricky, but the essence of it is that the "building blocks" of any learning algorithm would be values of where A is a set in feature space and Y is the truth value of the case. The expected empirical value of this is equal to the proportion in the complete sample in both sampling methods; and the variance is lower with stratification. -- Meni Rosenfeld ( talk) 11:52, 3 December 2012 (UTC)
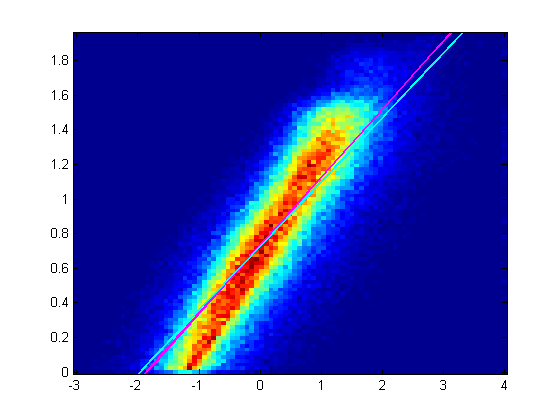
The following is a heat map 2D histogram of X,Y values. It's a linear heat map. Magenta is the bisquare robust fit, cyan is the least squares fit. The robust fit makes the previous least squares fit only slightly better. How do I make it fit the trend? The plot is a log-log plot comparing log velocity (on the Y axis) against log of curvature (on the X-axis), where the data apparently follows a power law. However, using linear regression seems to obtain the wrong slope. John Riemann Soong ( talk) 05:18, 3 December 2012 (UTC)
I've also read on the internet the maximum likelihood estimator might be better than linear regression for power laws. However, this seems to be based on X-frequency rather than X-Y (velocity-curvature). Is there a way to use maximum likelihood estimators between two variables? Why are the discussions biased to frequency? John Riemann Soong ( talk) 05:36, 3 December 2012 (UTC)
- Something is wrong, even the least squares fit should be better than that. Check your data. StuRat ( talk) 06:09, 3 December 2012 (UTC)
- I agree with StuRat that simple least squares should do better than that. Given that this is a histogram, what exactly are you using as input to your least squares fit? Traditionally least squares would require a set of (X, Y) pairs, with this histogram every pair has some value. You would have to use the histogram's value at each (X, Y) pair as your weight in a weigted least square, but then if the values at the sides (blue areas) aren't exactly zero then you would introduce a bias by the rectangle shape of the region you're fitting on, which clearly seems to be happening. 41.164.7.242 ( talk) 09:52, 3 December 2012 (UTC) Eon
- I haven't uploaded the logarithmic histogram (where frequency / colormap is on a logarithmic scale) but there is a larger spread of values below the line than above the line, so rather than going through what would be the "median distribution" it goes through the average. How do I make it go through the median? John Riemann Soong ( talk) 06:33, 3 December 2012 (UTC)
- The problem seems to be with the slope. Given that slope, this might be about as good of a fit as can be expected. However, that's clearly not the correct slope. StuRat ( talk) 08:11, 3 December 2012 (UTC)
- Can you give a legend with the frequency represented by each color?
- If you want a "median" line (which could help if you have outliers), you want the sum of absolute (not squared) distances from the line to be minimal. I don't know of a standard technique, but you could try this: Pick some angle. Rotate all data by this angle. Find the median of y-values (using x-values is equivalent with a different angle), and the sum of absolute differences of y-values from this median. This gives you the loss function for this angle. Try this for different angles an use some numerical optimization technique for the best angle. Rotate the line y=median back, and you get your optimal line. (Everything should be weighted by frequency, of course). --
Meni Rosenfeld (
talk) 10:30, 3 December 2012 (UTC)
- See
Least absolute deviations.
Duoduoduo (
talk) 13:57, 3 December 2012 (UTC)
- Right. One difference between this and what I discussed is something the OP alluded to - it focuses on finding y as a function of x rather than treating x and y symmetrically. This makes it biased against vertical lines (which can be fixed by multiplying the loss by the cosine of the angle). -- Meni Rosenfeld ( talk) 14:14, 3 December 2012 (UTC)
- See
Least absolute deviations.
Duoduoduo (
talk) 13:57, 3 December 2012 (UTC)
- Deming regression treats x and y symmetrically if is set equal to 1. It minimizes the sum of squared perpendicular residuals, but I would imagine that one could recast the objective function in terms of perpendicular absolute deviations and then use one of the solution techniques discussed in Least absolute deviations. Duoduoduo ( talk) 14:42, 3 December 2012 (UTC)
- This is absurd, but needs saying; you could actually provide a better fit, by drawing a line through the center of the point cloud and figuring out the equation of the line. Gzuckier ( talk) 18:30, 3 December 2012 (UTC)
- I think maybe what Gzuckier has in mind is that, when you look at the graph above, the least squares line and the other fitted line "can't be right" because the fitted lines don't have the same slope as the heat map of the data. But those fitted lines are optimal according to their own criteria. My guess from looking at the graph is that if you do a plot of
x or ofy horizontally and of the corresponding residuals vertically, you find thatthere is serial correlation of the residualsthe residuals are poorly behaved -- that is, higherx ory values are associated with numerically higher values of the residuals -- higher values ofx ory tend to be more likely to have positive residuals (= actual y minus predicted y). That's always a hint that the regression has been misspecified. To figure out what is misspecified about it, you have to examine the theory that says you should be running this regression in the first place, and see if a more appropriate variant of the theory suggests a different specification of the regression -- one that might have a more random mapping of the regression residuals. Duoduoduo ( talk) 21:53, 3 December 2012 (UTC) - But I agree with 41.164.7.242 and StuRat above -- while the heat map could be misleading to the eye about where there are more points than elsewhere, it seems to me that the least squares regression line and probably the other one have been plotted wrong, or maybe the heat map has been plotted wrong, or maybe there was a coding error in running the regressions. Duoduoduo ( talk) 00:06, 4 December 2012 (UTC)
- I think maybe what Gzuckier has in mind is that, when you look at the graph above, the least squares line and the other fitted line "can't be right" because the fitted lines don't have the same slope as the heat map of the data. But those fitted lines are optimal according to their own criteria. My guess from looking at the graph is that if you do a plot of


- What Gzuckier is describing is exactly total least squares. You find the mean, subtract it, then take the outer product of all of the vectors with themselves to get a 2x2 matrix, take the SVD of that matrix to find the slope of the line. Proving it is pretty involved, but doing it is pretty easy. —Ben FrantzDale ( talk) 00:43, 4 December 2012 (UTC)
- Well, as StuRat pointed out, Gzuckier said we should "draw a line" through the point cloud, which sounds like hand-drawing and isn't the same thing as doing it mathematically. In any event, total least squares in this simple context is Deming regression, which I linked to. Duoduoduo ( talk) 14:00, 4 December 2012 (UTC)
- This is a common problem with linear regression -- the ordinary formula essentially assumes that the X values are perfectly accurate and all the error is in the Y values. For this particular case, based on my personal experience, you would get much better results regressing X as a function of Y rather than the other way around -- but even better is to use Deming regression, as Duoduoduo suggested. Looie496 ( talk) 20:15, 3 December 2012 (UTC)
- Are you fitting a line to a log x + b = log y, or are you doing y = a x + b? That said, I agree with other posters: It doesn't look like it's not robust, it looks like it is wrong. If it were an issue of robustness, you would see bright red dots pulling the regression lines away from the "backbone" of your histogram. —Ben FrantzDale ( talk) 00:34, 4 December 2012 (UTC)
Newer (better) slopes and intercepts, but would still like to optimize further
I really can't draw the slopes myself. I have to measure the slopes between genotypes, and measure the differences. The background to this is that I have to distinguish between genotypes, which I have found to have statistically significantly different slopes. The separation is good, at least decent. I was trying to achieve an even better separation and see if the effect size was even greater by having the regression line actually fit the data. The data is log y = b log x + log K (to fit the power law y = K*x^beta). I modified the robust regression parameters to have a really low tuning constant (1.2) for the bisquare weighting algorithm. Here are some new histograms. (Not really important but I just realized the x-axis labels are wrong, R should be in mm, not rad/mm.)
By lowering the tuning constant, the slopes (and intercepts) found are slightly better, but they are still not optimally fit. The problem is that there seems to be a point where lowering the tuning constant doesn't make any difference, because robustfit() hits the iteration limit if I set the tuning constant below a certain point. If I set the iteration limit to 150 (changing it from 50), I still get roughly the same slopes. Which is fine, it still returns me valid slopes, but it appears that there is a limit to how much the slopes can be corrected.
John Riemann Soong (
talk) 01:27, 4 December 2012 (UTC)
You can see from the logarithmic histogram a better clearer picture about where the errors might be coming from. I did notice initially that the line seemed to fit the logarithmic histogram (where the frequency z is scaled logarithmically) whereas the linear scale hides the low frequency (but seemingly high-influencce) values. Fumin has this interesting subdistribution which is why I saved the image for my lab presentation this morning, the other genotypes doesn't. They all have a "long tail" (frequencywise) in the (x,y) coordinates "below" the line though (the tail falloff trend runs almost "normal" or "orthogonal" to the main trendline).
I think this is why robust regression might be having a hard time even with ridiculously low tuning constants because the slow dropoff of the long tail (away from the "median" or "mode") make them not seen as outliers so their their r (error/distance with respect to the last iteration) is still seen as low. How do I obtain a "median" line fit in MATLAB?
Specific to fumin: I note that the straight robust line in the "fumin" histogram goes through the main distribution, which is good, but is still somewhat influenced by the "smaller" subdistribution (the local frequency maximum in that region is unique to fumin), and this lowers its slope somewhat to lower than the other genotypes. Individually, each of the fumin flies' regressions have higher slopes than the other genotypes, within at least a 95% degree of confidence, if not more. John Riemann Soong ( talk) 01:35, 4 December 2012 (UTC)
- what's your model/mechanism? as a naive observer, it looks to me more like the fumin subpop you mention could be described as a distinct subpop/distribution with a similar slope to the main pop but a lower intercept, rather than a continuous tail to the main distribution. is that physically/realistically possible? Gzuckier ( talk) 17:26, 4 December 2012 (UTC)
- It sounds to me like the OP is searching through numerous specifications in order to pick the one that best confirms his desired result (in this case, the the different genotypes have markedly different slopes). This is known as "torturing the data until it confesses", and is considered bad form and potentially even unprofessional if only the "best" results are reported. With any data set, one can fiddle around with the technique of analysis until some approach rejects the null and confirms the pre-chosen result; if you run, say, 20 different specifications at the 95% confidence level (alpha = .05), then typically you'll find one specification that rejects the null (one out of 20, or 5% of the specifications) even when the null is true. (In this case the null is no difference among the phenotypes.)
- In order to find out whether this is what you have done, or whether significant results really do reflect an actual departure of reality from the null, you should split the sample for each genotype in, say, half; run your procedures on half the data, get the results, and then see how well your regression results predict for the other half of the data -- the half that was not used in getting the results. This is known as checking the out-of-sample fit and is a vital part of empirical work. Duoduoduo ( talk) 14:52, 5 December 2012 (UTC)
- My problem is not significance levels, it is effect size. I have significance (essentially) no matter how I analyze the data, I simply want to get linearity. I think nonlinearity is masking the true effect size. Also my personal standard for alpha is 0.005, not 0.05.
71.207.151.227 (
talk) 21:42, 5 December 2012 (UTC)
- See
Box–Cox transformation.
Duoduoduo (
talk) 12:42, 6 December 2012 (UTC)
- Or: From your latest graph, it looks like maybe you could split the data points into those for which x<k and those for which x≥k, for some k chosen by looking at the graph, and then run a linear regression separately on each of the two resulting data sets. Then you could do an F-test on those regressions to test the null hypothesis that splitting the data sets gives no significant improvement to the fit. Or, judging from the graph maybe the individual slopes of the two data sets are identical, but only the intercepts are different. Then maybe you should run a single regression on all the data but including as a regressor a dummy variable which =0 for x<k and =1 for x≥k. Duoduoduo ( talk) 17:04, 6 December 2012 (UTC)
- See
Box–Cox transformation.
Duoduoduo (
talk) 12:42, 6 December 2012 (UTC)
- My problem is not significance levels, it is effect size. I have significance (essentially) no matter how I analyze the data, I simply want to get linearity. I think nonlinearity is masking the true effect size. Also my personal standard for alpha is 0.005, not 0.05.
71.207.151.227 (
talk) 21:42, 5 December 2012 (UTC)
| Mathematics desk | ||
|---|---|---|
| < December 2 | << Nov | December | Jan >> | December 4 > |
| Welcome to the Wikipedia Mathematics Reference Desk Archives |
|---|
| The page you are currently viewing is an archive page. While you can leave answers for any questions shown below, please ask new questions on one of the current reference desk pages. |
December 3 Information
Kind of an esoteric question; I'm modeling something rare, about 1000 cases in about a million population. (SAS, proc genmod, poisson distribution). Intending to do the more or less routine thing; model sample of 1/3, run that model on second third to check for overfitting, use final third to validate/score. Works fine, but.... i sampled cases and controls separately then combined each (i.e. 1/3 of cases + 1/3 of controls), rather than taking 1/3 sample of total controls+cases together, because I was worried that 1/3 of the whole thing might get a small number of cases. Anyway, practicality aside, I'd like to hear any arguments re the theoretical validity or lack of same of the aforementioned separate sampling of controls and cases and combining? Thanks. Gzuckier ( talk) 04:46, 3 December 2012 (UTC)
- This is basically stratified sampling, and is valid (and efficient) for the same reasons that it is. A full theoretical justification would be tricky, but the essence of it is that the "building blocks" of any learning algorithm would be values of where A is a set in feature space and Y is the truth value of the case. The expected empirical value of this is equal to the proportion in the complete sample in both sampling methods; and the variance is lower with stratification. -- Meni Rosenfeld ( talk) 11:52, 3 December 2012 (UTC)
The following is a heat map 2D histogram of X,Y values. It's a linear heat map. Magenta is the bisquare robust fit, cyan is the least squares fit. The robust fit makes the previous least squares fit only slightly better. How do I make it fit the trend? The plot is a log-log plot comparing log velocity (on the Y axis) against log of curvature (on the X-axis), where the data apparently follows a power law. However, using linear regression seems to obtain the wrong slope. John Riemann Soong ( talk) 05:18, 3 December 2012 (UTC)
I've also read on the internet the maximum likelihood estimator might be better than linear regression for power laws. However, this seems to be based on X-frequency rather than X-Y (velocity-curvature). Is there a way to use maximum likelihood estimators between two variables? Why are the discussions biased to frequency? John Riemann Soong ( talk) 05:36, 3 December 2012 (UTC)
- Something is wrong, even the least squares fit should be better than that. Check your data. StuRat ( talk) 06:09, 3 December 2012 (UTC)
- I agree with StuRat that simple least squares should do better than that. Given that this is a histogram, what exactly are you using as input to your least squares fit? Traditionally least squares would require a set of (X, Y) pairs, with this histogram every pair has some value. You would have to use the histogram's value at each (X, Y) pair as your weight in a weigted least square, but then if the values at the sides (blue areas) aren't exactly zero then you would introduce a bias by the rectangle shape of the region you're fitting on, which clearly seems to be happening. 41.164.7.242 ( talk) 09:52, 3 December 2012 (UTC) Eon
- I haven't uploaded the logarithmic histogram (where frequency / colormap is on a logarithmic scale) but there is a larger spread of values below the line than above the line, so rather than going through what would be the "median distribution" it goes through the average. How do I make it go through the median? John Riemann Soong ( talk) 06:33, 3 December 2012 (UTC)
- The problem seems to be with the slope. Given that slope, this might be about as good of a fit as can be expected. However, that's clearly not the correct slope. StuRat ( talk) 08:11, 3 December 2012 (UTC)
- Can you give a legend with the frequency represented by each color?
- If you want a "median" line (which could help if you have outliers), you want the sum of absolute (not squared) distances from the line to be minimal. I don't know of a standard technique, but you could try this: Pick some angle. Rotate all data by this angle. Find the median of y-values (using x-values is equivalent with a different angle), and the sum of absolute differences of y-values from this median. This gives you the loss function for this angle. Try this for different angles an use some numerical optimization technique for the best angle. Rotate the line y=median back, and you get your optimal line. (Everything should be weighted by frequency, of course). --
Meni Rosenfeld (
talk) 10:30, 3 December 2012 (UTC)
- See
Least absolute deviations.
Duoduoduo (
talk) 13:57, 3 December 2012 (UTC)
- Right. One difference between this and what I discussed is something the OP alluded to - it focuses on finding y as a function of x rather than treating x and y symmetrically. This makes it biased against vertical lines (which can be fixed by multiplying the loss by the cosine of the angle). -- Meni Rosenfeld ( talk) 14:14, 3 December 2012 (UTC)
- See
Least absolute deviations.
Duoduoduo (
talk) 13:57, 3 December 2012 (UTC)
- Deming regression treats x and y symmetrically if is set equal to 1. It minimizes the sum of squared perpendicular residuals, but I would imagine that one could recast the objective function in terms of perpendicular absolute deviations and then use one of the solution techniques discussed in Least absolute deviations. Duoduoduo ( talk) 14:42, 3 December 2012 (UTC)
- This is absurd, but needs saying; you could actually provide a better fit, by drawing a line through the center of the point cloud and figuring out the equation of the line. Gzuckier ( talk) 18:30, 3 December 2012 (UTC)
- I think maybe what Gzuckier has in mind is that, when you look at the graph above, the least squares line and the other fitted line "can't be right" because the fitted lines don't have the same slope as the heat map of the data. But those fitted lines are optimal according to their own criteria. My guess from looking at the graph is that if you do a plot of
x or ofy horizontally and of the corresponding residuals vertically, you find thatthere is serial correlation of the residualsthe residuals are poorly behaved -- that is, higherx ory values are associated with numerically higher values of the residuals -- higher values ofx ory tend to be more likely to have positive residuals (= actual y minus predicted y). That's always a hint that the regression has been misspecified. To figure out what is misspecified about it, you have to examine the theory that says you should be running this regression in the first place, and see if a more appropriate variant of the theory suggests a different specification of the regression -- one that might have a more random mapping of the regression residuals. Duoduoduo ( talk) 21:53, 3 December 2012 (UTC) - But I agree with 41.164.7.242 and StuRat above -- while the heat map could be misleading to the eye about where there are more points than elsewhere, it seems to me that the least squares regression line and probably the other one have been plotted wrong, or maybe the heat map has been plotted wrong, or maybe there was a coding error in running the regressions. Duoduoduo ( talk) 00:06, 4 December 2012 (UTC)
- I think maybe what Gzuckier has in mind is that, when you look at the graph above, the least squares line and the other fitted line "can't be right" because the fitted lines don't have the same slope as the heat map of the data. But those fitted lines are optimal according to their own criteria. My guess from looking at the graph is that if you do a plot of


- What Gzuckier is describing is exactly total least squares. You find the mean, subtract it, then take the outer product of all of the vectors with themselves to get a 2x2 matrix, take the SVD of that matrix to find the slope of the line. Proving it is pretty involved, but doing it is pretty easy. —Ben FrantzDale ( talk) 00:43, 4 December 2012 (UTC)
- Well, as StuRat pointed out, Gzuckier said we should "draw a line" through the point cloud, which sounds like hand-drawing and isn't the same thing as doing it mathematically. In any event, total least squares in this simple context is Deming regression, which I linked to. Duoduoduo ( talk) 14:00, 4 December 2012 (UTC)
- This is a common problem with linear regression -- the ordinary formula essentially assumes that the X values are perfectly accurate and all the error is in the Y values. For this particular case, based on my personal experience, you would get much better results regressing X as a function of Y rather than the other way around -- but even better is to use Deming regression, as Duoduoduo suggested. Looie496 ( talk) 20:15, 3 December 2012 (UTC)
- Are you fitting a line to a log x + b = log y, or are you doing y = a x + b? That said, I agree with other posters: It doesn't look like it's not robust, it looks like it is wrong. If it were an issue of robustness, you would see bright red dots pulling the regression lines away from the "backbone" of your histogram. —Ben FrantzDale ( talk) 00:34, 4 December 2012 (UTC)
Newer (better) slopes and intercepts, but would still like to optimize further
I really can't draw the slopes myself. I have to measure the slopes between genotypes, and measure the differences. The background to this is that I have to distinguish between genotypes, which I have found to have statistically significantly different slopes. The separation is good, at least decent. I was trying to achieve an even better separation and see if the effect size was even greater by having the regression line actually fit the data. The data is log y = b log x + log K (to fit the power law y = K*x^beta). I modified the robust regression parameters to have a really low tuning constant (1.2) for the bisquare weighting algorithm. Here are some new histograms. (Not really important but I just realized the x-axis labels are wrong, R should be in mm, not rad/mm.)
By lowering the tuning constant, the slopes (and intercepts) found are slightly better, but they are still not optimally fit. The problem is that there seems to be a point where lowering the tuning constant doesn't make any difference, because robustfit() hits the iteration limit if I set the tuning constant below a certain point. If I set the iteration limit to 150 (changing it from 50), I still get roughly the same slopes. Which is fine, it still returns me valid slopes, but it appears that there is a limit to how much the slopes can be corrected.
John Riemann Soong (
talk) 01:27, 4 December 2012 (UTC)
You can see from the logarithmic histogram a better clearer picture about where the errors might be coming from. I did notice initially that the line seemed to fit the logarithmic histogram (where the frequency z is scaled logarithmically) whereas the linear scale hides the low frequency (but seemingly high-influencce) values. Fumin has this interesting subdistribution which is why I saved the image for my lab presentation this morning, the other genotypes doesn't. They all have a "long tail" (frequencywise) in the (x,y) coordinates "below" the line though (the tail falloff trend runs almost "normal" or "orthogonal" to the main trendline).
I think this is why robust regression might be having a hard time even with ridiculously low tuning constants because the slow dropoff of the long tail (away from the "median" or "mode") make them not seen as outliers so their their r (error/distance with respect to the last iteration) is still seen as low. How do I obtain a "median" line fit in MATLAB?
Specific to fumin: I note that the straight robust line in the "fumin" histogram goes through the main distribution, which is good, but is still somewhat influenced by the "smaller" subdistribution (the local frequency maximum in that region is unique to fumin), and this lowers its slope somewhat to lower than the other genotypes. Individually, each of the fumin flies' regressions have higher slopes than the other genotypes, within at least a 95% degree of confidence, if not more. John Riemann Soong ( talk) 01:35, 4 December 2012 (UTC)
- what's your model/mechanism? as a naive observer, it looks to me more like the fumin subpop you mention could be described as a distinct subpop/distribution with a similar slope to the main pop but a lower intercept, rather than a continuous tail to the main distribution. is that physically/realistically possible? Gzuckier ( talk) 17:26, 4 December 2012 (UTC)
- It sounds to me like the OP is searching through numerous specifications in order to pick the one that best confirms his desired result (in this case, the the different genotypes have markedly different slopes). This is known as "torturing the data until it confesses", and is considered bad form and potentially even unprofessional if only the "best" results are reported. With any data set, one can fiddle around with the technique of analysis until some approach rejects the null and confirms the pre-chosen result; if you run, say, 20 different specifications at the 95% confidence level (alpha = .05), then typically you'll find one specification that rejects the null (one out of 20, or 5% of the specifications) even when the null is true. (In this case the null is no difference among the phenotypes.)
- In order to find out whether this is what you have done, or whether significant results really do reflect an actual departure of reality from the null, you should split the sample for each genotype in, say, half; run your procedures on half the data, get the results, and then see how well your regression results predict for the other half of the data -- the half that was not used in getting the results. This is known as checking the out-of-sample fit and is a vital part of empirical work. Duoduoduo ( talk) 14:52, 5 December 2012 (UTC)
- My problem is not significance levels, it is effect size. I have significance (essentially) no matter how I analyze the data, I simply want to get linearity. I think nonlinearity is masking the true effect size. Also my personal standard for alpha is 0.005, not 0.05.
71.207.151.227 (
talk) 21:42, 5 December 2012 (UTC)
- See
Box–Cox transformation.
Duoduoduo (
talk) 12:42, 6 December 2012 (UTC)
- Or: From your latest graph, it looks like maybe you could split the data points into those for which x<k and those for which x≥k, for some k chosen by looking at the graph, and then run a linear regression separately on each of the two resulting data sets. Then you could do an F-test on those regressions to test the null hypothesis that splitting the data sets gives no significant improvement to the fit. Or, judging from the graph maybe the individual slopes of the two data sets are identical, but only the intercepts are different. Then maybe you should run a single regression on all the data but including as a regressor a dummy variable which =0 for x<k and =1 for x≥k. Duoduoduo ( talk) 17:04, 6 December 2012 (UTC)
- See
Box–Cox transformation.
Duoduoduo (
talk) 12:42, 6 December 2012 (UTC)
- My problem is not significance levels, it is effect size. I have significance (essentially) no matter how I analyze the data, I simply want to get linearity. I think nonlinearity is masking the true effect size. Also my personal standard for alpha is 0.005, not 0.05.
71.207.151.227 (
talk) 21:42, 5 December 2012 (UTC)