| Adenylate kinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
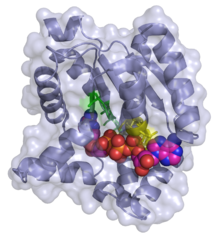 3D ribbon/surface model of adenylate kinase in complex with bis(adenosine)tetraphosphate (ADP-ADP) | |||||||||
| Identifiers | |||||||||
| Symbol | ADK | ||||||||
| Pfam | PF00406 | ||||||||
| InterPro | IPR000850 | ||||||||
| PROSITE | PDOC00104 | ||||||||
| SCOP2 | 1ake / SCOPe / SUPFAM | ||||||||
| |||||||||
Adenylate kinase ( EC 2.7.4.3) (also known as ADK or myokinase) is a phosphotransferase enzyme that catalyzes the interconversion of adenine nucleotides (ATP, ADP, and AMP). By constantly monitoring phosphate nucleotide levels inside the cell, ADK plays an important role in cellular energy homeostasis.
| ADK_lid | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Bacillus stearothermophilus adenylate kinase | |||||||||
| Identifiers | |||||||||
| Symbol | ADK_lid | ||||||||
| Pfam | PF05191 | ||||||||
| InterPro | IPR007862 | ||||||||
| PROSITE | PDOC00104 | ||||||||
| SCOP2 | 1ake / SCOPe / SUPFAM | ||||||||
| |||||||||
| Adenylate kinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 2.7.4.3 | ||||||||
| CAS no. | 2598011 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
The reaction catalyzed is:
The equilibrium constant varies with condition, but it is close to 1. [1] Thus, ΔGo for this reaction is close to zero. In muscle from a variety of species of vertebrates and invertebrates, the concentration of ATP is typically 7-10 times that of ADP, and usually greater than 100 times that of AMP. [2] The rate of oxidative phosphorylation is controlled by the availability of ADP. Thus, the mitochondrion attempts to keep ATP levels high due to the combined action of adenylate kinase and the controls on oxidative phosphorylation.
To date there have been nine human ADK protein isoforms identified. While some of these are ubiquitous throughout the body, some are localized into specific tissues. For example, ADK7 and ADK8 are both only found in the cytosol of cells; and ADK7 is found in skeletal muscle whereas ADK8 is not [3]. Not only do the locations of the various isoforms within the cell vary, but the binding of substrate to the enzyme and kinetics of the phosphoryl transfer are different as well. ADK1, the most abundant cytosolic ADK isozyme, has a Km about a thousand times higher than the Km of ADK7 and 8, indicating a much weaker binding of ADK1 to AMP [4]. Sub-cellular localization of the ADK enzymes is done by including a targeting sequence in the protein [3]. Each isoform also has different preference for NTP's. Some will only use ATP, whereas others will accept GTP, UTP, and CTP as the phosphoryl carrier.
Some of these isoforms prefer other NTP's entirely. There is a mitochondrial GTP:AMP phosphotransferase, also specific for the phosphorylation of AMP, that can only use GTP or ITP as the phosphoryl donor [5]. ADK has also been identified in different bacterial species and in yeast [6]. Two further enzymes are known to be related to the ADK family, i.e. yeast uridine monophosphokinase and slime mold UMP-CMP kinase. Some rfesidues are conserved across these isoforms, indicating how essential they are for catalysis. One of the most conserved areas includes an Arg residue, whose modification inactivates the enzyme, together with an Asp that resides in the catalytic cleft of the enzyme and participates in a salt bridge.
 | This is a user sandbox of
Snodgrah. You can use it for testing or practicing edits. This is not the sandbox where you should draft your assigned article for a dashboard.wikiedu.org course. To find the right sandbox for your assignment, visit your Dashboard course page and follow the Sandbox Draft link for your assigned article in the My Articles section. |
- Adenylate kinase, subfamily InterPro: IPR006259
- UMP-CMP kinase InterPro: IPR006266
- Adenylate kinase, isozyme 1 InterPro: IPR006267
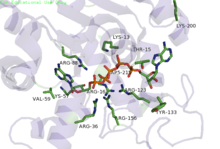
Phosphoryl transfer only occurs on closing of the 'open lid'. This causes an exclusion of water molecules that brings the substrates in proximity to each other [7], lowering the energy barrier for the nucleophilic attack by the γ-phosphoryl group of ATP on the α-phosphoryl of AMP. In the crystal structure of the ADK enzyme from E. Coli with inhibitor Ap5A, the Arg88 residue binds the Ap5A at the α-phosphate group. It has been shown that the mutation R88G results in 99% loss of catalytic activity of this enzyme, suggesting that this residue is intimately involved in the phosphoryl transfer [8]. Another highly conserved residue is Arg119, which lies in the adenosine binding region of the ADK, and acts to sandwich the adenine in the active site. It has been suggested that the promiscuity of these enzymes in accepting other NTP's is due to this relatively inconsequential interactions of the base in the ATP binding pocket [9]. A network of positive, conserved residues (Lys13, Arg123, Arg156, and Arg167 in ADK from E. Coli) stabilize the build up of negative charge on phosphoryl group during the transfer. Two distal aspartate residues bind to the arginine network, causing the enzyme to fold and reduces its flexibility. A magnesium cofactor is also required, essential for increasing the electrophilicity of the phosphate on AMP, though this magnesium ion is only held in the active pocket by electrostatic interactions and dissociates e
asily [9].

Flexibility and plasticity allow proteins to bind to ligands, form oligomers, aggregate, and perform mechanical work [10]. Large conformational changes in proteins play an important role in cellular signaling. Adenylate Kinase is a signal transducing protein; thus, the balance between conformations regulates protein activity. ADK has a locally unfolded state that becomes depopulated upon binding [11].

A 2007 study by Whitford et al. shows the conformations of ADK when binding with ATP or AMP [10]. The study shows that there are three relevant conformations or structures of ADK—CORE, Open, and Closed. In ADK, there are two small domains called the LID and NMP [12]. ATP binds in the pocket formed by the LID and CORE domains. AMP binds in the pocket formed by the NMP and CORE domains. The Whitford study also reported findings that show that localized regions of a protein unfold during conformational transitions. This mechanism reduces the strain and enhances catalytic efficiency. Local unfolding is the result of competing strain energies in the protein [10].
The local (thermodynamic) stability of the substrate-binding domains ATPlid and AMPlid has been shown to be significantly lower when compared with the CORE domain in ADKE. Coli. [13] Furthermore, it has been shown that the two subdomains (ATPlid and AMPlid) can fold and unfold in a "non-cooperative manner." [13] Binding of the substrates causes preference for 'closed' conformations amongst those that are sampled by ADK. These 'closed' conformations are hypothesized to help with removal of water from the active site to avoid wasteful hydrolysis of ATP in addition to helping optimize alignment of substrates for phosphoryl-transfer. [14] Furthermore, it has been shown that the apoenzyme will still sample the 'closed' conformations of the ATPlid and AMPlid domains in the absence of substrates [15]. When comparing the rate of opening of the enzyme (which allows for product release) and the rate of closing that accompanies substrate binding, closing was found to be the slower process.
The ability for a cell to dynamically measure energetic levels provides it with a method to monitor metabolic processes. [16] By continually monitoring and altering the levels of ATP and the other adenyl phosphates (ADP and AMP levels) adenylate kinase is an important regulator of energy expenditure at the cellular level. [17] As energy levels change under different metabolic stresses adenylate kinase is then able to generate AMP; which itself acts as a signaling molecule in further signaling cascades. This generated AMP can, for example, stimulate various AMP-dependent receptors such as those involved in glycolytic pathways, K-ATP channels, and 5' AMP-activated protein kinase ( AMPK). [16] Common factors that influence adenine nucleotide levels, and therefore ADK activity are exercise, stress, changes in hormone levels, and diet. [16] It facilitates decoding of cellular information by catalyzing nucleotide exchange in the intimate “sensing zone” of metabolic sensors [18].
Adenylate kinase is present in mitochondrial and myofibrillar compartments in the cell, and it makes two high-energy phosphoryls (β and γ) of ATP available to be transferred between adenine nucleotide molecules. [16] [17] In essence, adenylate kinase shuttles ATP to sites of high energy consumption and removes the AMP generated over the course of those reactions. These sequential phosphotransfer relays ultimately result in propagation of the phosphoryl groups along collections of ADK molecules. [16] This process can be thought of as a bucket brigade of ADK molecules that results in changes in local intracellular metabolic flux without apparent global changes in metabolite concentrations. [16] This process is extremely important for overall homeostasis of the cell. [16]
Nucleoside diphosphate (NDP) kinase catalyzes in vivo ATP-dependent synthesis of ribo- and deoxyribonucleoside triphosphates. In mutated Escherichia coli that had a disrupted nucleoside diphosphate kinase, adenylate kinase performed dual enzymatic functions. ADK complements nucleoside diphosphate kinase deficiency. [19]
Adenylate kinase deficiency in the erythrocyte is associated with hemolytic anemia. [20] This is a rare hereditary erythroenzymopathy that, in some cases, is associated with mental retardation and psychomotor impairment. [21] At least two patients have exhibited neonatal icterus and splenomegaly and required blood transfusions due to this deficiency. [22] In another patient, an abnormal fragment with homozygous and heterozygous A-->G substitutions at codon 164 caused severe erythrocyte ADK deficiency. [23] Two siblings had erythrocyte ADK deficiency, but one did not have evidence of hemolysis. [24]
Knock out of AK1 disrupts the synchrony between inorganic phosphate and turnover at ATP-consuming sites and ATP synthesis sites. This reduces the energetic signal communication in the post-ischemic heart and precipitates inadequate coronary reflow flowing ischemia-reperfusion. [25]
Adenylate Kinase 2 ( AK2) deficiency in humans causes hematopoietic defects associated with sensorineural deafness. [26] Recticular dysgenesis is an autosomal recessive form of human combined immunodeficiency. It is also characterized by an impaired lymphoid maturation and early differentiation arrest in the myeloid lineage. AK2 deficiency results in absent or a large decrease in the expression of proteins. AK2 is specifically expressed in the stria vascularis of the inner ear which indicates why individuals with an AK2 deficiency will have sensorineural deafness. [26]
AK1 genetic ablation decreases tolerance to metabolic stress. AK1 deficiency induces fiber-type specific variation in groups of transcripts in glycolysis and mitochondrial metabolism. [27] This supports muscle energy metabolism.
-Need to check for consistency, be sure to label everything ADK (NOT AK)
The mechanism link for the Adenylate kinase page is extremely weak. Not only does it not provide any of the actual structures or residues involved in the catalysis but in general only describes the reagents required for the catalysis. This may be because of larger articles under the umbrella of kinases that do go into more depth but if that is the case I think it is important that the article at least go into detail about why adenylate kinase is active on particular substrates, what gives it its specificity. The information from this mechanism page also only includes detail from a single study that was published in 2005. A quick search on google scholar shows that 'adenylate kinase mechanism' gives ~16,000 hits from articles exclusively published in 2013 and later. Ideally, in my article I would like to include what amino acids are directly involved in the catalysis, any adjacent ones that may be contributing electrostatic effects, and the substrate specificity of the adenylate kinase. Also, the structure and function portions could be expanded/rewritten.
 | This is a user sandbox of
Snodgrah. You can use it for testing or practicing edits. This is not the sandbox where you should draft your assigned article for a dashboard.wikiedu.org course. To find the right sandbox for your assignment, visit your Dashboard course page and follow the Sandbox Draft link for your assigned article in the My Articles section. |
Hi guys! I think the revisions and additions you have made are strong and fill in many of the missing gaps from the original article. I just have a couple of suggestions. First, watch your sentence structure. There are a couple of places that are really wordy (ex: Metabolic monitoring first sentence) or places where you could use more professional wording (ex: In "ADK Shuttle" section, "It all depends what stimuli..."). Second, add links to existing Wikipedia pages for terms that the reader of this page might not know. For example, the first time e. coli, Km, and CORE and LID domains are mentioned. If pages don't exist, it might be helpful to give a brief definition. Lastly, if the structures exist and if there's a significant difference visually, it might be cool to include images of the open and closed conformations. Otherwise, looks great, your mothers would be proud.
~Julia
It might be a good idea to completely rewrite the sections that you are adding to; this will be more consistent and easier to read. Also, it might be helpful to add a general reaction scheme at the beginning of the article so the function of the enzyme is immediately apparent. One more thing you could do would be to include pymol structures of the 'open' and 'closed' conformations that highlight the differences (if they're available), in addition to the binding of the substrate with a cartoon of the enzyme (instead of just the active site residues).
Hey Rago. We've toyed around with the idea of rewriting the sections before, it will probably happen during the final write up. I felt bad taking away from someone elses hard work :( . See above about the open/closed conformation PDB structures.
-AJR
Yo fam. Good edits overall. I agree with Julia and Rago's comments above. Additionally, at the beginning of your mechanism section you should clarify on "closing of the 'open lid'". Either that or moving the structure section before the mechanism (which would probably be the better fix seeing as mechanism is intricately dependent on structure) and touching up the wording for consistency. I'm not sure if this is possible, but enlarging the ChemDraw figures in the mechanism and structure sentences would also be helpful as they're really small and impossible to see. Otherwise, good job!
~Bates =)
Hey Julia! First off, thanks but my mother has never been proud. For the grammatical criticism, those will be revised. We can include some more links to the more technical terms too. There is a significant difference in the open and closed conformation, but unfortunately the only entry in the PDB under adenylate kinase in the open conformation was sent in in 2012 under a 'to be published' paper that was never published and is only a "Partly open" conformation so I'm a bit hesitant to include this.
Hey Rago. We've toyed around with the idea of rewriting the sections before, it will probably happen during the final write up. I felt bad taking away from someone elses hard work :( . See above about the open/closed conformation PDB structures.
Hey QT(3.14). We will definitely consider moving the sections around. This was more in order of what contributions we made first, not what we think the final order in the article should be.
Looking forward to incorporating these changes :D
~HS
- ^ The NIST Thermodynamics of Enzyme-Catalyzed Reactions database, http://xpdb.nist.gov/enzyme_thermodynamics/enzyme1.pl, Goldberg RN, Tewari YB, Bhat TN (November 2004). "Thermodynamics of enzyme-catalyzed reactions--a database for quantitative biochemistry". Bioinformatics. 20 (16): 2874–7. doi: 10.1093/bioinformatics/bth314. PMID 15145806., gives equilibrium constants, search for adenylate kinase under enzymes
- ^ Beis I, Newsholme EA (October 1975). "The contents of adenine nucleotides, phosphagens and some glycolytic intermediates in resting muscles from vertebrates and invertebrates". The Biochemical Journal. 152 (1): 23–32. doi: 10.1042/bj1520023. PMC 1172435. PMID 1212224.
- ^ a b Panayiotou, Christakis; Solaroli, Nicola; Karlsson, Anna (2014-04-01). "The many isoforms of human adenylate kinases". The International Journal of Biochemistry & Cell Biology. 49: 75–83. doi: 10.1016/j.biocel.2014.01.014.
- ^ Panayiotou, Christakis; Solaroli, Nicola; Xu, Yunjian; Johansson, Magnus; Karlsson, Anna (2011-02-01). "The characterization of human adenylate kinases 7 and 8 demonstrates differences in kinetic parameters and structural organization among the family of adenylate kinase isoenzymes". Biochemical Journal. 433 (3): 527–534. doi: 10.1042/BJ20101443. ISSN 0264-6021. PMID 21080915.
- ^ Tomasselli, Alfredo G.; Noda, Lafayette H. (1979-01-01). "Mitochondrial GTP-AMP Phosphotransferase". European Journal of Biochemistry. 93 (2): 263–270. doi: 10.1111/j.1432-1033.1979.tb12819.x. ISSN 1432-1033.
- ^ Jane Cooper, A.; Friedberg, Errol C. (1992-05-01). "A putative second adenylate kinase-encoding gene from the yeast Saccharomyces cerevisiae". Gene. 114 (1): 145–148. doi: 10.1016/0378-1119(92)90721-Z.
- ^ Henzler-Wildman, Katherine A.; Thai, Vu; Lei, Ming; Ott, Maria; Wolf-Watz, Magnus; Fenn, Tim; Pozharski, Ed; Wilson, Mark A.; Petsko, Gregory A. "Intrinsic motions along an enzymatic reaction trajectory". Nature. 450 (7171): 838–844. doi: 10.1038/nature06410.
- ^ Reinstein, J.; Gilles, A. M.; Rose, T.; Wittinghofer, A.; Girons, I. Saint; Bârzu, O.; Surewicz, W. K.; Mantsch, H. H. (1989-05-15). "Structural and catalytic role of arginine 88 in Escherichia coli adenylate kinase as evidenced by chemical modification and site-directed mutagenesis". Journal of Biological Chemistry. 264 (14): 8107–8112. ISSN 0021-9258. PMID 2542263.
- ^
a
b Muller, C., Schulz, G.E. (1991).
"Structure of the complex between adenylate kinase from Escherichia coli and the inhibitor Ap5A refined at 1.9 A resolution: A model for a catalytic transition state". Journal of Molecular Biology. 224: 159–177 – via Elsevier.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Whitford, Paul C.; Miyashita, Osamu; Levy, Yaakov; Onuchic, José N. (2007-03-09).
"Conformational Transitions of Adenylate Kinase: Switching by Cracking". Journal of Molecular Biology. 366 (5): 1661–1671.
doi:
10.1016/j.jmb.2006.11.085.
PMC
2561047.
PMID
17217965.
{{ cite journal}}: CS1 maint: PMC format ( link) -
^ Schrank, Travis P.; Bolen, D. Wayne; Hilser, Vincent J. (2009-10-06).
"Rational modulation of conformational fluctuations in adenylate kinase reveals a local unfolding mechanism for allostery and functional adaptation in proteins". Proceedings of the National Academy of Sciences. 106 (40): 16984–16989.
doi:
10.1073/pnas.0906510106.
ISSN
0027-8424.
PMC
2761315.
PMID
19805185.
{{ cite journal}}: CS1 maint: PMC format ( link) -
^ Daily, Michael D.; Phillips Jr., George N.; Cui, Qiang (2010-07-16).
"Many Local Motions Cooperate to Produce the Adenylate Kinase Conformational Transition". Journal of Molecular Biology. 400 (3): 618–631.
doi:
10.1016/j.jmb.2010.05.015.
PMC
2902635.
PMID
20471396.
{{ cite journal}}: CS1 maint: PMC format ( link) - ^ a b Rundqvist, Louise; Ådén, Jörgen; Sparrman, Tobias; Wallgren, Marcus; Olsson, Ulrika; Wolf-Watz, Magnus (2009-03-10). "Noncooperative Folding of Subdomains in Adenylate Kinase". Biochemistry. 48 (9): 1911–1927. doi: 10.1021/bi8018042. ISSN 0006-2960.
- ^ Olsson, Ulrika; Wolf-Watz, Magnus (2010-11-16). "Overlap between folding and functional energy landscapes for adenylate kinase conformational change". Nature Communications. 1 (8). doi: 10.1038/ncomms1106. ISSN 2041-1723.
- ^ Henzler-Wildman, Katherine A.; Thai, Vu; Lei, Ming; Ott, Maria; Wolf-Watz, Magnus; Fenn, Tim; Pozharski, Ed; Wilson, Mark A.; Petsko, Gregory A. "Intrinsic motions along an enzymatic reaction trajectory". Nature. 450 (7171): 838–844. doi: 10.1038/nature06410.
- ^
a
b
c
d
e
f
g Dzeja, Petras; Terzic, Andre (2009-04-17).
"Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing". International Journal of Molecular Sciences. 10 (4): 1729–1772.
doi:
10.3390/ijms10041729.
ISSN
1422-0067.
PMC
2680645.
PMID
19468337.
{{ cite journal}}: CS1 maint: PMC format ( link) CS1 maint: unflagged free DOI ( link) - ^
a
b Dzeja, Petras P.; Chung, Susan; Faustino, Randolph S.; Behfar, Atta; Terzic, Andre (2011-04-27).
"Developmental Enhancement of Adenylate Kinase-AMPK Metabolic Signaling Axis Supports Stem Cell Cardiac Differentiation". PLOS ONE. 6 (4): e19300.
doi:
10.1371/journal.pone.0019300.
ISSN
1932-6203.
PMC
3083437.
PMID
21556322.
{{ cite journal}}: CS1 maint: PMC format ( link) CS1 maint: unflagged free DOI ( link) -
^ Dzeja, Petras; Terzic, Andre (2009-04-17).
"Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing". International Journal of Molecular Sciences. 10 (4): 1729–1772.
doi:
10.3390/ijms10041729.
PMC
2680645.
PMID
19468337.
{{ cite journal}}: CS1 maint: PMC format ( link) CS1 maint: unflagged free DOI ( link) - ^ Lu Q, Inouye M (June 1996). "Adenylate kinase complements nucleoside diphosphate kinase deficiency in nucleotide metabolism". Proceedings of the National Academy of Sciences of the United States of America. 93 (12): 5720–5. doi: 10.1073/pnas.93.12.5720. PMC 39127. PMID 8650159.
- ^ Matsuura, S.; Igarashi, M.; Tanizawa, Y.; Yamada, M.; Kishi, F.; Kajii, T.; Fujii, H.; Miwa, S.; Sakurai, M.; Nakazawa, A. (Jun 1989). "Human adenylate kinase deficiency associated with hemolytic anemia. A single base substitution affecting solubility and catalytic activity of the cytosolic adenylate kinase". J Biol Chem. 264 (17): 10148–55. PMID 2542324.
- ^ Abrusci P, Chiarelli LR, Galizzi A, Fermo E, Bianchi P, Zanella A, Valentini G (August 2007). "Erythrocyte adenylate kinase deficiency: characterization of recombinant mutant forms and relationship with nonspherocytic hemolytic anemia". Experimental Hematology. 35 (8): 1182–9. doi: 10.1016/j.exphem.2007.05.004. PMID 17662886.
- ^ Corrons JL, Garcia E, Tusell JJ, Varughese KI, West C, Beutler E (July 2003). "Red cell adenylate kinase deficiency: molecular study of 3 new mutations (118G>A, 190G>A, and GAC deletion) associated with hereditary nonspherocytic hemolytic anemia". Blood. 102 (1): 353–6. doi: 10.1182/blood-2002-07-2288. PMID 12649162.
- ^ Qualtieri, A.; Pedace, V.; Bisconte, MG.; Bria, M.; Gulino, B.; Andreoli, V.; Brancati, C. (Dec 1997). "Severe erythrocyte adenylate kinase deficiency due to homozygous A-->G substitution at codon 164 of human AK1 gene associated with chronic haemolytic anaemia". Br J Haematol. 99 (4): 770–6. doi: 10.1046/j.1365-2141.1997.4953299.x. PMID 9432020.
- ^ Beutler E, Carson D, Dannawi H, Forman L, Kuhl W, West C, Westwood B (August 1983). "Metabolic compensation for profound erythrocyte adenylate kinase deficiency. A hereditary enzyme defect without hemolytic anemia". The Journal of Clinical Investigation. 72 (2): 648–55. doi: 10.1172/JCI111014. PMC 1129224. PMID 6308059.
-
^ Dzeja PP, Bast P, Pucar D, Wieringa B, Terzic A (October 2007).
"Defective metabolic signaling in adenylate kinase AK1 gene knock-out hearts compromises post-ischemic coronary reflow". The Journal of Biological Chemistry. 282 (43): 31366–72.
doi:
10.1074/jbc.M705268200.
PMC
3232003.
PMID
17704060.
{{ cite journal}}: CS1 maint: unflagged free DOI ( link) - ^ a b Lagresle-Peyrou C, Six EM, Picard C, Rieux-Laucat F, Michel V, Ditadi A, Demerens-de Chappedelaine C, Morillon E, Valensi F, Simon-Stoos KL, Mullikin JC, Noroski LM, Besse C, Wulffraat NM, Ferster A, Abecasis MM, Calvo F, Petit C, Candotti F, Abel L, Fischer A, Cavazzana-Calvo M (January 2009). "Human adenylate kinase 2 deficiency causes a profound hematopoietic defect associated with sensorineural deafness". Nature Genetics. 41 (1): 106–11. doi: 10.1038/ng.278. PMC 2612090. PMID 19043416.
-
^ Janssen E, de Groof A, Wijers M, Fransen J, Dzeja PP, Terzic A, Wieringa B (April 2003). "Adenylate kinase 1 deficiency induces molecular and structural adaptations to support muscle energy metabolism". The Journal of Biological Chemistry. 278 (15): 12937–45.
doi:
10.1074/jbc.M211465200.
PMID
12562761.
{{ cite journal}}: CS1 maint: unflagged free DOI ( link)
| Adenylate kinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 3D ribbon/surface model of adenylate kinase in complex with bis(adenosine)tetraphosphate (ADP-ADP) | |||||||||
| Identifiers | |||||||||
| Symbol | ADK | ||||||||
| Pfam | PF00406 | ||||||||
| InterPro | IPR000850 | ||||||||
| PROSITE | PDOC00104 | ||||||||
| SCOP2 | 1ake / SCOPe / SUPFAM | ||||||||
| |||||||||
Adenylate kinase ( EC 2.7.4.3) (also known as ADK or myokinase) is a phosphotransferase enzyme that catalyzes the interconversion of adenine nucleotides (ATP, ADP, and AMP). By constantly monitoring phosphate nucleotide levels inside the cell, ADK plays an important role in cellular energy homeostasis.
| ADK_lid | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Bacillus stearothermophilus adenylate kinase | |||||||||
| Identifiers | |||||||||
| Symbol | ADK_lid | ||||||||
| Pfam | PF05191 | ||||||||
| InterPro | IPR007862 | ||||||||
| PROSITE | PDOC00104 | ||||||||
| SCOP2 | 1ake / SCOPe / SUPFAM | ||||||||
| |||||||||
| Adenylate kinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 2.7.4.3 | ||||||||
| CAS no. | 2598011 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||

The reaction catalyzed is:
The equilibrium constant varies with condition, but it is close to 1. [1] Thus, ΔGo for this reaction is close to zero. In muscle from a variety of species of vertebrates and invertebrates, the concentration of ATP is typically 7-10 times that of ADP, and usually greater than 100 times that of AMP. [2] The rate of oxidative phosphorylation is controlled by the availability of ADP. Thus, the mitochondrion attempts to keep ATP levels high due to the combined action of adenylate kinase and the controls on oxidative phosphorylation.
To date there have been nine human ADK protein isoforms identified. While some of these are ubiquitous throughout the body, some are localized into specific tissues. For example, ADK7 and ADK8 are both only found in the cytosol of cells; and ADK7 is found in skeletal muscle whereas ADK8 is not [3]. Not only do the locations of the various isoforms within the cell vary, but the binding of substrate to the enzyme and kinetics of the phosphoryl transfer are different as well. ADK1, the most abundant cytosolic ADK isozyme, has a Km about a thousand times higher than the Km of ADK7 and 8, indicating a much weaker binding of ADK1 to AMP [4]. Sub-cellular localization of the ADK enzymes is done by including a targeting sequence in the protein [3]. Each isoform also has different preference for NTP's. Some will only use ATP, whereas others will accept GTP, UTP, and CTP as the phosphoryl carrier.
Some of these isoforms prefer other NTP's entirely. There is a mitochondrial GTP:AMP phosphotransferase, also specific for the phosphorylation of AMP, that can only use GTP or ITP as the phosphoryl donor [5]. ADK has also been identified in different bacterial species and in yeast [6]. Two further enzymes are known to be related to the ADK family, i.e. yeast uridine monophosphokinase and slime mold UMP-CMP kinase. Some rfesidues are conserved across these isoforms, indicating how essential they are for catalysis. One of the most conserved areas includes an Arg residue, whose modification inactivates the enzyme, together with an Asp that resides in the catalytic cleft of the enzyme and participates in a salt bridge.
 | This is a user sandbox of
Snodgrah. You can use it for testing or practicing edits. This is not the sandbox where you should draft your assigned article for a dashboard.wikiedu.org course. To find the right sandbox for your assignment, visit your Dashboard course page and follow the Sandbox Draft link for your assigned article in the My Articles section. |
- Adenylate kinase, subfamily InterPro: IPR006259
- UMP-CMP kinase InterPro: IPR006266
- Adenylate kinase, isozyme 1 InterPro: IPR006267
Phosphoryl transfer only occurs on closing of the 'open lid'. This causes an exclusion of water molecules that brings the substrates in proximity to each other [7], lowering the energy barrier for the nucleophilic attack by the γ-phosphoryl group of ATP on the α-phosphoryl of AMP. In the crystal structure of the ADK enzyme from E. Coli with inhibitor Ap5A, the Arg88 residue binds the Ap5A at the α-phosphate group. It has been shown that the mutation R88G results in 99% loss of catalytic activity of this enzyme, suggesting that this residue is intimately involved in the phosphoryl transfer [8]. Another highly conserved residue is Arg119, which lies in the adenosine binding region of the ADK, and acts to sandwich the adenine in the active site. It has been suggested that the promiscuity of these enzymes in accepting other NTP's is due to this relatively inconsequential interactions of the base in the ATP binding pocket [9]. A network of positive, conserved residues (Lys13, Arg123, Arg156, and Arg167 in ADK from E. Coli) stabilize the build up of negative charge on phosphoryl group during the transfer. Two distal aspartate residues bind to the arginine network, causing the enzyme to fold and reduces its flexibility. A magnesium cofactor is also required, essential for increasing the electrophilicity of the phosphate on AMP, though this magnesium ion is only held in the active pocket by electrostatic interactions and dissociates e
asily [9].

Flexibility and plasticity allow proteins to bind to ligands, form oligomers, aggregate, and perform mechanical work [10]. Large conformational changes in proteins play an important role in cellular signaling. Adenylate Kinase is a signal transducing protein; thus, the balance between conformations regulates protein activity. ADK has a locally unfolded state that becomes depopulated upon binding [11].

A 2007 study by Whitford et al. shows the conformations of ADK when binding with ATP or AMP [10]. The study shows that there are three relevant conformations or structures of ADK—CORE, Open, and Closed. In ADK, there are two small domains called the LID and NMP [12]. ATP binds in the pocket formed by the LID and CORE domains. AMP binds in the pocket formed by the NMP and CORE domains. The Whitford study also reported findings that show that localized regions of a protein unfold during conformational transitions. This mechanism reduces the strain and enhances catalytic efficiency. Local unfolding is the result of competing strain energies in the protein [10].
The local (thermodynamic) stability of the substrate-binding domains ATPlid and AMPlid has been shown to be significantly lower when compared with the CORE domain in ADKE. Coli. [13] Furthermore, it has been shown that the two subdomains (ATPlid and AMPlid) can fold and unfold in a "non-cooperative manner." [13] Binding of the substrates causes preference for 'closed' conformations amongst those that are sampled by ADK. These 'closed' conformations are hypothesized to help with removal of water from the active site to avoid wasteful hydrolysis of ATP in addition to helping optimize alignment of substrates for phosphoryl-transfer. [14] Furthermore, it has been shown that the apoenzyme will still sample the 'closed' conformations of the ATPlid and AMPlid domains in the absence of substrates [15]. When comparing the rate of opening of the enzyme (which allows for product release) and the rate of closing that accompanies substrate binding, closing was found to be the slower process.
The ability for a cell to dynamically measure energetic levels provides it with a method to monitor metabolic processes. [16] By continually monitoring and altering the levels of ATP and the other adenyl phosphates (ADP and AMP levels) adenylate kinase is an important regulator of energy expenditure at the cellular level. [17] As energy levels change under different metabolic stresses adenylate kinase is then able to generate AMP; which itself acts as a signaling molecule in further signaling cascades. This generated AMP can, for example, stimulate various AMP-dependent receptors such as those involved in glycolytic pathways, K-ATP channels, and 5' AMP-activated protein kinase ( AMPK). [16] Common factors that influence adenine nucleotide levels, and therefore ADK activity are exercise, stress, changes in hormone levels, and diet. [16] It facilitates decoding of cellular information by catalyzing nucleotide exchange in the intimate “sensing zone” of metabolic sensors [18].
Adenylate kinase is present in mitochondrial and myofibrillar compartments in the cell, and it makes two high-energy phosphoryls (β and γ) of ATP available to be transferred between adenine nucleotide molecules. [16] [17] In essence, adenylate kinase shuttles ATP to sites of high energy consumption and removes the AMP generated over the course of those reactions. These sequential phosphotransfer relays ultimately result in propagation of the phosphoryl groups along collections of ADK molecules. [16] This process can be thought of as a bucket brigade of ADK molecules that results in changes in local intracellular metabolic flux without apparent global changes in metabolite concentrations. [16] This process is extremely important for overall homeostasis of the cell. [16]
Nucleoside diphosphate (NDP) kinase catalyzes in vivo ATP-dependent synthesis of ribo- and deoxyribonucleoside triphosphates. In mutated Escherichia coli that had a disrupted nucleoside diphosphate kinase, adenylate kinase performed dual enzymatic functions. ADK complements nucleoside diphosphate kinase deficiency. [19]
Adenylate kinase deficiency in the erythrocyte is associated with hemolytic anemia. [20] This is a rare hereditary erythroenzymopathy that, in some cases, is associated with mental retardation and psychomotor impairment. [21] At least two patients have exhibited neonatal icterus and splenomegaly and required blood transfusions due to this deficiency. [22] In another patient, an abnormal fragment with homozygous and heterozygous A-->G substitutions at codon 164 caused severe erythrocyte ADK deficiency. [23] Two siblings had erythrocyte ADK deficiency, but one did not have evidence of hemolysis. [24]
Knock out of AK1 disrupts the synchrony between inorganic phosphate and turnover at ATP-consuming sites and ATP synthesis sites. This reduces the energetic signal communication in the post-ischemic heart and precipitates inadequate coronary reflow flowing ischemia-reperfusion. [25]
Adenylate Kinase 2 ( AK2) deficiency in humans causes hematopoietic defects associated with sensorineural deafness. [26] Recticular dysgenesis is an autosomal recessive form of human combined immunodeficiency. It is also characterized by an impaired lymphoid maturation and early differentiation arrest in the myeloid lineage. AK2 deficiency results in absent or a large decrease in the expression of proteins. AK2 is specifically expressed in the stria vascularis of the inner ear which indicates why individuals with an AK2 deficiency will have sensorineural deafness. [26]
AK1 genetic ablation decreases tolerance to metabolic stress. AK1 deficiency induces fiber-type specific variation in groups of transcripts in glycolysis and mitochondrial metabolism. [27] This supports muscle energy metabolism.
-Need to check for consistency, be sure to label everything ADK (NOT AK)
The mechanism link for the Adenylate kinase page is extremely weak. Not only does it not provide any of the actual structures or residues involved in the catalysis but in general only describes the reagents required for the catalysis. This may be because of larger articles under the umbrella of kinases that do go into more depth but if that is the case I think it is important that the article at least go into detail about why adenylate kinase is active on particular substrates, what gives it its specificity. The information from this mechanism page also only includes detail from a single study that was published in 2005. A quick search on google scholar shows that 'adenylate kinase mechanism' gives ~16,000 hits from articles exclusively published in 2013 and later. Ideally, in my article I would like to include what amino acids are directly involved in the catalysis, any adjacent ones that may be contributing electrostatic effects, and the substrate specificity of the adenylate kinase. Also, the structure and function portions could be expanded/rewritten.
 | This is a user sandbox of
Snodgrah. You can use it for testing or practicing edits. This is not the sandbox where you should draft your assigned article for a dashboard.wikiedu.org course. To find the right sandbox for your assignment, visit your Dashboard course page and follow the Sandbox Draft link for your assigned article in the My Articles section. |
Hi guys! I think the revisions and additions you have made are strong and fill in many of the missing gaps from the original article. I just have a couple of suggestions. First, watch your sentence structure. There are a couple of places that are really wordy (ex: Metabolic monitoring first sentence) or places where you could use more professional wording (ex: In "ADK Shuttle" section, "It all depends what stimuli..."). Second, add links to existing Wikipedia pages for terms that the reader of this page might not know. For example, the first time e. coli, Km, and CORE and LID domains are mentioned. If pages don't exist, it might be helpful to give a brief definition. Lastly, if the structures exist and if there's a significant difference visually, it might be cool to include images of the open and closed conformations. Otherwise, looks great, your mothers would be proud.
~Julia
It might be a good idea to completely rewrite the sections that you are adding to; this will be more consistent and easier to read. Also, it might be helpful to add a general reaction scheme at the beginning of the article so the function of the enzyme is immediately apparent. One more thing you could do would be to include pymol structures of the 'open' and 'closed' conformations that highlight the differences (if they're available), in addition to the binding of the substrate with a cartoon of the enzyme (instead of just the active site residues).
Hey Rago. We've toyed around with the idea of rewriting the sections before, it will probably happen during the final write up. I felt bad taking away from someone elses hard work :( . See above about the open/closed conformation PDB structures.
-AJR
Yo fam. Good edits overall. I agree with Julia and Rago's comments above. Additionally, at the beginning of your mechanism section you should clarify on "closing of the 'open lid'". Either that or moving the structure section before the mechanism (which would probably be the better fix seeing as mechanism is intricately dependent on structure) and touching up the wording for consistency. I'm not sure if this is possible, but enlarging the ChemDraw figures in the mechanism and structure sentences would also be helpful as they're really small and impossible to see. Otherwise, good job!
~Bates =)
Hey Julia! First off, thanks but my mother has never been proud. For the grammatical criticism, those will be revised. We can include some more links to the more technical terms too. There is a significant difference in the open and closed conformation, but unfortunately the only entry in the PDB under adenylate kinase in the open conformation was sent in in 2012 under a 'to be published' paper that was never published and is only a "Partly open" conformation so I'm a bit hesitant to include this.
Hey Rago. We've toyed around with the idea of rewriting the sections before, it will probably happen during the final write up. I felt bad taking away from someone elses hard work :( . See above about the open/closed conformation PDB structures.
Hey QT(3.14). We will definitely consider moving the sections around. This was more in order of what contributions we made first, not what we think the final order in the article should be.
Looking forward to incorporating these changes :D
~HS
- ^ The NIST Thermodynamics of Enzyme-Catalyzed Reactions database, http://xpdb.nist.gov/enzyme_thermodynamics/enzyme1.pl, Goldberg RN, Tewari YB, Bhat TN (November 2004). "Thermodynamics of enzyme-catalyzed reactions--a database for quantitative biochemistry". Bioinformatics. 20 (16): 2874–7. doi: 10.1093/bioinformatics/bth314. PMID 15145806., gives equilibrium constants, search for adenylate kinase under enzymes
- ^ Beis I, Newsholme EA (October 1975). "The contents of adenine nucleotides, phosphagens and some glycolytic intermediates in resting muscles from vertebrates and invertebrates". The Biochemical Journal. 152 (1): 23–32. doi: 10.1042/bj1520023. PMC 1172435. PMID 1212224.
- ^ a b Panayiotou, Christakis; Solaroli, Nicola; Karlsson, Anna (2014-04-01). "The many isoforms of human adenylate kinases". The International Journal of Biochemistry & Cell Biology. 49: 75–83. doi: 10.1016/j.biocel.2014.01.014.
- ^ Panayiotou, Christakis; Solaroli, Nicola; Xu, Yunjian; Johansson, Magnus; Karlsson, Anna (2011-02-01). "The characterization of human adenylate kinases 7 and 8 demonstrates differences in kinetic parameters and structural organization among the family of adenylate kinase isoenzymes". Biochemical Journal. 433 (3): 527–534. doi: 10.1042/BJ20101443. ISSN 0264-6021. PMID 21080915.
- ^ Tomasselli, Alfredo G.; Noda, Lafayette H. (1979-01-01). "Mitochondrial GTP-AMP Phosphotransferase". European Journal of Biochemistry. 93 (2): 263–270. doi: 10.1111/j.1432-1033.1979.tb12819.x. ISSN 1432-1033.
- ^ Jane Cooper, A.; Friedberg, Errol C. (1992-05-01). "A putative second adenylate kinase-encoding gene from the yeast Saccharomyces cerevisiae". Gene. 114 (1): 145–148. doi: 10.1016/0378-1119(92)90721-Z.
- ^ Henzler-Wildman, Katherine A.; Thai, Vu; Lei, Ming; Ott, Maria; Wolf-Watz, Magnus; Fenn, Tim; Pozharski, Ed; Wilson, Mark A.; Petsko, Gregory A. "Intrinsic motions along an enzymatic reaction trajectory". Nature. 450 (7171): 838–844. doi: 10.1038/nature06410.
- ^ Reinstein, J.; Gilles, A. M.; Rose, T.; Wittinghofer, A.; Girons, I. Saint; Bârzu, O.; Surewicz, W. K.; Mantsch, H. H. (1989-05-15). "Structural and catalytic role of arginine 88 in Escherichia coli adenylate kinase as evidenced by chemical modification and site-directed mutagenesis". Journal of Biological Chemistry. 264 (14): 8107–8112. ISSN 0021-9258. PMID 2542263.
- ^
a
b Muller, C., Schulz, G.E. (1991).
"Structure of the complex between adenylate kinase from Escherichia coli and the inhibitor Ap5A refined at 1.9 A resolution: A model for a catalytic transition state". Journal of Molecular Biology. 224: 159–177 – via Elsevier.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Whitford, Paul C.; Miyashita, Osamu; Levy, Yaakov; Onuchic, José N. (2007-03-09).
"Conformational Transitions of Adenylate Kinase: Switching by Cracking". Journal of Molecular Biology. 366 (5): 1661–1671.
doi:
10.1016/j.jmb.2006.11.085.
PMC
2561047.
PMID
17217965.
{{ cite journal}}: CS1 maint: PMC format ( link) -
^ Schrank, Travis P.; Bolen, D. Wayne; Hilser, Vincent J. (2009-10-06).
"Rational modulation of conformational fluctuations in adenylate kinase reveals a local unfolding mechanism for allostery and functional adaptation in proteins". Proceedings of the National Academy of Sciences. 106 (40): 16984–16989.
doi:
10.1073/pnas.0906510106.
ISSN
0027-8424.
PMC
2761315.
PMID
19805185.
{{ cite journal}}: CS1 maint: PMC format ( link) -
^ Daily, Michael D.; Phillips Jr., George N.; Cui, Qiang (2010-07-16).
"Many Local Motions Cooperate to Produce the Adenylate Kinase Conformational Transition". Journal of Molecular Biology. 400 (3): 618–631.
doi:
10.1016/j.jmb.2010.05.015.
PMC
2902635.
PMID
20471396.
{{ cite journal}}: CS1 maint: PMC format ( link) - ^ a b Rundqvist, Louise; Ådén, Jörgen; Sparrman, Tobias; Wallgren, Marcus; Olsson, Ulrika; Wolf-Watz, Magnus (2009-03-10). "Noncooperative Folding of Subdomains in Adenylate Kinase". Biochemistry. 48 (9): 1911–1927. doi: 10.1021/bi8018042. ISSN 0006-2960.
- ^ Olsson, Ulrika; Wolf-Watz, Magnus (2010-11-16). "Overlap between folding and functional energy landscapes for adenylate kinase conformational change". Nature Communications. 1 (8). doi: 10.1038/ncomms1106. ISSN 2041-1723.
- ^ Henzler-Wildman, Katherine A.; Thai, Vu; Lei, Ming; Ott, Maria; Wolf-Watz, Magnus; Fenn, Tim; Pozharski, Ed; Wilson, Mark A.; Petsko, Gregory A. "Intrinsic motions along an enzymatic reaction trajectory". Nature. 450 (7171): 838–844. doi: 10.1038/nature06410.
- ^
a
b
c
d
e
f
g Dzeja, Petras; Terzic, Andre (2009-04-17).
"Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing". International Journal of Molecular Sciences. 10 (4): 1729–1772.
doi:
10.3390/ijms10041729.
ISSN
1422-0067.
PMC
2680645.
PMID
19468337.
{{ cite journal}}: CS1 maint: PMC format ( link) CS1 maint: unflagged free DOI ( link) - ^
a
b Dzeja, Petras P.; Chung, Susan; Faustino, Randolph S.; Behfar, Atta; Terzic, Andre (2011-04-27).
"Developmental Enhancement of Adenylate Kinase-AMPK Metabolic Signaling Axis Supports Stem Cell Cardiac Differentiation". PLOS ONE. 6 (4): e19300.
doi:
10.1371/journal.pone.0019300.
ISSN
1932-6203.
PMC
3083437.
PMID
21556322.
{{ cite journal}}: CS1 maint: PMC format ( link) CS1 maint: unflagged free DOI ( link) -
^ Dzeja, Petras; Terzic, Andre (2009-04-17).
"Adenylate Kinase and AMP Signaling Networks: Metabolic Monitoring, Signal Communication and Body Energy Sensing". International Journal of Molecular Sciences. 10 (4): 1729–1772.
doi:
10.3390/ijms10041729.
PMC
2680645.
PMID
19468337.
{{ cite journal}}: CS1 maint: PMC format ( link) CS1 maint: unflagged free DOI ( link) - ^ Lu Q, Inouye M (June 1996). "Adenylate kinase complements nucleoside diphosphate kinase deficiency in nucleotide metabolism". Proceedings of the National Academy of Sciences of the United States of America. 93 (12): 5720–5. doi: 10.1073/pnas.93.12.5720. PMC 39127. PMID 8650159.
- ^ Matsuura, S.; Igarashi, M.; Tanizawa, Y.; Yamada, M.; Kishi, F.; Kajii, T.; Fujii, H.; Miwa, S.; Sakurai, M.; Nakazawa, A. (Jun 1989). "Human adenylate kinase deficiency associated with hemolytic anemia. A single base substitution affecting solubility and catalytic activity of the cytosolic adenylate kinase". J Biol Chem. 264 (17): 10148–55. PMID 2542324.
- ^ Abrusci P, Chiarelli LR, Galizzi A, Fermo E, Bianchi P, Zanella A, Valentini G (August 2007). "Erythrocyte adenylate kinase deficiency: characterization of recombinant mutant forms and relationship with nonspherocytic hemolytic anemia". Experimental Hematology. 35 (8): 1182–9. doi: 10.1016/j.exphem.2007.05.004. PMID 17662886.
- ^ Corrons JL, Garcia E, Tusell JJ, Varughese KI, West C, Beutler E (July 2003). "Red cell adenylate kinase deficiency: molecular study of 3 new mutations (118G>A, 190G>A, and GAC deletion) associated with hereditary nonspherocytic hemolytic anemia". Blood. 102 (1): 353–6. doi: 10.1182/blood-2002-07-2288. PMID 12649162.
- ^ Qualtieri, A.; Pedace, V.; Bisconte, MG.; Bria, M.; Gulino, B.; Andreoli, V.; Brancati, C. (Dec 1997). "Severe erythrocyte adenylate kinase deficiency due to homozygous A-->G substitution at codon 164 of human AK1 gene associated with chronic haemolytic anaemia". Br J Haematol. 99 (4): 770–6. doi: 10.1046/j.1365-2141.1997.4953299.x. PMID 9432020.
- ^ Beutler E, Carson D, Dannawi H, Forman L, Kuhl W, West C, Westwood B (August 1983). "Metabolic compensation for profound erythrocyte adenylate kinase deficiency. A hereditary enzyme defect without hemolytic anemia". The Journal of Clinical Investigation. 72 (2): 648–55. doi: 10.1172/JCI111014. PMC 1129224. PMID 6308059.
-
^ Dzeja PP, Bast P, Pucar D, Wieringa B, Terzic A (October 2007).
"Defective metabolic signaling in adenylate kinase AK1 gene knock-out hearts compromises post-ischemic coronary reflow". The Journal of Biological Chemistry. 282 (43): 31366–72.
doi:
10.1074/jbc.M705268200.
PMC
3232003.
PMID
17704060.
{{ cite journal}}: CS1 maint: unflagged free DOI ( link) - ^ a b Lagresle-Peyrou C, Six EM, Picard C, Rieux-Laucat F, Michel V, Ditadi A, Demerens-de Chappedelaine C, Morillon E, Valensi F, Simon-Stoos KL, Mullikin JC, Noroski LM, Besse C, Wulffraat NM, Ferster A, Abecasis MM, Calvo F, Petit C, Candotti F, Abel L, Fischer A, Cavazzana-Calvo M (January 2009). "Human adenylate kinase 2 deficiency causes a profound hematopoietic defect associated with sensorineural deafness". Nature Genetics. 41 (1): 106–11. doi: 10.1038/ng.278. PMC 2612090. PMID 19043416.
-
^ Janssen E, de Groof A, Wijers M, Fransen J, Dzeja PP, Terzic A, Wieringa B (April 2003). "Adenylate kinase 1 deficiency induces molecular and structural adaptations to support muscle energy metabolism". The Journal of Biological Chemistry. 278 (15): 12937–45.
doi:
10.1074/jbc.M211465200.
PMID
12562761.
{{ cite journal}}: CS1 maint: unflagged free DOI ( link)