Proto-oncogene protein Wnt-1 protein, short for Wingless-Type MMTV Integration Site Family Member 1, is encoded by the Wnt-1 gene. [1] Wnt-1 is part of the Wnt-gene family, together forming a highly conserved secreted signaling pathway that regulate cell-to-cell interaction during embryogenesis. [2]
Wnt-1 is a secreted glycoprotein, usually around 350-400 amino acids in length with conserved pattern of 23-25 cysteine residues across species. [3]
During embryonic development, Wnt-1 plays a role in generation of cell polarity, body axis formation, [4] and development of the central nervous system. [5] It also has a role in the regeneration of tissues in adult organisms. [6] A mis-expression of Wnt-1 is correlated with tumorogenesis. [3]
Wnt signals through cell-surface receptors in the Frizzled and LRP families. [7] [2] Binding of Wnt protein to cell-surface receptor triggers a signaling cascade involving intracellular signaling protein beta-catenin, which leads to the increased expression of 50+ genes. [7]
Wnt-1 protein can be referred to as Int-1 or Wg in literature, varying by the organism studied and the time of research. Int-1 (Integration site-1) was first described by Nusse and Varmus in 1982 as the proto-oncogene activated by mouse mammary tumor virus (MMTV). [8] In 1976, the ortholog gene in Drosophila melanogaster was described as Wg (Wingless), from the wingless phenotype of mutants. [9] To reduce confusion, Int-1 and Wg were combined and renamed as Wnt-1 around the 1990s. [10]
Wnt-1 is required for proper formation of the midbrain and anterior hindbrain in the mouse. [5] Mutations of the Wnt-1 gene in embryonic mice results in a wide range of phenotypes: some die at birth lacking an entire cerebellum and parts of the midbrain, while some live through adulthood suffering from ataxia and lacking the anterior half of the cerebellum. [5] The malformed parts of the brain in Wnt-1 mutants correspond to not only the region where Wnt-1 would normally be highly expressed, but also the surrounding tissues. [11] This suggests Wnt-1 protein’s importance in inducing development of surrounding tissues. [12] No explanation for the variability of Wnt-1 mutant phenotype expressed is known, but it may be due to several factors, including genetic background. [13]
The role of Wnt-1 protein in body-axis formation was studied in the model organism Xenopus laevis. The injection of Wnt-1 mRNA into early Xenopus embryo results in duplicated dorsal axis. This suggests the role of Wnt-1 as an organizer molecule and embryonic signal inducer for pattern formation. [14]
Wnt proteins are involved in a multitude of pathways. In most pathways, Wnt is found to interact directly with transmembrane proteins of the Frizzled family. [15] Wnt1 is specifically found to interact with Fzd8. The binding of Wnt to Fzd8 requires a dimerization of two CRD domains of Fzd. [16]
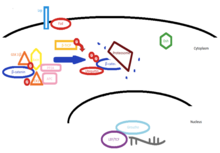
In the absence of Wnt, β-catenin is phosphorylated and marked for degradation by ubiquitin mediated proteosomes. [17] This phosphorylation is catalyzed by serine/threonine glycogen synthase kinase-3beta (GSK-3β) in complex with casein kinase 1 (CK1), the adenomatous polyposis coli (APC) protein, axin and protein phosphatase 2A (PP2A). [15] [17]This process of phosphorylation by GSK-3 β occurs at serines-33 and -37, as well as at serine-45. [18] The process is continued by β-TrCP in conjunction with ubiquitin ligase in order to mark the protein with ubiquitin for degradation. [19]

A change in the cytoplasmic conformation of the proteins occurs when Wnt binds to the Fzd/LRP complex, resulting in the binding of Axin to LRP and the binding of the Dishevelled protein (Dsh) to Fzd. [20] Other transmembrane proteins found to interact with Fzd8 and assist in the signal transmission across the membrane are two low density lipoprotein receptor-related proteins, LRP5 and LRP6. [20] [15]The deactivation of GSK-3β is coupled to the interaction between axin and dishevelled (DIX) domain, thus preventing the formation of the axin dependant phosphorylation complex. [21] [22] This prevents phosphorylation of β-catenin and allows for its accumulation.
In the nucleus without β-catenin, two transcription factors,
T-cell factor (TCF) and
Lymphoid enhancer factor (LEF) in conjugation with the protein
Groucho, act as
inhibitors of Wnt controlled genes.
[17] Once build up of β-catenin occurs, it acts to from a complex with TCF/LEF. The complex then acts to induce the same genes that TCF/LEF inhibited in the absence of β-catenin, specifically those relating to
cell growth and
differentiation.
[17]
The two major non-canonical pathways are the planar cell polarity (PCP) pathway and the Wnt/Ca2+ pathway. Both of these pathways are also reliant on the Frizzled transmembrane protein. [17] It is suggested that the activation of these alternate pathways is dependent on the specific Fzd related membrane proteins present, such as LRP5 or LRP6. The Wnt/Ca2+ pathway is reliant on a reaction between dishevelled protein and Fzd, causing the activation of a G-protein coupled signal transduction cascade.The ultimate release of calcium from this mechanism activates calcium/calmodium-dependent protein kinase II (CaMKII). [17] In developmental processes, the release of such kinases results in a decrease in the effect of the canonical beta catenin pathway, and plays a role in cell migration and axis development. Planar Cell Polarity (PCP) is a pathway involving many of the same proteins such as Fzd and Dishevelled , but primarily plays a role in altering cell morphology. [23] This pathway involves the activation of Rho- and Rac-GTPases at the membrane, followed by the activation of different Myosins and Kinases which alter cell structure. [15] [23]
Wnt-1 mis-expression are of clinical significance since mutations are associated chronic diseases.
An overexpression of Wnt-1 is correlated with tumorogenesis. [3] Wnt-1 was first identified in 1982 by Nesse and Varmus as a proto-oncogene activated by integration of mouse mammary tumor virus (MMTV). [8] Today, the mis-expression of Wnt-1 is correlated with diseases such as oral squamous cell carcinoma (OSCC), [24] breast tumours in human breast cancer [25] (19), non-small cell lung cancer (NSCLC), [26] and gastric cancer. [27]
The up-regulation of Wnt proteins can enhance the process of osteocyte proliferation. [28] Injecting mice with purified Wnt3a protein resulted in proliferation of liposomes at the sites of bone damage. This not only suggests an additional area of therapeutic research using Wnt pathways, but also the potential positive effects of up-regulated Wnt protein expressions in tissue/organ regeneration. Wnt-1 over-expression can suppress the lymphangiogenesis and potentiates the delay of metastasis in melanoma. [29] In addition, Wnt-1 expression has shown to aid the restoration of cardiac functions by up-regulating the formation of new cardiac fibroblast cells, along the Wnt-1/β-catenin pathway in the cases of acute cardiac ischemia. [30]
Inhibition studies of the Wnt-gene family were long focused due to their cancer-causing properties when over-expressed. Over-expression of Wnt-1 has demonstrated a strong causational relationship to the formation of mammary gland tumor, [25] a deeper understanding of the specific Wnt-1 pathway in relevant organs will potentially bring a more effective treatment therapy for breast cancer.
The highly conserved nature of Wnt gene throughout evolution creates an opportunity for research to take place in multiple model organisms, including worms, flies, and mice. [31] [2] The human Wnt-1 protein sequence is 99% identical to that of the mouse homologue. [32] Wnt-1 is usually around 350-400 amino acids in length with conserved pattern of 23-25 cysteine residues across species. [3]
Evolutionary biologists speculate the early amplification and divergence of the Wnt family were the cause for increasing complexity of animal body plans. Wnt-1 has been found conserved throughout the entire animal kingdom, often expressed in regionalized patterns. [33]
- ^ "WNT1 wingless-type MMTV integration site family, member 1 [homo sapiens]". NCBI. Retrieved 10 November 2012.
- ^ a b c Nusse, Roel. "The Wnt Homepage". Retrieved 10 November 2012.
- ^
a
b
c
d Cardigan, Ken (1997).
"Wnt signaling: a common theme in animal development". Genes & Development. 11: 3286–3305.
doi:
10.1101/gad.11.24.3286 Genes & Dev. 1997. 11: 3286-3305 (inactive 2023-08-02). Retrieved 17 November 2012.
{{ cite journal}}: Check|doi=value ( help); Unknown parameter|coauthors=ignored (|author=suggested) ( help)CS1 maint: DOI inactive as of August 2023 ( link) -
^ McMahon, Andrew P.; Moon, Randall T. (1989 Sep 22).
"Ectopic expression of the proto-oncogene int-1 in Xenopus embryos leads to duplication of the embryonic axis". Cell. 58 (6): 1075–84.
doi:
10.1016/0092-8674(89)90506-0.
PMID
[PubMed - indexed for MEDLINE 2673541 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^
a
b
c Thomas, Kirk R.; Capecchi, Mario R. (NaN undefined NaN).
"Targeted disruption of the murine int-1 proto-oncogene resulting in severe abnormalities in midbrain and cerebellar development". Nature. 346 (6287): 847–850.
doi:
10.1038/346847a0.
PMID
2202907.
{{ cite journal}}: Check date values in:|date=( help) -
^ Gurley, Kyle A.; Elliott, Sarah A.; Simakov, Oleg; Schmidt, Heiko A.; Holstein, Thomas W.; Alvarado, Alejandro Sánchez (2010 Nov 1).
"Expression of secreted Wnt pathway components reveals unexpected complexity of the planarian amputation response". Developmental Biology. 347 (1): 24–39.
doi:
10.1016/j.ydbio.2010.08.007.
PMID
[PubMed - indexed for MEDLINE 20707997 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^ a b "Wnt1 protein (30R-AW002)". Fitzgerald-FII Supplier of Antibodies Protein and ELISA kits. Retrieved 18 November 2012.
- ^
a
b Nusse, Roel; Varmus, Harold E. (November 1982).
"Many tumors induced by the mouse mammary tumor virus contain a provirus integrated in the same region of the host genome". Cell. 31 (1): 99–109.
doi:
10.1016/0092-8674(82)90409-3.
PMID
6297757.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Sharma, R.P.; Chopra, V.L. (Feb).
"Effect of the Wingless (wg1) mutation on wing and haltere development in Drosophila melanogaster". Developmental Biology. 48 (2): 461–5.
doi:
10.1016/0012-1606(76)90108-1.
PMID
[PubMed - indexed for MEDLINE 815114 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=and|year=/|date=mismatch ( help) -
^ Nusse, Roel; Varmus, Harold (2012 May 22).
"Three decades of Wnts: a personal perspective on how a scientific field developed". The EMBO Journal. 31 (12): 2670–84.
doi:
10.1038/emboj.2012.146.
PMID
[PubMed - indexed for MEDLINE 22617420 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ McMahon, Andrew P.; Bradley, Allan (NaN undefined NaN).
"The Wnt-1 (int-1) proto-oncogene is required for development of a large region of the mouse brain". Cell. 62 (6): 1073–1085.
doi:
10.1016/0092-8674(90)90385-R.
PMID
2205396.
{{ cite journal}}: Check date values in:|date=( help) -
^ Nusse, R (1992 Sep).
"The Wnt gene family in tumorigenesis and in normal development". The Journal of Steroid Biochemistry and Molecular Biology. 43 (1–3): 9–12.
doi:
10.1016/0960-0760(92)90181-H.
PMID
[PubMed - indexed for MEDLINE 1388050 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Nusse, R.; Varmus, H. E. (26).
"Wnt Genes Review" (PDF). Cell. 69 (7): 1073–1087.
doi:
10.1016/0092-8674(92)90630-U.
PMID
1617723.
{{ cite journal}}: Check date values in:|date=and|year=/|date=mismatch ( help); Unknown parameter|month=ignored ( help) -
^ McMahon, Andrew P.; Moon, Randall T. (1989 Sep 22).
"Ectopic expression of the proto-oncogene int-1 in Xenopus embryos leads to duplication of the embryonic axis". Cell. 58 (6): 1075–84.
doi:
10.1016/0092-8674(89)90506-0.
PMID
[PubMed - indexed for MEDLINE 2673541 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^
a
b
c
d Nusse, R. (NaN undefined NaN).
"Wnt Signaling". Cold Spring Harbor Perspectives in Biology. 4 (5): a011163.
doi:
10.1101/cshperspect.a011163.
PMC
3331700.
PMID
22550232.
{{ cite journal}}: Check date values in:|date=( help) -
^ Voronkov, A. E.; Baskin, I. I.; Palyulin, V. A.; Zefirov, N. S. (NaN undefined NaN).
"Molecular model of the Wnt protein binding site on the surface of dimeric CRD domain of the hFzd8 receptor". Doklady Biochemistry and Biophysics. 419 (1): 75–78.
doi:
10.1134/S1607672908020087.
PMID
18505162.
{{ cite journal}}: Check date values in:|date=( help) - ^ a b c d e f "Wnt Signaling Pathway". R and D Systems, Inc. Retrieved 18 November 2012.
-
^ Sakanaka, C (2002 Nov).
"Phosphorylation and regulation of beta-catenin by casein kinase I epsilon". Journal of Biochemistry. 132 (5): 697–703.
doi:
10.1093/oxfordjournals.jbchem.a003276.
PMID
[PubMed - indexed for MEDLINE 12417018 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Liu, Chunming; Kato, Yoichi; Zhang, Zhuohua; Do, Viet Minh; Yankner, Bruce A.; He, Xi (1999 May 25).
"beta-Trcp couples beta-catenin phosphorylation-degradation and regulates Xenopus axis formation". Proceedings of the National Academy of Sciences of the United States of America. 96 (11): 6273–8.
doi:
10.1073/pnas.96.11.6273.
PMID
[PubMed - indexed for MEDLINE 10339577 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^
a
b Smalley, MJ (15 November 2005).
"Dishevelled (Dvl-2) activates canonical Wnt signalling in the absence of cytoplasmic puncta". Journal of Cell Science. 118: 5278–5289.
doi:
10.1242/jcs.02647 (inactive 2023-08-02).
{{ cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) ( help); zero width space character in|doi=at position 9 ( help)CS1 maint: DOI inactive as of August 2023 ( link) -
^ Fiedler, M.; Mendoza-Topaz, C.; Rutherford, T. J.; Mieszczanek, J.; Bienz, M. (NaN undefined NaN).
"Dishevelled interacts with the DIX domain polymerization interface of Axin to interfere with its function in down-regulating -catenin". Proceedings of the National Academy of Sciences. 108 (5): 1937–1942.
doi:
10.1073/pnas.1017063108.
PMC
3033301.
PMID
21245303.
{{ cite journal}}: Check date values in:|date=and|year=/|date=mismatch ( help); Unknown parameter|month=ignored ( help) -
^ Li, L. (NaN undefined NaN).
"Axin and Frat1 interact with Dvl and GSK, bridging Dvl to GSK in Wnt-mediated regulation of LEF-1". The EMBO Journal. 18 (15): 4233–4240.
doi:
10.1093/emboj/18.15.4233.
PMC
1171499.
PMID
10428961.
{{ cite journal}}: Check date values in:|date=( help) - ^
a
b Lapébie, P (2011 Oct).
"Dissecting the PCP pathway: one or more pathways?: Does a separate Wnt-Fz-Rho pathway drive morphogenesis?". BioEssays : News and Reviews in Molecular, Cellular and Developmental Biology. 33 (10): 759–68.
doi:
10.1002/bies.201100023.
PMID
[PubMed - indexed for MEDLINE 21919026 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help); Unknown parameter|coauthors=ignored (|author=suggested) ( help) -
^ Zhang, WM (2005 Jun).
"Effect of WNT-1 on beta-catenin expression and its relation to Ki-67 and tumor differentiation in oral squamous cell carcinoma". Oncology Reports. 13 (6): 1095–9.
PMID
[PubMed - indexed for MEDLINE 15870927 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help); Unknown parameter|coauthors=ignored (|author=suggested) ( help) - ^
a
b Choi, A-Ram; Park, Jeong-Ran; Kim, Ran-Ju; Kim, Soo-Rim; Cho, Sung-Dae; Jung, Ji-Youn; Nam, Jeong-Seok (2012 Aug 24).
"Inhibition of Wnt1 expression reduces the enrichment of cancer stem cells in a mouse model of breast cancer". Biochemical and Biophysical Research Communications. 425 (2): 436–42.
doi:
10.1016/j.bbrc.2012.07.120.
PMID
[PubMed - in process 22846569 [PubMed - in process]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Huang, Cheng-Long; Liu, Dage; Ishikawa, Shinya; Nakashima, Takashi; Nakashima, Nariyasu; Yokomise, Hiroyasu; Kadota, Kyuichi; Ueno, Masaki (2008 Nov).
"Wnt1 overexpression promotes tumour progression in non-small cell lung cancer". European Journal of Cancer (Oxford, England : 1990). 44 (17): 2680–8.
doi:
10.1016/j.ejca.2008.08.004.
PMID
[PubMed - indexed for MEDLINE 18790633 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Katoh, M (2003 Jan).
"Expression and regulation of WNT1 in human cancer: up-regulation of WNT1 by beta-estradiol in MCF-7 cells". International Journal of Oncology. 22 (1): 209–12.
PMID
[PubMed - indexed for MEDLINE 12469206 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Minear, Steven; Leucht, Philipp; Jiang, Jie; Liu, Bo; Zeng, Arial; Fuerer, Christophe; Nusse, Roel; Helms, Jill A. (NaN undefined NaN). "Wnt Proteins Promote Bone Regeneration". Science Translational Medicine. 2 (29): 29ra30.
doi:
10.1126/scitranslmed.3000231.
PMID
20427820.
{{ cite journal}}: Check date values in:|date=( help) -
^ Niederleithner, Heide; Heinz, Magdalena; Tauber, Stefanie; Bilban, Martin; Pehamberger, Hubert; Sonderegger, Stefan; Knöfler, Martin; Bracher, Andreas; Berger, Walter; Loewe, Robert; Petzelbauer, Peter (2012 Sep).
"Wnt1 is anti-lymphangiogenic in a melanoma mouse model". The Journal of Investigative Dermatology. 132 (9): 2235–44.
doi:
10.1038/jid.2012.138.
PMID
[PubMed - indexed for MEDLINE 22572818 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Duan, Jinzhu; Gherghe, Costin; Liu, Dianxin; Hamlett, Eric; Srikantha, Luxman; Rodgers, Laurel; Regan, Jenna N.; Rojas, Mauricio; Willis, Monte; Leask, Andrew; Majesky, Mark; Deb, Arjun (2011 Nov 15).
"Wnt1/βcatenin injury response activates the epicardium and cardiac fibroblasts to promote cardiac repair". The EMBO Journal. 31 (2): 429–42.
doi:
10.1038/emboj.2011.418.
PMID
[PubMed - indexed for MEDLINE 22085926 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Sullivan, James C.; Ryan, Joseph F.; Mullikin, James C.; Finnerty, John R. (2007 Mar).
"Conserved and novel Wnt clusters in the basal eumetazoan Nematostella vectensis". Development Genes and Evolution. 217 (3): 235–9.
doi:
10.1007/s00427-007-0136-5.
PMID
[PubMed - indexed for MEDLINE 17310352 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Van Ooyen, A.; Kwee, V.; Nusse, R. (1985).
"The nucleotide sequence of the human int-1 mammary oncogene; evolutionary conservation of coding and non-coding sequences". EMBO. 4 (11): 2095–2909.
doi:
10.1002/j.1460-2075.1985.tb04021.x.
PMC
554596.
PMID
2998762.
{{ cite journal}}: Unknown parameter|month=ignored ( help)CS1 maint: date and year ( link) -
^ Kusserow, Arne; Pang, Kevin; Sturm, Carsten; Hrouda, Martina; Lentfer, Jan; Schmidt, Heiko A.; Technau, Ulrich; von Haeseler, Arndt; Hobmayer, Bert; Martindale, Mark Q.; Holstein, Thomas W. (2005 Jan 13).
"Unexpected complexity of the Wnt gene family in a sea anemone". Nature. 433 (7022): 156–60.
doi:
10.1038/nature03158.
PMID
[PubMed - indexed for MEDLINE 15650739 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help)
Proto-oncogene protein Wnt-1 protein, short for Wingless-Type MMTV Integration Site Family Member 1, is encoded by the Wnt-1 gene. [1] Wnt-1 is part of the Wnt-gene family, together forming a highly conserved secreted signaling pathway that regulate cell-to-cell interaction during embryogenesis. [2]
Wnt-1 is a secreted glycoprotein, usually around 350-400 amino acids in length with conserved pattern of 23-25 cysteine residues across species. [3]
During embryonic development, Wnt-1 plays a role in generation of cell polarity, body axis formation, [4] and development of the central nervous system. [5] It also has a role in the regeneration of tissues in adult organisms. [6] A mis-expression of Wnt-1 is correlated with tumorogenesis. [3]
Wnt signals through cell-surface receptors in the Frizzled and LRP families. [7] [2] Binding of Wnt protein to cell-surface receptor triggers a signaling cascade involving intracellular signaling protein beta-catenin, which leads to the increased expression of 50+ genes. [7]
Wnt-1 protein can be referred to as Int-1 or Wg in literature, varying by the organism studied and the time of research. Int-1 (Integration site-1) was first described by Nusse and Varmus in 1982 as the proto-oncogene activated by mouse mammary tumor virus (MMTV). [8] In 1976, the ortholog gene in Drosophila melanogaster was described as Wg (Wingless), from the wingless phenotype of mutants. [9] To reduce confusion, Int-1 and Wg were combined and renamed as Wnt-1 around the 1990s. [10]
Wnt-1 is required for proper formation of the midbrain and anterior hindbrain in the mouse. [5] Mutations of the Wnt-1 gene in embryonic mice results in a wide range of phenotypes: some die at birth lacking an entire cerebellum and parts of the midbrain, while some live through adulthood suffering from ataxia and lacking the anterior half of the cerebellum. [5] The malformed parts of the brain in Wnt-1 mutants correspond to not only the region where Wnt-1 would normally be highly expressed, but also the surrounding tissues. [11] This suggests Wnt-1 protein’s importance in inducing development of surrounding tissues. [12] No explanation for the variability of Wnt-1 mutant phenotype expressed is known, but it may be due to several factors, including genetic background. [13]
The role of Wnt-1 protein in body-axis formation was studied in the model organism Xenopus laevis. The injection of Wnt-1 mRNA into early Xenopus embryo results in duplicated dorsal axis. This suggests the role of Wnt-1 as an organizer molecule and embryonic signal inducer for pattern formation. [14]
Wnt proteins are involved in a multitude of pathways. In most pathways, Wnt is found to interact directly with transmembrane proteins of the Frizzled family. [15] Wnt1 is specifically found to interact with Fzd8. The binding of Wnt to Fzd8 requires a dimerization of two CRD domains of Fzd. [16]

In the absence of Wnt, β-catenin is phosphorylated and marked for degradation by ubiquitin mediated proteosomes. [17] This phosphorylation is catalyzed by serine/threonine glycogen synthase kinase-3beta (GSK-3β) in complex with casein kinase 1 (CK1), the adenomatous polyposis coli (APC) protein, axin and protein phosphatase 2A (PP2A). [15] [17]This process of phosphorylation by GSK-3 β occurs at serines-33 and -37, as well as at serine-45. [18] The process is continued by β-TrCP in conjunction with ubiquitin ligase in order to mark the protein with ubiquitin for degradation. [19]

A change in the cytoplasmic conformation of the proteins occurs when Wnt binds to the Fzd/LRP complex, resulting in the binding of Axin to LRP and the binding of the Dishevelled protein (Dsh) to Fzd. [20] Other transmembrane proteins found to interact with Fzd8 and assist in the signal transmission across the membrane are two low density lipoprotein receptor-related proteins, LRP5 and LRP6. [20] [15]The deactivation of GSK-3β is coupled to the interaction between axin and dishevelled (DIX) domain, thus preventing the formation of the axin dependant phosphorylation complex. [21] [22] This prevents phosphorylation of β-catenin and allows for its accumulation.
In the nucleus without β-catenin, two transcription factors,
T-cell factor (TCF) and
Lymphoid enhancer factor (LEF) in conjugation with the protein
Groucho, act as
inhibitors of Wnt controlled genes.
[17] Once build up of β-catenin occurs, it acts to from a complex with TCF/LEF. The complex then acts to induce the same genes that TCF/LEF inhibited in the absence of β-catenin, specifically those relating to
cell growth and
differentiation.
[17]
The two major non-canonical pathways are the planar cell polarity (PCP) pathway and the Wnt/Ca2+ pathway. Both of these pathways are also reliant on the Frizzled transmembrane protein. [17] It is suggested that the activation of these alternate pathways is dependent on the specific Fzd related membrane proteins present, such as LRP5 or LRP6. The Wnt/Ca2+ pathway is reliant on a reaction between dishevelled protein and Fzd, causing the activation of a G-protein coupled signal transduction cascade.The ultimate release of calcium from this mechanism activates calcium/calmodium-dependent protein kinase II (CaMKII). [17] In developmental processes, the release of such kinases results in a decrease in the effect of the canonical beta catenin pathway, and plays a role in cell migration and axis development. Planar Cell Polarity (PCP) is a pathway involving many of the same proteins such as Fzd and Dishevelled , but primarily plays a role in altering cell morphology. [23] This pathway involves the activation of Rho- and Rac-GTPases at the membrane, followed by the activation of different Myosins and Kinases which alter cell structure. [15] [23]
Wnt-1 mis-expression are of clinical significance since mutations are associated chronic diseases.
An overexpression of Wnt-1 is correlated with tumorogenesis. [3] Wnt-1 was first identified in 1982 by Nesse and Varmus as a proto-oncogene activated by integration of mouse mammary tumor virus (MMTV). [8] Today, the mis-expression of Wnt-1 is correlated with diseases such as oral squamous cell carcinoma (OSCC), [24] breast tumours in human breast cancer [25] (19), non-small cell lung cancer (NSCLC), [26] and gastric cancer. [27]
The up-regulation of Wnt proteins can enhance the process of osteocyte proliferation. [28] Injecting mice with purified Wnt3a protein resulted in proliferation of liposomes at the sites of bone damage. This not only suggests an additional area of therapeutic research using Wnt pathways, but also the potential positive effects of up-regulated Wnt protein expressions in tissue/organ regeneration. Wnt-1 over-expression can suppress the lymphangiogenesis and potentiates the delay of metastasis in melanoma. [29] In addition, Wnt-1 expression has shown to aid the restoration of cardiac functions by up-regulating the formation of new cardiac fibroblast cells, along the Wnt-1/β-catenin pathway in the cases of acute cardiac ischemia. [30]
Inhibition studies of the Wnt-gene family were long focused due to their cancer-causing properties when over-expressed. Over-expression of Wnt-1 has demonstrated a strong causational relationship to the formation of mammary gland tumor, [25] a deeper understanding of the specific Wnt-1 pathway in relevant organs will potentially bring a more effective treatment therapy for breast cancer.
The highly conserved nature of Wnt gene throughout evolution creates an opportunity for research to take place in multiple model organisms, including worms, flies, and mice. [31] [2] The human Wnt-1 protein sequence is 99% identical to that of the mouse homologue. [32] Wnt-1 is usually around 350-400 amino acids in length with conserved pattern of 23-25 cysteine residues across species. [3]
Evolutionary biologists speculate the early amplification and divergence of the Wnt family were the cause for increasing complexity of animal body plans. Wnt-1 has been found conserved throughout the entire animal kingdom, often expressed in regionalized patterns. [33]
- ^ "WNT1 wingless-type MMTV integration site family, member 1 [homo sapiens]". NCBI. Retrieved 10 November 2012.
- ^ a b c Nusse, Roel. "The Wnt Homepage". Retrieved 10 November 2012.
- ^
a
b
c
d Cardigan, Ken (1997).
"Wnt signaling: a common theme in animal development". Genes & Development. 11: 3286–3305.
doi:
10.1101/gad.11.24.3286 Genes & Dev. 1997. 11: 3286-3305 (inactive 2023-08-02). Retrieved 17 November 2012.
{{ cite journal}}: Check|doi=value ( help); Unknown parameter|coauthors=ignored (|author=suggested) ( help)CS1 maint: DOI inactive as of August 2023 ( link) -
^ McMahon, Andrew P.; Moon, Randall T. (1989 Sep 22).
"Ectopic expression of the proto-oncogene int-1 in Xenopus embryos leads to duplication of the embryonic axis". Cell. 58 (6): 1075–84.
doi:
10.1016/0092-8674(89)90506-0.
PMID
[PubMed - indexed for MEDLINE 2673541 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^
a
b
c Thomas, Kirk R.; Capecchi, Mario R. (NaN undefined NaN).
"Targeted disruption of the murine int-1 proto-oncogene resulting in severe abnormalities in midbrain and cerebellar development". Nature. 346 (6287): 847–850.
doi:
10.1038/346847a0.
PMID
2202907.
{{ cite journal}}: Check date values in:|date=( help) -
^ Gurley, Kyle A.; Elliott, Sarah A.; Simakov, Oleg; Schmidt, Heiko A.; Holstein, Thomas W.; Alvarado, Alejandro Sánchez (2010 Nov 1).
"Expression of secreted Wnt pathway components reveals unexpected complexity of the planarian amputation response". Developmental Biology. 347 (1): 24–39.
doi:
10.1016/j.ydbio.2010.08.007.
PMID
[PubMed - indexed for MEDLINE 20707997 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^ a b "Wnt1 protein (30R-AW002)". Fitzgerald-FII Supplier of Antibodies Protein and ELISA kits. Retrieved 18 November 2012.
- ^
a
b Nusse, Roel; Varmus, Harold E. (November 1982).
"Many tumors induced by the mouse mammary tumor virus contain a provirus integrated in the same region of the host genome". Cell. 31 (1): 99–109.
doi:
10.1016/0092-8674(82)90409-3.
PMID
6297757.
{{ cite journal}}: CS1 maint: date and year ( link) -
^ Sharma, R.P.; Chopra, V.L. (Feb).
"Effect of the Wingless (wg1) mutation on wing and haltere development in Drosophila melanogaster". Developmental Biology. 48 (2): 461–5.
doi:
10.1016/0012-1606(76)90108-1.
PMID
[PubMed - indexed for MEDLINE 815114 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=and|year=/|date=mismatch ( help) -
^ Nusse, Roel; Varmus, Harold (2012 May 22).
"Three decades of Wnts: a personal perspective on how a scientific field developed". The EMBO Journal. 31 (12): 2670–84.
doi:
10.1038/emboj.2012.146.
PMID
[PubMed - indexed for MEDLINE 22617420 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ McMahon, Andrew P.; Bradley, Allan (NaN undefined NaN).
"The Wnt-1 (int-1) proto-oncogene is required for development of a large region of the mouse brain". Cell. 62 (6): 1073–1085.
doi:
10.1016/0092-8674(90)90385-R.
PMID
2205396.
{{ cite journal}}: Check date values in:|date=( help) -
^ Nusse, R (1992 Sep).
"The Wnt gene family in tumorigenesis and in normal development". The Journal of Steroid Biochemistry and Molecular Biology. 43 (1–3): 9–12.
doi:
10.1016/0960-0760(92)90181-H.
PMID
[PubMed - indexed for MEDLINE 1388050 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Nusse, R.; Varmus, H. E. (26).
"Wnt Genes Review" (PDF). Cell. 69 (7): 1073–1087.
doi:
10.1016/0092-8674(92)90630-U.
PMID
1617723.
{{ cite journal}}: Check date values in:|date=and|year=/|date=mismatch ( help); Unknown parameter|month=ignored ( help) -
^ McMahon, Andrew P.; Moon, Randall T. (1989 Sep 22).
"Ectopic expression of the proto-oncogene int-1 in Xenopus embryos leads to duplication of the embryonic axis". Cell. 58 (6): 1075–84.
doi:
10.1016/0092-8674(89)90506-0.
PMID
[PubMed - indexed for MEDLINE 2673541 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^
a
b
c
d Nusse, R. (NaN undefined NaN).
"Wnt Signaling". Cold Spring Harbor Perspectives in Biology. 4 (5): a011163.
doi:
10.1101/cshperspect.a011163.
PMC
3331700.
PMID
22550232.
{{ cite journal}}: Check date values in:|date=( help) -
^ Voronkov, A. E.; Baskin, I. I.; Palyulin, V. A.; Zefirov, N. S. (NaN undefined NaN).
"Molecular model of the Wnt protein binding site on the surface of dimeric CRD domain of the hFzd8 receptor". Doklady Biochemistry and Biophysics. 419 (1): 75–78.
doi:
10.1134/S1607672908020087.
PMID
18505162.
{{ cite journal}}: Check date values in:|date=( help) - ^ a b c d e f "Wnt Signaling Pathway". R and D Systems, Inc. Retrieved 18 November 2012.
-
^ Sakanaka, C (2002 Nov).
"Phosphorylation and regulation of beta-catenin by casein kinase I epsilon". Journal of Biochemistry. 132 (5): 697–703.
doi:
10.1093/oxfordjournals.jbchem.a003276.
PMID
[PubMed - indexed for MEDLINE 12417018 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Liu, Chunming; Kato, Yoichi; Zhang, Zhuohua; Do, Viet Minh; Yankner, Bruce A.; He, Xi (1999 May 25).
"beta-Trcp couples beta-catenin phosphorylation-degradation and regulates Xenopus axis formation". Proceedings of the National Academy of Sciences of the United States of America. 96 (11): 6273–8.
doi:
10.1073/pnas.96.11.6273.
PMID
[PubMed - indexed for MEDLINE 10339577 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) - ^
a
b Smalley, MJ (15 November 2005).
"Dishevelled (Dvl-2) activates canonical Wnt signalling in the absence of cytoplasmic puncta". Journal of Cell Science. 118: 5278–5289.
doi:
10.1242/jcs.02647 (inactive 2023-08-02).
{{ cite journal}}: Unknown parameter|coauthors=ignored (|author=suggested) ( help); zero width space character in|doi=at position 9 ( help)CS1 maint: DOI inactive as of August 2023 ( link) -
^ Fiedler, M.; Mendoza-Topaz, C.; Rutherford, T. J.; Mieszczanek, J.; Bienz, M. (NaN undefined NaN).
"Dishevelled interacts with the DIX domain polymerization interface of Axin to interfere with its function in down-regulating -catenin". Proceedings of the National Academy of Sciences. 108 (5): 1937–1942.
doi:
10.1073/pnas.1017063108.
PMC
3033301.
PMID
21245303.
{{ cite journal}}: Check date values in:|date=and|year=/|date=mismatch ( help); Unknown parameter|month=ignored ( help) -
^ Li, L. (NaN undefined NaN).
"Axin and Frat1 interact with Dvl and GSK, bridging Dvl to GSK in Wnt-mediated regulation of LEF-1". The EMBO Journal. 18 (15): 4233–4240.
doi:
10.1093/emboj/18.15.4233.
PMC
1171499.
PMID
10428961.
{{ cite journal}}: Check date values in:|date=( help) - ^
a
b Lapébie, P (2011 Oct).
"Dissecting the PCP pathway: one or more pathways?: Does a separate Wnt-Fz-Rho pathway drive morphogenesis?". BioEssays : News and Reviews in Molecular, Cellular and Developmental Biology. 33 (10): 759–68.
doi:
10.1002/bies.201100023.
PMID
[PubMed - indexed for MEDLINE 21919026 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help); Unknown parameter|coauthors=ignored (|author=suggested) ( help) -
^ Zhang, WM (2005 Jun).
"Effect of WNT-1 on beta-catenin expression and its relation to Ki-67 and tumor differentiation in oral squamous cell carcinoma". Oncology Reports. 13 (6): 1095–9.
PMID
[PubMed - indexed for MEDLINE 15870927 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help); Unknown parameter|coauthors=ignored (|author=suggested) ( help) - ^
a
b Choi, A-Ram; Park, Jeong-Ran; Kim, Ran-Ju; Kim, Soo-Rim; Cho, Sung-Dae; Jung, Ji-Youn; Nam, Jeong-Seok (2012 Aug 24).
"Inhibition of Wnt1 expression reduces the enrichment of cancer stem cells in a mouse model of breast cancer". Biochemical and Biophysical Research Communications. 425 (2): 436–42.
doi:
10.1016/j.bbrc.2012.07.120.
PMID
[PubMed - in process 22846569 [PubMed - in process]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Huang, Cheng-Long; Liu, Dage; Ishikawa, Shinya; Nakashima, Takashi; Nakashima, Nariyasu; Yokomise, Hiroyasu; Kadota, Kyuichi; Ueno, Masaki (2008 Nov).
"Wnt1 overexpression promotes tumour progression in non-small cell lung cancer". European Journal of Cancer (Oxford, England : 1990). 44 (17): 2680–8.
doi:
10.1016/j.ejca.2008.08.004.
PMID
[PubMed - indexed for MEDLINE 18790633 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Katoh, M (2003 Jan).
"Expression and regulation of WNT1 in human cancer: up-regulation of WNT1 by beta-estradiol in MCF-7 cells". International Journal of Oncology. 22 (1): 209–12.
PMID
[PubMed - indexed for MEDLINE 12469206 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Minear, Steven; Leucht, Philipp; Jiang, Jie; Liu, Bo; Zeng, Arial; Fuerer, Christophe; Nusse, Roel; Helms, Jill A. (NaN undefined NaN). "Wnt Proteins Promote Bone Regeneration". Science Translational Medicine. 2 (29): 29ra30.
doi:
10.1126/scitranslmed.3000231.
PMID
20427820.
{{ cite journal}}: Check date values in:|date=( help) -
^ Niederleithner, Heide; Heinz, Magdalena; Tauber, Stefanie; Bilban, Martin; Pehamberger, Hubert; Sonderegger, Stefan; Knöfler, Martin; Bracher, Andreas; Berger, Walter; Loewe, Robert; Petzelbauer, Peter (2012 Sep).
"Wnt1 is anti-lymphangiogenic in a melanoma mouse model". The Journal of Investigative Dermatology. 132 (9): 2235–44.
doi:
10.1038/jid.2012.138.
PMID
[PubMed - indexed for MEDLINE 22572818 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Duan, Jinzhu; Gherghe, Costin; Liu, Dianxin; Hamlett, Eric; Srikantha, Luxman; Rodgers, Laurel; Regan, Jenna N.; Rojas, Mauricio; Willis, Monte; Leask, Andrew; Majesky, Mark; Deb, Arjun (2011 Nov 15).
"Wnt1/βcatenin injury response activates the epicardium and cardiac fibroblasts to promote cardiac repair". The EMBO Journal. 31 (2): 429–42.
doi:
10.1038/emboj.2011.418.
PMID
[PubMed - indexed for MEDLINE 22085926 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Sullivan, James C.; Ryan, Joseph F.; Mullikin, James C.; Finnerty, John R. (2007 Mar).
"Conserved and novel Wnt clusters in the basal eumetazoan Nematostella vectensis". Development Genes and Evolution. 217 (3): 235–9.
doi:
10.1007/s00427-007-0136-5.
PMID
[PubMed - indexed for MEDLINE 17310352 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help) -
^ Van Ooyen, A.; Kwee, V.; Nusse, R. (1985).
"The nucleotide sequence of the human int-1 mammary oncogene; evolutionary conservation of coding and non-coding sequences". EMBO. 4 (11): 2095–2909.
doi:
10.1002/j.1460-2075.1985.tb04021.x.
PMC
554596.
PMID
2998762.
{{ cite journal}}: Unknown parameter|month=ignored ( help)CS1 maint: date and year ( link) -
^ Kusserow, Arne; Pang, Kevin; Sturm, Carsten; Hrouda, Martina; Lentfer, Jan; Schmidt, Heiko A.; Technau, Ulrich; von Haeseler, Arndt; Hobmayer, Bert; Martindale, Mark Q.; Holstein, Thomas W. (2005 Jan 13).
"Unexpected complexity of the Wnt gene family in a sea anemone". Nature. 433 (7022): 156–60.
doi:
10.1038/nature03158.
PMID
[PubMed - indexed for MEDLINE 15650739 [PubMed - indexed for MEDLINE]].
{{ cite journal}}: Check|pmid=value ( help); Check date values in:|date=( help)
