Recombination signal sequences are conserved sequences of noncoding DNA that are recognized by the RAG1/RAG2 enzyme complex during V(D)J recombination in immature B cells and T cells. [1] Recombination signal sequences guide the enzyme complex to the V, D, and J gene segments that will undergo recombination during the formation of the heavy and light-chain variable regions in T-cell receptors and immunoglobulin molecules. [1]
Structure
RSSs are made up of highly conserved
heptamer sequences (7 base pairs),
spacer sequences, and conserved
nonamer sequences (9 base pairs) that are adjacent to the V, D and J sequences in the
heavy-chain region of DNA and the V and J sequences in the
light-chain DNA region.
[1]
[2] Spacer sequences are located between heptamer and nonamer sequences and exhibit base pair variety but are always either 12 base pairs or 23 base pairs long.
[3] Heptamer sequences are usually CACAGTG and nonamers are usually ACAAAAACC. The nucleotides in bold are more highly conserved.
[3] The RAG1/RAG2 enzyme complex follows the 12-23 rule when joining V, D, and J segments, pairing 12-bp spacer RSSs to 23-bp spacer RSSs.
[1]
[2] This prevents two different genes coding for the same region from recombining (ex. V-V recombination).
[1] RSSs are located between V, D, and J segments of the germ-line DNA of maturing B and T lymphocytes and are permanently spliced out of the final Ig mRNA product after V(D)J recombination is complete.
[1]
Function
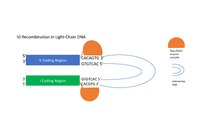
The RAG1/RAG2 enzyme complex recognizes the heptamer sequences flanking the V and J coding regions and nicks their 5' end, releasing the intervening DNA between the V and J coding regions. [1] In the heavy-chain coding region of DNA, the RAG1/RAG2 enzyme complex recognizes the RSSs flanking the D and J segments and brings them together, forming a loop containing intervening DNA. [1] [4] The RAG1/RAG2 complex then introduces a nick at the 5' end of the RSS heptamers adjacent to the coding regions on both the D and J segments, permanently removing the loop of intervening DNA and creating a double-stranded break that is repaired by VDJ recombinase enzymes. [1] [4] This process is repeated for the joining of V to DJ. [1] In light-chain rearrangement, only V and J segments are brought together. [1]
Related Diseases & Disorders
cRSS
Cryptic RSSs are gene sequences that resemble authentic RSSs and are occasionally mistaken for them by the RAG1/RAG2 enzyme complex. [3] Recombining an RSS with a cRSS can lead to chromosome translocations, which can lead to cancer. [3]
Omenn's Syndrome
Some infants born with autosomal recessive SCIDS lack a functional copies of the genes that code for the RAG1/RAG2 enzyme complex because of missense mutations. [5] [6] These infants will produce a non-functional RAG1/RAG2 enzyme complex that cannot recognize RSSs and therefore cannot initiate V(D)J recombination effectively. [5] [6] This disorder is characterized by a lack of functioning B and T cells. [1] [5]
References
- ^ a b c d e f g h i j k l Owen, Judith A.; Punt, Jenni; Stranford, Sharon A. (2013). Kuby Immunology. New York: W.H. Freeman and Company. pp. 233–235.
- ^ a b Schatz, David G.; Oettinger, Marjorie A.; Schlissel, Mark S. (1992). "V(D)J Recombination: Molecular Biology and Regulation". Annual Review of Immunology. 10: 359–383. doi: 10.1146/annurev.immunol.10.1.359. PMID 1590991.
- ^ a b c d Roth, David B. (2014). "V(D)J Recombination: Mechanism, Errors, and Fidelity". Microbiology Spectrum. 2 (6): 1–11. doi: 10.1128/microbiolspec.MDNA3-0041-2014. ISBN 9781555819200. PMC 5089068. PMID 26104458.
- ^ a b Rodgers, Karla (2017). "Riches in RAGs: Revealing the V(D)J Recombinase Through High-Resolution Structures". Trends in Biochemical Sciences. 42 (1): 72–84. doi: 10.1016/j.tibs.2016.10.003. PMC 5182142. PMID 27825771.
- ^ a b c Buckley, Rebecca H. (2004). "Molecular Defects in Human Severe Combined Immunodeficiency And Approaches to Immune Reconstitution". Annual Review of Immunology. 22: 625–655. doi: 10.1146/annurev.immunol.22.012703.104614. PMID 15032591.
- ^ a b Elnour, Ibtisam B; Ahmed, Shakeel; Halim, Kamel; Nirmala, V (2007). "Omenn's Syndrome: A rare primary immunodeficiency disorder". Sultan Qaboos University Medical Journal. 7 (2): 1–6. PMC 3074865. PMID 21748095.
Recombination signal sequences are conserved sequences of noncoding DNA that are recognized by the RAG1/RAG2 enzyme complex during V(D)J recombination in immature B cells and T cells. [1] Recombination signal sequences guide the enzyme complex to the V, D, and J gene segments that will undergo recombination during the formation of the heavy and light-chain variable regions in T-cell receptors and immunoglobulin molecules. [1]
Structure
RSSs are made up of highly conserved
heptamer sequences (7 base pairs),
spacer sequences, and conserved
nonamer sequences (9 base pairs) that are adjacent to the V, D and J sequences in the
heavy-chain region of DNA and the V and J sequences in the
light-chain DNA region.
[1]
[2] Spacer sequences are located between heptamer and nonamer sequences and exhibit base pair variety but are always either 12 base pairs or 23 base pairs long.
[3] Heptamer sequences are usually CACAGTG and nonamers are usually ACAAAAACC. The nucleotides in bold are more highly conserved.
[3] The RAG1/RAG2 enzyme complex follows the 12-23 rule when joining V, D, and J segments, pairing 12-bp spacer RSSs to 23-bp spacer RSSs.
[1]
[2] This prevents two different genes coding for the same region from recombining (ex. V-V recombination).
[1] RSSs are located between V, D, and J segments of the germ-line DNA of maturing B and T lymphocytes and are permanently spliced out of the final Ig mRNA product after V(D)J recombination is complete.
[1]
Function

The RAG1/RAG2 enzyme complex recognizes the heptamer sequences flanking the V and J coding regions and nicks their 5' end, releasing the intervening DNA between the V and J coding regions. [1] In the heavy-chain coding region of DNA, the RAG1/RAG2 enzyme complex recognizes the RSSs flanking the D and J segments and brings them together, forming a loop containing intervening DNA. [1] [4] The RAG1/RAG2 complex then introduces a nick at the 5' end of the RSS heptamers adjacent to the coding regions on both the D and J segments, permanently removing the loop of intervening DNA and creating a double-stranded break that is repaired by VDJ recombinase enzymes. [1] [4] This process is repeated for the joining of V to DJ. [1] In light-chain rearrangement, only V and J segments are brought together. [1]
Related Diseases & Disorders
cRSS
Cryptic RSSs are gene sequences that resemble authentic RSSs and are occasionally mistaken for them by the RAG1/RAG2 enzyme complex. [3] Recombining an RSS with a cRSS can lead to chromosome translocations, which can lead to cancer. [3]
Omenn's Syndrome
Some infants born with autosomal recessive SCIDS lack a functional copies of the genes that code for the RAG1/RAG2 enzyme complex because of missense mutations. [5] [6] These infants will produce a non-functional RAG1/RAG2 enzyme complex that cannot recognize RSSs and therefore cannot initiate V(D)J recombination effectively. [5] [6] This disorder is characterized by a lack of functioning B and T cells. [1] [5]
References
- ^ a b c d e f g h i j k l Owen, Judith A.; Punt, Jenni; Stranford, Sharon A. (2013). Kuby Immunology. New York: W.H. Freeman and Company. pp. 233–235.
- ^ a b Schatz, David G.; Oettinger, Marjorie A.; Schlissel, Mark S. (1992). "V(D)J Recombination: Molecular Biology and Regulation". Annual Review of Immunology. 10: 359–383. doi: 10.1146/annurev.immunol.10.1.359. PMID 1590991.
- ^ a b c d Roth, David B. (2014). "V(D)J Recombination: Mechanism, Errors, and Fidelity". Microbiology Spectrum. 2 (6): 1–11. doi: 10.1128/microbiolspec.MDNA3-0041-2014. ISBN 9781555819200. PMC 5089068. PMID 26104458.
- ^ a b Rodgers, Karla (2017). "Riches in RAGs: Revealing the V(D)J Recombinase Through High-Resolution Structures". Trends in Biochemical Sciences. 42 (1): 72–84. doi: 10.1016/j.tibs.2016.10.003. PMC 5182142. PMID 27825771.
- ^ a b c Buckley, Rebecca H. (2004). "Molecular Defects in Human Severe Combined Immunodeficiency And Approaches to Immune Reconstitution". Annual Review of Immunology. 22: 625–655. doi: 10.1146/annurev.immunol.22.012703.104614. PMID 15032591.
- ^ a b Elnour, Ibtisam B; Ahmed, Shakeel; Halim, Kamel; Nirmala, V (2007). "Omenn's Syndrome: A rare primary immunodeficiency disorder". Sultan Qaboos University Medical Journal. 7 (2): 1–6. PMC 3074865. PMID 21748095.