Within the field of molecular biology, a protein-fragment complementation assay, or PCA, is a method for the identification and quantification of protein–protein interactions. In the PCA, the proteins of interest ("bait" and "prey") are each covalently linked to fragments of a third protein (e.g. DHFR, which acts as a "reporter"). Interaction between the bait and the prey proteins brings the fragments of the reporter protein in close proximity to allow them to form a functional reporter protein whose activity can be measured. This principle can be applied to many different reporter proteins and is also the basis for the yeast two-hybrid system, an archetypical PCA assay.
Split protein assays
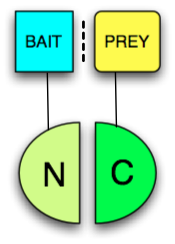
Any protein that can be split into two parts and reconstituted non-covalently to form a functional protein may be used in a PCA. The two fragments however have low affinity for each other and must be brought together by other interacting proteins fused to them (often called "bait" and "prey" since the bait protein can be used to identify a prey protein, see figure). The protein that produces a detectable readout is called "reporter". Usually enzymes which confer resistance to nutrient deprivation or antibiotics, such as dihydrofolate reductase or beta-lactamase respectively, or proteins that give colorimetric or fluorescent signals are used as reporters. When fluorescent proteins are reconstituted the PCA is called Bimolecular fluorescence complementation assay. The following proteins have been used in split protein PCAs:
- Beta-lactamase [1] [2]
- Dihydrofolate reductase (DHFR) [3]
- Focal adhesion kinase (FAK) [4]
- Gal4, a yeast transcription factor (as in the classical yeast two-hybrid system)
- GFP (split-GFP), e.g. EGFP (enhanced green fluorescent protein) [5] [6] [7]
- Horseradish peroxidase [8]
- Infrared fluorescent protein IFP1.4, an engineered chromophore-binding domain (CBD) of a bacteriophytochrome from Deinococcus radiodurans [9]
- LacZ ( beta-galactosidase) [10]
- Luciferase, [11] [12] including ReBiL (recombinase enhanced bimolecular luciferase) [13] and Gaussia princeps luciferase. [14] Commercial products using luciferase include NanoLuc and NanoBIT. [15] A modification has also been developed for lipid droplet-associated interactions. [16]
- TEV ( Tobacco etch virus protease) [17]
- Ubiquitin [18]
Genome-wide applications
The methods mentioned above have been applied to whole genomes, e.g. yeast [3] or syphilis bacteria. [19]
References
- ^ Park JH, Back JH, Hahm SH, Shim HY, Park MJ, Ko SI, Han YS (October 2007). "Bacterial beta-lactamase fragmentation complementation strategy can be used as a method for identifying interacting protein pairs". Journal of Microbiology and Biotechnology. 17 (10): 1607–15. PMID 18156775.
- ^ Remy I, Ghaddar G, Michnick SW (2007). "Using the beta-lactamase protein-fragment complementation assay to probe dynamic protein-protein interactions". Nature Protocols. 2 (9): 2302–6. doi: 10.1038/nprot.2007.356. PMID 17853887. S2CID 7607566.
- ^ a b Tarassov K, Messier V, Landry CR, Radinovic S, Serna Molina MM, Shames I, Malitskaya Y, Vogel J, Bussey H, Michnick SW (June 2008). "An in vivo map of the yeast protein interactome" (PDF). Science. 320 (5882): 1465–70. Bibcode: 2008Sci...320.1465T. doi: 10.1126/science.1153878. PMID 18467557. S2CID 1732896.
- ^ Ma Y, Nagamune T, Kawahara M (September 2014). "Split focal adhesion kinase for probing protein–protein interactions". Biochemical Engineering Journal. 90: 272–278. Bibcode: 2014BioEJ..90..272M. doi: 10.1016/j.bej.2014.06.022.
- ^ Barnard E, Timson DJ (2010). "Split-EGFP Screens for the Detection and Localisation of Protein–Protein Interactions in Living Yeast Cells". Molecular and Cell Biology Methods for Fungi. Methods in Molecular Biology. Vol. 638. pp. 303–17. doi: 10.1007/978-1-60761-611-5_23. ISBN 978-1-60761-610-8. PMID 20238279.
- ^ Blakeley BD, Chapman AM, McNaughton BR (August 2012). "Split-superpositive GFP reassembly is a fast, efficient, and robust method for detecting protein-protein interactions in vivo". Molecular BioSystems. 8 (8): 2036–40. doi: 10.1039/c2mb25130b. PMID 22692102.
- ^ Cabantous S, Nguyen HB, Pedelacq JD, Koraïchi F, Chaudhary A, Ganguly K, Lockard MA, Favre G, Terwilliger TC, Waldo GS (October 2013). "A new protein-protein interaction sensor based on tripartite split-GFP association". Scientific Reports. 3: 2854. Bibcode: 2013NatSR...3E2854C. doi: 10.1038/srep02854. PMC 3790201. PMID 24092409.
- ^ Martell JD, Yamagata M, Deerinck TJ, Phan S, Kwa CG, Ellisman MH, Sanes JR, Ting AY (July 2016). "A split horseradish peroxidase for the detection of intercellular protein-protein interactions and sensitive visualization of synapses" (PDF). Nature Biotechnology. 34 (7): 774–80. doi: 10.1038/nbt.3563. PMC 4942342. PMID 27240195.
- ^ Tchekanda E, Sivanesan D, Michnick SW (June 2014). "An infrared reporter to detect spatiotemporal dynamics of protein-protein interactions". Nature Methods. 11 (6): 641–4. doi: 10.1038/nmeth.2934. PMID 24747815. S2CID 1958433.
- ^ Rossi F, Charlton CA, Blau HM (August 1997). "Monitoring protein-protein interactions in intact eukaryotic cells by beta-galactosidase complementation". Proceedings of the National Academy of Sciences of the United States of America. 94 (16): 8405–10. Bibcode: 1997PNAS...94.8405R. doi: 10.1073/pnas.94.16.8405. PMC 22934. PMID 9237989.
- ^ Cassonnet P, Rolloy C, Neveu G, Vidalain PO, Chantier T, Pellet J, Jones L, Muller M, Demeret C, Gaud G, Vuillier F, Lotteau V, Tangy F, Favre M, Jacob Y (November 2011). "Benchmarking a luciferase complementation assay for detecting protein complexes". Nature Methods. 8 (12): 990–2. doi: 10.1038/nmeth.1773. PMID 22127214. S2CID 9377872.
- ^ Fujikawa, Y. et al. (2014) Split luciferase complementation assay to detect regulated protein-protein interactions in rice protoplasts in a large-scale format. Rice 7:11
- ^ Li YC, Rodewald LW, Hoppmann C, Wong ET, Lebreton S, Safar P, Patek M, Wang L, Wertman KF, Wahl GM (December 2014). "A versatile platform to analyze low-affinity and transient protein-protein interactions in living cells in real time". Cell Reports. 9 (5): 1946–58. doi: 10.1016/j.celrep.2014.10.058. PMC 4269221. PMID 25464845.
- ^ Neveu G, Cassonnet P, Vidalain PO, Rolloy C, Mendoza J, Jones L, Tangy F, Muller M, Demeret C, Tafforeau L, Lotteau V, Rabourdin-Combe C, Travé G, Dricot A, Hill DE, Vidal M, Favre M, Jacob Y (December 2012). "Comparative analysis of virus-host interactomes with a mammalian high-throughput protein complementation assay based on Gaussia princeps luciferase". Methods. 58 (4): 349–59. doi: 10.1016/j.ymeth.2012.07.029. PMC 3546263. PMID 22898364.
- ^ Binkowski B, Eggers C, Butler B, Schwinn M, Slater M, Machleidt T, Cong M, Wood K, Fan F (May 2016). "Monitoring intracellular protein interactions using NanoLuc® Binary Technology (NanoBiTTM)" (PDF). Promega.
- ^ Kolkhof P, Werthebach M, van de Venn A, Poschmann G, Chen L, Welte M, Stühler K, Beller M (March 2017). "A Luciferase-fragment Complementation Assay to Detect Lipid Droplet-associated Protein-Protein Interactions". Molecular & Cellular Proteomics. 16 (3): 329–345. doi: 10.1074/mcp.M116.061499. PMC 5340998. PMID 27956707.
- ^ Wehr MC, Laage R, Bolz U, Fischer TM, Grünewald S, Scheek S, Bach A, Nave KA, Rossner MJ (December 2006). "Monitoring regulated protein-protein interactions using split TEV". Nature Methods. 3 (12): 985–93. doi: 10.1038/nmeth967. PMID 17072307. S2CID 37120401.
- ^ Dünkler A, Müller J, Johnsson N (2012). "Detecting Protein–Protein Interactions with the Split-Ubiquitin Sensor". Gene Regulatory Networks. Methods in Molecular Biology. Vol. 786. pp. 115–30. doi: 10.1007/978-1-61779-292-2_7. ISBN 978-1-61779-291-5. PMID 21938623.
- ^ Titz, Björn; Rajagopala, Seesandra V.; Goll, Johannes; Häuser, Roman; McKevitt, Matthew T.; Palzkill, Timothy; Uetz, Peter (2008-05-28). "The binary protein interactome of Treponema pallidum--the syphilis spirochete". PLOS ONE. 3 (5): e2292. Bibcode: 2008PLoSO...3.2292T. doi: 10.1371/journal.pone.0002292. ISSN 1932-6203. PMC 2386257. PMID 18509523.
Further reading
- Rochette S, Diss G, Filteau M, Leducq JB, Dubé AK, Landry CR (March 2015). "Genome-wide protein-protein interaction screening by protein-fragment complementation assay (PCA) in living cells". Journal of Visualized Experiments (97). doi: 10.3791/52255. PMC 4401175. PMID 25867901.
Within the field of molecular biology, a protein-fragment complementation assay, or PCA, is a method for the identification and quantification of protein–protein interactions. In the PCA, the proteins of interest ("bait" and "prey") are each covalently linked to fragments of a third protein (e.g. DHFR, which acts as a "reporter"). Interaction between the bait and the prey proteins brings the fragments of the reporter protein in close proximity to allow them to form a functional reporter protein whose activity can be measured. This principle can be applied to many different reporter proteins and is also the basis for the yeast two-hybrid system, an archetypical PCA assay.
Split protein assays

Any protein that can be split into two parts and reconstituted non-covalently to form a functional protein may be used in a PCA. The two fragments however have low affinity for each other and must be brought together by other interacting proteins fused to them (often called "bait" and "prey" since the bait protein can be used to identify a prey protein, see figure). The protein that produces a detectable readout is called "reporter". Usually enzymes which confer resistance to nutrient deprivation or antibiotics, such as dihydrofolate reductase or beta-lactamase respectively, or proteins that give colorimetric or fluorescent signals are used as reporters. When fluorescent proteins are reconstituted the PCA is called Bimolecular fluorescence complementation assay. The following proteins have been used in split protein PCAs:
- Beta-lactamase [1] [2]
- Dihydrofolate reductase (DHFR) [3]
- Focal adhesion kinase (FAK) [4]
- Gal4, a yeast transcription factor (as in the classical yeast two-hybrid system)
- GFP (split-GFP), e.g. EGFP (enhanced green fluorescent protein) [5] [6] [7]
- Horseradish peroxidase [8]
- Infrared fluorescent protein IFP1.4, an engineered chromophore-binding domain (CBD) of a bacteriophytochrome from Deinococcus radiodurans [9]
- LacZ ( beta-galactosidase) [10]
- Luciferase, [11] [12] including ReBiL (recombinase enhanced bimolecular luciferase) [13] and Gaussia princeps luciferase. [14] Commercial products using luciferase include NanoLuc and NanoBIT. [15] A modification has also been developed for lipid droplet-associated interactions. [16]
- TEV ( Tobacco etch virus protease) [17]
- Ubiquitin [18]
Genome-wide applications
The methods mentioned above have been applied to whole genomes, e.g. yeast [3] or syphilis bacteria. [19]
References
- ^ Park JH, Back JH, Hahm SH, Shim HY, Park MJ, Ko SI, Han YS (October 2007). "Bacterial beta-lactamase fragmentation complementation strategy can be used as a method for identifying interacting protein pairs". Journal of Microbiology and Biotechnology. 17 (10): 1607–15. PMID 18156775.
- ^ Remy I, Ghaddar G, Michnick SW (2007). "Using the beta-lactamase protein-fragment complementation assay to probe dynamic protein-protein interactions". Nature Protocols. 2 (9): 2302–6. doi: 10.1038/nprot.2007.356. PMID 17853887. S2CID 7607566.
- ^ a b Tarassov K, Messier V, Landry CR, Radinovic S, Serna Molina MM, Shames I, Malitskaya Y, Vogel J, Bussey H, Michnick SW (June 2008). "An in vivo map of the yeast protein interactome" (PDF). Science. 320 (5882): 1465–70. Bibcode: 2008Sci...320.1465T. doi: 10.1126/science.1153878. PMID 18467557. S2CID 1732896.
- ^ Ma Y, Nagamune T, Kawahara M (September 2014). "Split focal adhesion kinase for probing protein–protein interactions". Biochemical Engineering Journal. 90: 272–278. Bibcode: 2014BioEJ..90..272M. doi: 10.1016/j.bej.2014.06.022.
- ^ Barnard E, Timson DJ (2010). "Split-EGFP Screens for the Detection and Localisation of Protein–Protein Interactions in Living Yeast Cells". Molecular and Cell Biology Methods for Fungi. Methods in Molecular Biology. Vol. 638. pp. 303–17. doi: 10.1007/978-1-60761-611-5_23. ISBN 978-1-60761-610-8. PMID 20238279.
- ^ Blakeley BD, Chapman AM, McNaughton BR (August 2012). "Split-superpositive GFP reassembly is a fast, efficient, and robust method for detecting protein-protein interactions in vivo". Molecular BioSystems. 8 (8): 2036–40. doi: 10.1039/c2mb25130b. PMID 22692102.
- ^ Cabantous S, Nguyen HB, Pedelacq JD, Koraïchi F, Chaudhary A, Ganguly K, Lockard MA, Favre G, Terwilliger TC, Waldo GS (October 2013). "A new protein-protein interaction sensor based on tripartite split-GFP association". Scientific Reports. 3: 2854. Bibcode: 2013NatSR...3E2854C. doi: 10.1038/srep02854. PMC 3790201. PMID 24092409.
- ^ Martell JD, Yamagata M, Deerinck TJ, Phan S, Kwa CG, Ellisman MH, Sanes JR, Ting AY (July 2016). "A split horseradish peroxidase for the detection of intercellular protein-protein interactions and sensitive visualization of synapses" (PDF). Nature Biotechnology. 34 (7): 774–80. doi: 10.1038/nbt.3563. PMC 4942342. PMID 27240195.
- ^ Tchekanda E, Sivanesan D, Michnick SW (June 2014). "An infrared reporter to detect spatiotemporal dynamics of protein-protein interactions". Nature Methods. 11 (6): 641–4. doi: 10.1038/nmeth.2934. PMID 24747815. S2CID 1958433.
- ^ Rossi F, Charlton CA, Blau HM (August 1997). "Monitoring protein-protein interactions in intact eukaryotic cells by beta-galactosidase complementation". Proceedings of the National Academy of Sciences of the United States of America. 94 (16): 8405–10. Bibcode: 1997PNAS...94.8405R. doi: 10.1073/pnas.94.16.8405. PMC 22934. PMID 9237989.
- ^ Cassonnet P, Rolloy C, Neveu G, Vidalain PO, Chantier T, Pellet J, Jones L, Muller M, Demeret C, Gaud G, Vuillier F, Lotteau V, Tangy F, Favre M, Jacob Y (November 2011). "Benchmarking a luciferase complementation assay for detecting protein complexes". Nature Methods. 8 (12): 990–2. doi: 10.1038/nmeth.1773. PMID 22127214. S2CID 9377872.
- ^ Fujikawa, Y. et al. (2014) Split luciferase complementation assay to detect regulated protein-protein interactions in rice protoplasts in a large-scale format. Rice 7:11
- ^ Li YC, Rodewald LW, Hoppmann C, Wong ET, Lebreton S, Safar P, Patek M, Wang L, Wertman KF, Wahl GM (December 2014). "A versatile platform to analyze low-affinity and transient protein-protein interactions in living cells in real time". Cell Reports. 9 (5): 1946–58. doi: 10.1016/j.celrep.2014.10.058. PMC 4269221. PMID 25464845.
- ^ Neveu G, Cassonnet P, Vidalain PO, Rolloy C, Mendoza J, Jones L, Tangy F, Muller M, Demeret C, Tafforeau L, Lotteau V, Rabourdin-Combe C, Travé G, Dricot A, Hill DE, Vidal M, Favre M, Jacob Y (December 2012). "Comparative analysis of virus-host interactomes with a mammalian high-throughput protein complementation assay based on Gaussia princeps luciferase". Methods. 58 (4): 349–59. doi: 10.1016/j.ymeth.2012.07.029. PMC 3546263. PMID 22898364.
- ^ Binkowski B, Eggers C, Butler B, Schwinn M, Slater M, Machleidt T, Cong M, Wood K, Fan F (May 2016). "Monitoring intracellular protein interactions using NanoLuc® Binary Technology (NanoBiTTM)" (PDF). Promega.
- ^ Kolkhof P, Werthebach M, van de Venn A, Poschmann G, Chen L, Welte M, Stühler K, Beller M (March 2017). "A Luciferase-fragment Complementation Assay to Detect Lipid Droplet-associated Protein-Protein Interactions". Molecular & Cellular Proteomics. 16 (3): 329–345. doi: 10.1074/mcp.M116.061499. PMC 5340998. PMID 27956707.
- ^ Wehr MC, Laage R, Bolz U, Fischer TM, Grünewald S, Scheek S, Bach A, Nave KA, Rossner MJ (December 2006). "Monitoring regulated protein-protein interactions using split TEV". Nature Methods. 3 (12): 985–93. doi: 10.1038/nmeth967. PMID 17072307. S2CID 37120401.
- ^ Dünkler A, Müller J, Johnsson N (2012). "Detecting Protein–Protein Interactions with the Split-Ubiquitin Sensor". Gene Regulatory Networks. Methods in Molecular Biology. Vol. 786. pp. 115–30. doi: 10.1007/978-1-61779-292-2_7. ISBN 978-1-61779-291-5. PMID 21938623.
- ^ Titz, Björn; Rajagopala, Seesandra V.; Goll, Johannes; Häuser, Roman; McKevitt, Matthew T.; Palzkill, Timothy; Uetz, Peter (2008-05-28). "The binary protein interactome of Treponema pallidum--the syphilis spirochete". PLOS ONE. 3 (5): e2292. Bibcode: 2008PLoSO...3.2292T. doi: 10.1371/journal.pone.0002292. ISSN 1932-6203. PMC 2386257. PMID 18509523.
Further reading
- Rochette S, Diss G, Filteau M, Leducq JB, Dubé AK, Landry CR (March 2015). "Genome-wide protein-protein interaction screening by protein-fragment complementation assay (PCA) in living cells". Journal of Visualized Experiments (97). doi: 10.3791/52255. PMC 4401175. PMID 25867901.