| PDLIM3 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | PDLIM3, ALP, PDZ and LIM domain 3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 605889; MGI: 1859274; HomoloGene: 8710; GeneCards: PDLIM3; OMA: PDLIM3 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
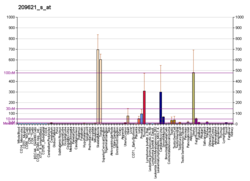
Actin-associated LIM protein (ALP), also known as PDZ and LIM domain protein 3 is a protein that in humans is encoded by the PDLIM3 gene. [5] [6] [7] ALP is highly expressed in cardiac and skeletal muscle, where it localizes to Z-discs and intercalated discs. ALP functions to enhance the crosslinking of actin by alpha-actinin-2 and also appears to be essential for right ventricular chamber formation and contractile function.
ALP exists primarily as two alternatively spliced variants; a 39.2 kDa (364 amino acids) protein in skeletal muscle and a 34.3 kDa (316 amino acids) protein in cardiac muscle and smooth muscle. [8] [9] [10] ALP has a N-terminal PDZ domain and a C-terminal LIM domain. In addition, the ALP subfamily contains a specific 34 amino acid domain named the ALP-like motif, containing protein kinase C consensus sequences. [11] The PDZ domain of ALP binds to alpha actinin-2, specifically to its spectrin-like repeats. [12] The PDZ domain is a motif composed of 80-120 amino acids with conserved four residue GLGF sequences that typically interact with C-termini of cytoskeletal proteins. [13] The region of heterogeneity in the two isoforms is between the PDZ domain and LIM domain. [10] ALP is localized to chromosome 4q35. [12] It has been shown that deletion of muscleblind-like 1 in mice can alter the splicing pattern of PDLIM3. [14]
Studies have shown that ALP is present at the first stage of myofibrilogenesis where it is bound to alpha actinin-2, and this association remains intact in mature myofibrils where ALP is localized to Z-discs and intercalated discs. Alpha actinin-2 is however not required for targeting ALP to Z-lines. [15] Studies in ALP knockout mice have shown that ALP facilitates the cross-linking of actin filaments by alpha actinin-2, and absence of ALP induces abnormal right ventricular chamber formation, dysplasia and cardiomyopathy. [16] Further studies using right ventricular epicardial systolic strain and geometric remodeling analysis in these animals unveiled that absence of ALP diminishes right ventricular contractile function and alters the pattern of cardiac hypertrophic remodeling. [17] Two studies using integrative genomic approaches to investigate genetic modifiers of collagen deposition [18] or intrinsic aerobic running capacity (ARC) [19] have mapped PDLIM3 to respective quantitative trait loci, suggesting that ALP may be involved in molecular networks related to these cardiac phenomena.
Chromosome 4 pericentric inversion has been observed in 10 patients, with associated cardiac defects linked to terminal 4q35.1 deletions, which may affect PDLIM3. [20]
ALP interacts with:
- ^ a b c GRCh38: Ensembl release 89: ENSG00000154553 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000031636 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Bouju S, Piétu G, Le Cunff M, Cros N, Malzac P, Pellissier JF, Pons F, Léger JJ, Auffray C, Dechesne CA (Jan 1999). "Exclusion of muscle specific actinin-associated LIM protein (ALP) gene from 4q35 facioscapulohumeral muscular dystrophy (FSHD) candidate genes". Neuromuscular Disorders. 9 (1): 3–10. doi: 10.1016/S0960-8966(98)00087-X. PMID 10063829. S2CID 24449614.
- ^ Piétu G, Alibert O, Guichard V, Lamy B, Bois F, Leroy E, Mariage-Sampson R, Houlgatte R, Soularue P, Auffray C (Jun 1996). "Novel gene transcripts preferentially expressed in human muscles revealed by quantitative hybridization of a high density cDNA array". Genome Research. 6 (6): 492–503. doi: 10.1101/gr.6.6.492. PMID 8828038.
- ^ "Entrez Gene: PDLIM3 PDZ and LIM domain 3".
- ^ "Protein sequence for human PDZ and LIM domain protein 3 (Uniprot: Q53GG5-2)". Cardiac Organellar Protein Atlas Knowledgebase (COPaKB). Archived from the original on 22 June 2015. Retrieved 22 June 2015.
- ^ "Protein sequence for human PDZ and LIM domain protein 3 (Uniprot: Q53GG5)". Cardiac Organellar Protein Atlas Knowledgebase (COPaKB). Archived from the original on 22 June 2015. Retrieved 22 June 2015.
- ^ a b Pomiès P, Macalma T, Beckerle MC (Oct 1999). "Purification and characterization of an alpha-actinin-binding PDZ-LIM protein that is up-regulated during muscle differentiation". The Journal of Biological Chemistry. 274 (41): 29242–50. doi: 10.1074/jbc.274.41.29242. PMID 10506181.
- ^ Te Velthuis AJ, Isogai T, Gerrits L, Bagowski CP (7 February 2007). "Insights into the molecular evolution of the PDZ/LIM family and identification of a novel conserved protein motif". PLOS ONE. 2 (2): e189. Bibcode: 2007PLoSO...2..189T. doi: 10.1371/journal.pone.0000189. PMC 1781342. PMID 17285143.
- ^ a b c Xia H, Winokur ST, Kuo WL, Altherr MR, Bredt DS (Oct 1997). "Actinin-associated LIM protein: identification of a domain interaction between PDZ and spectrin-like repeat motifs". The Journal of Cell Biology. 139 (2): 507–15. doi: 10.1083/jcb.139.2.507. PMC 2139795. PMID 9334352.
- ^ Ponting CP, Phillips C (Mar 1995). "DHR domains in syntrophins, neuronal NO synthases and other intracellular proteins". Trends in Biochemical Sciences. 20 (3): 102–3. doi: 10.1016/s0968-0004(00)88973-2. PMID 7535955.
- ^ Dixon DM, Choi J, El-Ghazali A, Park SY, Roos KP, Jordan MC, Fishbein MC, Comai L, Reddy S (12 March 2015). "Loss of muscleblind-like 1 results in cardiac pathology and persistence of embryonic splice isoforms". Scientific Reports. 5: 9042. Bibcode: 2015NatSR...5E9042D. doi: 10.1038/srep09042. PMC 4356957. PMID 25761764.
- ^ Henderson JR, Pomiès P, Auffray C, Beckerle MC (Mar 2003). "ALP and MLP distribution during myofibrillogenesis in cultured cardiomyocytes". Cell Motility and the Cytoskeleton. 54 (3): 254–65. doi: 10.1002/cm.10102. PMID 12589684.
- ^ a b Pashmforoush M, Pomiès P, Peterson KL, Kubalak S, Ross J, Hefti A, Aebi U, Beckerle MC, Chien KR (May 2001). "Adult mice deficient in actinin-associated LIM-domain protein reveal a developmental pathway for right ventricular cardiomyopathy". Nature Medicine. 7 (5): 591–7. doi: 10.1038/87920. PMID 11329061. S2CID 1781328.
- ^ Lorenzen-Schmidt I, McCulloch AD, Omens JH (Jul 2005). "Deficiency of actinin-associated LIM protein alters regional right ventricular function and hypertrophic remodeling". Annals of Biomedical Engineering. 33 (7): 888–96. doi: 10.1007/s10439-005-3604-y. PMC 4482468. PMID 16060528.
- ^ Lodder EM, Scicluna BP, Beekman L, Arends D, Moerland PD, Tanck MW, Adriaens ME, Bezzina CR (Dec 2014). "Integrative genomic approach identifies multiple genes involved in cardiac collagen deposition". Circulation: Cardiovascular Genetics. 7 (6): 790–8. doi: 10.1161/CIRCGENETICS.114.000537. PMID 25217174.
- ^ Lee SJ, Ways JA, Barbato JC, Essig D, Pettee K, DeRaedt SJ, Yang S, Weaver DA, Koch LG, Cicila GT (Sep 2005). "Gene expression profiling of the left ventricles in a rat model of intrinsic aerobic running capacity". Physiological Genomics. 23 (1): 62–71. CiteSeerX 10.1.1.321.9888. doi: 10.1152/physiolgenomics.00251.2004. PMID 16033863.
- ^ Maurin ML, Labrune P, Brisset S, Le Lorc'h M, Pineau D, Castel C, Romana S, Tachdjian G (Feb 2009). "Molecular cytogenetic characterization of a 4p15.1-pter duplication and a 4q35.1-qter deletion in a recombinant of chromosome 4 pericentric inversion". American Journal of Medical Genetics Part A. 149A (2): 226–31. doi: 10.1002/ajmg.a.32603. PMID 19161154. S2CID 205310317.
- Maruyama K, Sugano S (Jan 1994). "Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides". Gene. 138 (1–2): 171–4. doi: 10.1016/0378-1119(94)90802-8. PMID 8125298.
- Xia H, Winokur ST, Kuo WL, Altherr MR, Bredt DS (Oct 1997). "Actinin-associated LIM protein: identification of a domain interaction between PDZ and spectrin-like repeat motifs". The Journal of Cell Biology. 139 (2): 507–15. doi: 10.1083/jcb.139.2.507. PMC 2139795. PMID 9334352.
- Suzuki Y, Yoshitomo-Nakagawa K, Maruyama K, Suyama A, Sugano S (Oct 1997). "Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library". Gene. 200 (1–2): 149–56. doi: 10.1016/S0378-1119(97)00411-3. PMID 9373149.
- Klaavuniemi T, Kelloniemi A, Ylänne J (Jun 2004). "The ZASP-like motif in actinin-associated LIM protein is required for interaction with the alpha-actinin rod and for targeting to the muscle Z-line". The Journal of Biological Chemistry. 279 (25): 26402–10. doi: 10.1074/jbc.M401871200. PMID 15084604.
´* Brown AK, O'Connor PJ, Roberts TE, Wakefield RJ, Karim Z, Emery P (May 2006). "Ultrasonography for rheumatologists: the development of specific competency based educational outcomes". Annals of the Rheumatic Diseases. 65 (5): 629–36. doi: 10.1136/ard.2005.039974. PMC 1798129. PMID 16192291.
- Arola AM, Sanchez X, Murphy RT, Hasle E, Li H, Elliott PM, McKenna WJ, Towbin JA, Bowles NE (Apr 2007). "Mutations in PDLIM3 and MYOZ1 encoding myocyte Z line proteins are infrequently found in idiopathic dilated cardiomyopathy". Molecular Genetics and Metabolism. 90 (4): 435–40. doi: 10.1016/j.ymgme.2006.12.008. PMID 17254821.
| PDLIM3 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | PDLIM3, ALP, PDZ and LIM domain 3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 605889; MGI: 1859274; HomoloGene: 8710; GeneCards: PDLIM3; OMA: PDLIM3 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Actin-associated LIM protein (ALP), also known as PDZ and LIM domain protein 3 is a protein that in humans is encoded by the PDLIM3 gene. [5] [6] [7] ALP is highly expressed in cardiac and skeletal muscle, where it localizes to Z-discs and intercalated discs. ALP functions to enhance the crosslinking of actin by alpha-actinin-2 and also appears to be essential for right ventricular chamber formation and contractile function.
ALP exists primarily as two alternatively spliced variants; a 39.2 kDa (364 amino acids) protein in skeletal muscle and a 34.3 kDa (316 amino acids) protein in cardiac muscle and smooth muscle. [8] [9] [10] ALP has a N-terminal PDZ domain and a C-terminal LIM domain. In addition, the ALP subfamily contains a specific 34 amino acid domain named the ALP-like motif, containing protein kinase C consensus sequences. [11] The PDZ domain of ALP binds to alpha actinin-2, specifically to its spectrin-like repeats. [12] The PDZ domain is a motif composed of 80-120 amino acids with conserved four residue GLGF sequences that typically interact with C-termini of cytoskeletal proteins. [13] The region of heterogeneity in the two isoforms is between the PDZ domain and LIM domain. [10] ALP is localized to chromosome 4q35. [12] It has been shown that deletion of muscleblind-like 1 in mice can alter the splicing pattern of PDLIM3. [14]
Studies have shown that ALP is present at the first stage of myofibrilogenesis where it is bound to alpha actinin-2, and this association remains intact in mature myofibrils where ALP is localized to Z-discs and intercalated discs. Alpha actinin-2 is however not required for targeting ALP to Z-lines. [15] Studies in ALP knockout mice have shown that ALP facilitates the cross-linking of actin filaments by alpha actinin-2, and absence of ALP induces abnormal right ventricular chamber formation, dysplasia and cardiomyopathy. [16] Further studies using right ventricular epicardial systolic strain and geometric remodeling analysis in these animals unveiled that absence of ALP diminishes right ventricular contractile function and alters the pattern of cardiac hypertrophic remodeling. [17] Two studies using integrative genomic approaches to investigate genetic modifiers of collagen deposition [18] or intrinsic aerobic running capacity (ARC) [19] have mapped PDLIM3 to respective quantitative trait loci, suggesting that ALP may be involved in molecular networks related to these cardiac phenomena.
Chromosome 4 pericentric inversion has been observed in 10 patients, with associated cardiac defects linked to terminal 4q35.1 deletions, which may affect PDLIM3. [20]
ALP interacts with:
- ^ a b c GRCh38: Ensembl release 89: ENSG00000154553 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000031636 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Bouju S, Piétu G, Le Cunff M, Cros N, Malzac P, Pellissier JF, Pons F, Léger JJ, Auffray C, Dechesne CA (Jan 1999). "Exclusion of muscle specific actinin-associated LIM protein (ALP) gene from 4q35 facioscapulohumeral muscular dystrophy (FSHD) candidate genes". Neuromuscular Disorders. 9 (1): 3–10. doi: 10.1016/S0960-8966(98)00087-X. PMID 10063829. S2CID 24449614.
- ^ Piétu G, Alibert O, Guichard V, Lamy B, Bois F, Leroy E, Mariage-Sampson R, Houlgatte R, Soularue P, Auffray C (Jun 1996). "Novel gene transcripts preferentially expressed in human muscles revealed by quantitative hybridization of a high density cDNA array". Genome Research. 6 (6): 492–503. doi: 10.1101/gr.6.6.492. PMID 8828038.
- ^ "Entrez Gene: PDLIM3 PDZ and LIM domain 3".
- ^ "Protein sequence for human PDZ and LIM domain protein 3 (Uniprot: Q53GG5-2)". Cardiac Organellar Protein Atlas Knowledgebase (COPaKB). Archived from the original on 22 June 2015. Retrieved 22 June 2015.
- ^ "Protein sequence for human PDZ and LIM domain protein 3 (Uniprot: Q53GG5)". Cardiac Organellar Protein Atlas Knowledgebase (COPaKB). Archived from the original on 22 June 2015. Retrieved 22 June 2015.
- ^ a b Pomiès P, Macalma T, Beckerle MC (Oct 1999). "Purification and characterization of an alpha-actinin-binding PDZ-LIM protein that is up-regulated during muscle differentiation". The Journal of Biological Chemistry. 274 (41): 29242–50. doi: 10.1074/jbc.274.41.29242. PMID 10506181.
- ^ Te Velthuis AJ, Isogai T, Gerrits L, Bagowski CP (7 February 2007). "Insights into the molecular evolution of the PDZ/LIM family and identification of a novel conserved protein motif". PLOS ONE. 2 (2): e189. Bibcode: 2007PLoSO...2..189T. doi: 10.1371/journal.pone.0000189. PMC 1781342. PMID 17285143.
- ^ a b c Xia H, Winokur ST, Kuo WL, Altherr MR, Bredt DS (Oct 1997). "Actinin-associated LIM protein: identification of a domain interaction between PDZ and spectrin-like repeat motifs". The Journal of Cell Biology. 139 (2): 507–15. doi: 10.1083/jcb.139.2.507. PMC 2139795. PMID 9334352.
- ^ Ponting CP, Phillips C (Mar 1995). "DHR domains in syntrophins, neuronal NO synthases and other intracellular proteins". Trends in Biochemical Sciences. 20 (3): 102–3. doi: 10.1016/s0968-0004(00)88973-2. PMID 7535955.
- ^ Dixon DM, Choi J, El-Ghazali A, Park SY, Roos KP, Jordan MC, Fishbein MC, Comai L, Reddy S (12 March 2015). "Loss of muscleblind-like 1 results in cardiac pathology and persistence of embryonic splice isoforms". Scientific Reports. 5: 9042. Bibcode: 2015NatSR...5E9042D. doi: 10.1038/srep09042. PMC 4356957. PMID 25761764.
- ^ Henderson JR, Pomiès P, Auffray C, Beckerle MC (Mar 2003). "ALP and MLP distribution during myofibrillogenesis in cultured cardiomyocytes". Cell Motility and the Cytoskeleton. 54 (3): 254–65. doi: 10.1002/cm.10102. PMID 12589684.
- ^ a b Pashmforoush M, Pomiès P, Peterson KL, Kubalak S, Ross J, Hefti A, Aebi U, Beckerle MC, Chien KR (May 2001). "Adult mice deficient in actinin-associated LIM-domain protein reveal a developmental pathway for right ventricular cardiomyopathy". Nature Medicine. 7 (5): 591–7. doi: 10.1038/87920. PMID 11329061. S2CID 1781328.
- ^ Lorenzen-Schmidt I, McCulloch AD, Omens JH (Jul 2005). "Deficiency of actinin-associated LIM protein alters regional right ventricular function and hypertrophic remodeling". Annals of Biomedical Engineering. 33 (7): 888–96. doi: 10.1007/s10439-005-3604-y. PMC 4482468. PMID 16060528.
- ^ Lodder EM, Scicluna BP, Beekman L, Arends D, Moerland PD, Tanck MW, Adriaens ME, Bezzina CR (Dec 2014). "Integrative genomic approach identifies multiple genes involved in cardiac collagen deposition". Circulation: Cardiovascular Genetics. 7 (6): 790–8. doi: 10.1161/CIRCGENETICS.114.000537. PMID 25217174.
- ^ Lee SJ, Ways JA, Barbato JC, Essig D, Pettee K, DeRaedt SJ, Yang S, Weaver DA, Koch LG, Cicila GT (Sep 2005). "Gene expression profiling of the left ventricles in a rat model of intrinsic aerobic running capacity". Physiological Genomics. 23 (1): 62–71. CiteSeerX 10.1.1.321.9888. doi: 10.1152/physiolgenomics.00251.2004. PMID 16033863.
- ^ Maurin ML, Labrune P, Brisset S, Le Lorc'h M, Pineau D, Castel C, Romana S, Tachdjian G (Feb 2009). "Molecular cytogenetic characterization of a 4p15.1-pter duplication and a 4q35.1-qter deletion in a recombinant of chromosome 4 pericentric inversion". American Journal of Medical Genetics Part A. 149A (2): 226–31. doi: 10.1002/ajmg.a.32603. PMID 19161154. S2CID 205310317.
- Maruyama K, Sugano S (Jan 1994). "Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides". Gene. 138 (1–2): 171–4. doi: 10.1016/0378-1119(94)90802-8. PMID 8125298.
- Xia H, Winokur ST, Kuo WL, Altherr MR, Bredt DS (Oct 1997). "Actinin-associated LIM protein: identification of a domain interaction between PDZ and spectrin-like repeat motifs". The Journal of Cell Biology. 139 (2): 507–15. doi: 10.1083/jcb.139.2.507. PMC 2139795. PMID 9334352.
- Suzuki Y, Yoshitomo-Nakagawa K, Maruyama K, Suyama A, Sugano S (Oct 1997). "Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library". Gene. 200 (1–2): 149–56. doi: 10.1016/S0378-1119(97)00411-3. PMID 9373149.
- Klaavuniemi T, Kelloniemi A, Ylänne J (Jun 2004). "The ZASP-like motif in actinin-associated LIM protein is required for interaction with the alpha-actinin rod and for targeting to the muscle Z-line". The Journal of Biological Chemistry. 279 (25): 26402–10. doi: 10.1074/jbc.M401871200. PMID 15084604.
´* Brown AK, O'Connor PJ, Roberts TE, Wakefield RJ, Karim Z, Emery P (May 2006). "Ultrasonography for rheumatologists: the development of specific competency based educational outcomes". Annals of the Rheumatic Diseases. 65 (5): 629–36. doi: 10.1136/ard.2005.039974. PMC 1798129. PMID 16192291.
- Arola AM, Sanchez X, Murphy RT, Hasle E, Li H, Elliott PM, McKenna WJ, Towbin JA, Bowles NE (Apr 2007). "Mutations in PDLIM3 and MYOZ1 encoding myocyte Z line proteins are infrequently found in idiopathic dilated cardiomyopathy". Molecular Genetics and Metabolism. 90 (4): 435–40. doi: 10.1016/j.ymgme.2006.12.008. PMID 17254821.