This article has multiple issues. Please help
improve it or discuss these issues on the
talk page. (
Learn how and when to remove these template messages)
|

In molecular biology, hemagglutinins (alternatively spelt haemagglutinin, from the Greek haima, 'blood' + Latin gluten, 'glue') are receptor-binding membrane fusion glycoproteins produced by viruses in the Paramyxoviridae and Orthomyxoviridae families. [1] [2] Hemagglutinins are responsible for binding to receptors on host cells to initiate viral attachment and infection. [3]
Hemagglutinins recognize cell-surface glycoconjugates containing sialic acid on the surface of host red blood cells with a low affinity and use them to enter the endosome of host cells. [4] In the endosome, hemagglutinins are activated at a pH of 5–6.5 to undergo conformational changes that enable viral attachment through a fusion peptide. [5]
Virologist George K. Hirst discovered agglutination and hemagglutinins in 1941. [6] Alfred Gottschalk proved in 1957 that hemagglutinins bind a virus to a host cell by attaching to sialic acids on carbohydrate side chains of cell-membrane glycoproteins and glycolipids. [7]
The name "hemagglutinin" comes from the protein's ability to cause red blood cells (erythrocytes) to clump together (" agglutinate") in vitro. [8]
- Influenza hemagglutinin: a homotrimeric glycoprotein that is found on the surface of influenza viruses which is responsible for their infectivity. [9] Influenza strains are named for the specific hemagglutinin variant they produce, along with the specific variant of another surface protein, neuraminidase.
- Measles hemagglutinin: a hemagglutinin produced by the measles virus [10] that encodes six structural proteins, with hemagglutinin and fusion proteins being surface glycoproteins involved in attachment and entry. [11]
- Parainfluenza hemagglutinin-neuraminidase: a type of hemagglutinin-neuraminidase produced by parainfluenza, which is closely associated with both human and veterinary disease. [12]
- Mumps hemagglutinin-neuraminidase: a kind of hemagglutinin that the mumps virus (MuV) produces. [13]
- Hemagglutinin: the PH-E form of phytohaemagglutinin.
Hemagglutinins are small proteins that extend from the surface of the virus membrane as spikes that are 135 Angstroms (Å) in length and 30-50 Å in diameter.. [14] Each spike is composed of three identical monomer subunits, forming the protein a homotrimer. These monomers are formed of two glycopeptides, HA1 and HA2, and linked by two disulphide polypeptides, including membrane-distal HA1 and the smaller membrane-proximal HA2. X-ray crystallography and spectroscopy were used to identify that the majority of the protein structures is made of α-helical proteins. [15] In addition to the homotrimeric core structure, hemagglutinins have four subdomains: the membrane-distal receptor binding R subdomain, the vestigial domain E, that functions as a receptor-destroying esterase, the fusion domain F, and the membrane anchor subdomain M. The membrane anchor subdomain forms elastic protein chains linking the hemagglutinin to the ectodomain. [16]
- Hemagglutination Inhibition Assay: [17] A serologic assay which can be used either to screen for antibodies using RBCs with known surface antigens, or to identify RBCs surface antigens such as viruses or bacteria using a panel of known antibodies. This method, performed first by George K. Hirst in 1942, consists of mixing virus samples with serum dilutions so that antibodies bind to the virus before RBCs are added to the mix. Consequently, those viruses bound to antibodies are unable to link RBCs, meaning that a test’s positive result due to hemagglutination has been inhibited. On the contrary, if hemagglutination occurs, the test will result negative.
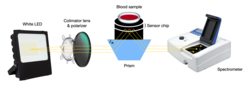
- Hemagglutination blood typing detection: [18] This method consists of measuring the blood’s reflectance spectrum alone (non-agglutination), and that of blood mixed with antibody reagents (agglutination) using a waveguide-mode sensor. As a result, some differences in reflectance between the samples are observed. Once antibodies are added, blood types and Rh(D) typing can be determined using the waveguide-mode sensor. This technique is able to detect weak agglutinations that are almost impossible to detect with the human eye.
- ABO blood group determination: Using anti-A and anti-B antibodies that bind specifically to either the A or to the B blood group surface antigens on RBCs, it is possible to test a small sample of blood and determine the ABO blood type of an individual. It does not identify the Rh(D) antigen ( Rh blood type).
- The bedside card method of blood grouping relies on visual agglutination to determine an individual's blood group. The card contains dried blood group antibody reagents fixed onto its surface. A drop of the individual's blood is placed on each blood group area on the card. The presence or absence of flocculation (visual agglutination) enables a quick and convenient method of determining the ABO and Rhesus status of the individual. As this technique depends on human eyes, it is less reliable than the blood typing based on waveguide-mode sensors.
- The agglutination of red blood cells is used in the Coombs test in diagnostic immunohematology to test for autoimmune hemolytic anemia. [19]
- In the case of red blood cells, transformed cells are known as kodecytes. Kode technology exposes exogenous antigens on the surface of cells, allowing antibody-antigen responses to be detected by the traditional hemagglutination test. [20]
- Cold agglutinin disease
- Hemagglutination assay
- Neuraminidase
- Influenza hemagglutinin (HA)
- Agglutination
- ^ Couch, Robert B. (1996), Baron, Samuel (ed.), "Orthomyxoviruses", Medical Microbiology (4th ed.), Galveston (TX): University of Texas Medical Branch at Galveston, ISBN 978-0-9631172-1-2, PMID 21413353, retrieved 30 January 2024
- ^ "Paramyxoviridae - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 30 January 2024.
- ^ Nobusawa, E. (October 1997). "[Structure and function of the hemagglutinin of influenza viruses]". Nihon Rinsho. Japanese Journal of Clinical Medicine. 55 (10): 2562–2569. ISSN 0047-1852. PMID 9360372.
- ^ Bangaru, Sandhya; Lang, Shanshan; Schotsaert, Michael; Vanderven, Hillary A.; Zhu, Xueyong; Kose, Nurgun; Bombardi, Robin; Finn, Jessica A.; Kent, Stephen J.; Gilchuk, Pavlo; Gilchuk, Iuliia (2019). "A Site of Vulnerability on the Influenza Virus Hemagglutinin Head Domain Trimer Interface". Cell. 177 (5): 1136–1152.e18. doi: 10.1016/j.cell.2019.04.011. PMC 6629437. PMID 31100268.
- ^ Medeiros, R.; Escriou, N.; Naffakh, N.; Manuguerra, J. C.; van der Werf, S. (10 October 2001). "Hemagglutinin residues of recent human A(H3N2) influenza viruses that contribute to the inability to agglutinate chicken erythrocytes". Virology. 289 (1): 74–85. doi: 10.1006/viro.2001.1121. ISSN 0042-6822. PMID 11601919.
- ^ Kolata, Gina (26 January 1994). "George Keble Hirst, 84, Is Dead; A Pioneer in Molecular Virology". The New York Times. ISSN 0362-4331. Retrieved 14 May 2024.
- ^ Henry, Ronnie; Murphy, Frederick A. (October 2018). "Etymologia: Hemagglutinin and Neuraminidase". Emerging Infectious Diseases. 24 (10): 1849. doi: 10.3201/eid2410.ET2410. PMC 6154157.
- ^ Nelson DL, Cox MM (2005). Lehninger's Principles of Biochemistry (4th ed.). New York: WH Freeman.
- ^ CDC (2 November 2021). "Types of Influenza Viruses". Centers for Disease Control and Prevention. Retrieved 18 October 2022.
- ^ Hashiguchi, Takao; Maenaka, Katsumi; Yanagi, Yusuke (16 December 2011). "Measles Virus Hemagglutinin: Structural Insights into Cell Entry and Measles Vaccine". Frontiers in Microbiology. 2: 247. doi: 10.3389/fmicb.2011.00247. ISSN 1664-302X. PMC 3267179. PMID 22319511.
-
^ Pan CH, Jimenez GS, Nair N (21 August 2014) [August, 2008].
"Use of Vaxfectin Adjuvant with DNA Vaccine Encoding the Measles Virus Hemagglutinin and Fusion Proteins Protects Juvenile and Infant Rhesus Macaques against Measles Virus". Clinical and Vaccine Immunology. 15 (8): 1214–1221.
doi:
10.1128/CVI.00120-08.
PMC
2519314.
PMID
18524884.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ Tappert, Mary M.; Porterfield, J. Zachary; Mehta-D'Souza, Padmaja; Gulati, Shelly; Air, Gillian M. (August 2013). "Quantitative Comparison of Human Parainfluenza Virus Hemagglutinin-Neuraminidase Receptor Binding and Receptor Cleavage". Journal of Virology. 87 (16): 8962–8970. doi: 10.1128/JVI.00739-13. ISSN 0022-538X. PMC 3754076. PMID 23740997.
- ^ Kubota, Marie; Hashiguchi, Takao (2020). "Large-Scale Expression and Purification of Mumps Virus Hemagglutinin-Neuraminidase for Structural Analyses and Glycan-Binding Assays". Lectin Purification and Analysis. Methods in Molecular Biology. Vol. 2132. pp. 641–652. doi: 10.1007/978-1-0716-0430-4_55. ISBN 978-1-0716-0429-8. ISSN 1940-6029. PMID 32306363. S2CID 216030421.
- ^ Gamblin, Steven J.; Vachieri, Sébastien G.; Xiong, Xiaoli; Zhang, Jie; Martin, Stephen R.; Skehel, John J. (1 October 2021). "Hemagglutinin Structure and Activities". Cold Spring Harbor Perspectives in Medicine. 11 (10): a038638. doi: 10.1101/cshperspect.a038638. ISSN 2157-1422. PMC 8485738. PMID 32513673.
- ^ Gamblin, Steven J.; Vachieri, Sébastien G.; Xiong, Xiaoli; Zhang, Jie; Martin, Stephen R.; Skehel, John J. (1 October 2021). "Hemagglutinin Structure and Activities". Cold Spring Harbor Perspectives in Medicine. 11 (10): a038638. doi: 10.1101/cshperspect.a038638. ISSN 2157-1422. PMC 8485738. PMID 32513673.
-
^ Donald J. Benton, Andrea Nans, Lesley J. Calder, Jack Turner, Ursula Neu, Yi Pu Lin, Esther Ketelaars, Nicole L. Kallewaard, Davide Corti, Antonio Lanzavecchia, Steven J. Gamblin, Peter B. Rosenthal, John J. Skehel (2 October 2018) [Sep 17, 2018].
"Hemagglutinin membrane anchor". Proceedings of the National Academy of Sciences of the United States of America. 115 (40): 10112–10117.
doi:
10.1073/pnas.1810927115.
PMC
6176637.
PMID
30224494.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ Payne, Susan (2017). "Methods to Study Viruses". Viruses. pp. 37–52. doi: 10.1016/B978-0-12-803109-4.00004-0. ISBN 978-0-12-803109-4. S2CID 89981392.
- ^ Ashiba, Hiroki; Fujimaki, Makoto; Awazu, Koichi; Fu, Mengying; Ohki, Yoshimichi; Tanaka, Torahiko; Makishima, Makoto (March 2015). "Hemagglutination detection for blood typing based on waveguide-mode sensors". Sensing and Bio-Sensing Research. 3: 59–64. doi: 10.1016/j.sbsr.2014.12.003.
- ^ Theis, Samuel R.; Hashmi, Muhammad F. (2022), "Coombs Test", StatPearls, Treasure Island (FL): StatPearls Publishing, PMID 31613487, retrieved 16 December 2022
- ^ Focosi, Daniele; Franchini, Massimo; Maggi, Fabrizio (8 March 2022). "Modified Hemagglutination Tests for COVID-19 Serology in Resource-Poor Settings: Ready for Prime-Time?". Vaccines. 10 (3): 406. doi: 10.3390/vaccines10030406. ISSN 2076-393X. PMC 8953758. PMID 35335038.
-
 Media related to
Haemagglutinin at Wikimedia Commons
Media related to
Haemagglutinin at Wikimedia Commons
This article has multiple issues. Please help
improve it or discuss these issues on the
talk page. (
Learn how and when to remove these template messages)
|

In molecular biology, hemagglutinins (alternatively spelt haemagglutinin, from the Greek haima, 'blood' + Latin gluten, 'glue') are receptor-binding membrane fusion glycoproteins produced by viruses in the Paramyxoviridae and Orthomyxoviridae families. [1] [2] Hemagglutinins are responsible for binding to receptors on host cells to initiate viral attachment and infection. [3]
Hemagglutinins recognize cell-surface glycoconjugates containing sialic acid on the surface of host red blood cells with a low affinity and use them to enter the endosome of host cells. [4] In the endosome, hemagglutinins are activated at a pH of 5–6.5 to undergo conformational changes that enable viral attachment through a fusion peptide. [5]
Virologist George K. Hirst discovered agglutination and hemagglutinins in 1941. [6] Alfred Gottschalk proved in 1957 that hemagglutinins bind a virus to a host cell by attaching to sialic acids on carbohydrate side chains of cell-membrane glycoproteins and glycolipids. [7]
The name "hemagglutinin" comes from the protein's ability to cause red blood cells (erythrocytes) to clump together (" agglutinate") in vitro. [8]
- Influenza hemagglutinin: a homotrimeric glycoprotein that is found on the surface of influenza viruses which is responsible for their infectivity. [9] Influenza strains are named for the specific hemagglutinin variant they produce, along with the specific variant of another surface protein, neuraminidase.
- Measles hemagglutinin: a hemagglutinin produced by the measles virus [10] that encodes six structural proteins, with hemagglutinin and fusion proteins being surface glycoproteins involved in attachment and entry. [11]
- Parainfluenza hemagglutinin-neuraminidase: a type of hemagglutinin-neuraminidase produced by parainfluenza, which is closely associated with both human and veterinary disease. [12]
- Mumps hemagglutinin-neuraminidase: a kind of hemagglutinin that the mumps virus (MuV) produces. [13]
- Hemagglutinin: the PH-E form of phytohaemagglutinin.
Hemagglutinins are small proteins that extend from the surface of the virus membrane as spikes that are 135 Angstroms (Å) in length and 30-50 Å in diameter.. [14] Each spike is composed of three identical monomer subunits, forming the protein a homotrimer. These monomers are formed of two glycopeptides, HA1 and HA2, and linked by two disulphide polypeptides, including membrane-distal HA1 and the smaller membrane-proximal HA2. X-ray crystallography and spectroscopy were used to identify that the majority of the protein structures is made of α-helical proteins. [15] In addition to the homotrimeric core structure, hemagglutinins have four subdomains: the membrane-distal receptor binding R subdomain, the vestigial domain E, that functions as a receptor-destroying esterase, the fusion domain F, and the membrane anchor subdomain M. The membrane anchor subdomain forms elastic protein chains linking the hemagglutinin to the ectodomain. [16]
- Hemagglutination Inhibition Assay: [17] A serologic assay which can be used either to screen for antibodies using RBCs with known surface antigens, or to identify RBCs surface antigens such as viruses or bacteria using a panel of known antibodies. This method, performed first by George K. Hirst in 1942, consists of mixing virus samples with serum dilutions so that antibodies bind to the virus before RBCs are added to the mix. Consequently, those viruses bound to antibodies are unable to link RBCs, meaning that a test’s positive result due to hemagglutination has been inhibited. On the contrary, if hemagglutination occurs, the test will result negative.

- Hemagglutination blood typing detection: [18] This method consists of measuring the blood’s reflectance spectrum alone (non-agglutination), and that of blood mixed with antibody reagents (agglutination) using a waveguide-mode sensor. As a result, some differences in reflectance between the samples are observed. Once antibodies are added, blood types and Rh(D) typing can be determined using the waveguide-mode sensor. This technique is able to detect weak agglutinations that are almost impossible to detect with the human eye.
- ABO blood group determination: Using anti-A and anti-B antibodies that bind specifically to either the A or to the B blood group surface antigens on RBCs, it is possible to test a small sample of blood and determine the ABO blood type of an individual. It does not identify the Rh(D) antigen ( Rh blood type).
- The bedside card method of blood grouping relies on visual agglutination to determine an individual's blood group. The card contains dried blood group antibody reagents fixed onto its surface. A drop of the individual's blood is placed on each blood group area on the card. The presence or absence of flocculation (visual agglutination) enables a quick and convenient method of determining the ABO and Rhesus status of the individual. As this technique depends on human eyes, it is less reliable than the blood typing based on waveguide-mode sensors.
- The agglutination of red blood cells is used in the Coombs test in diagnostic immunohematology to test for autoimmune hemolytic anemia. [19]
- In the case of red blood cells, transformed cells are known as kodecytes. Kode technology exposes exogenous antigens on the surface of cells, allowing antibody-antigen responses to be detected by the traditional hemagglutination test. [20]
- Cold agglutinin disease
- Hemagglutination assay
- Neuraminidase
- Influenza hemagglutinin (HA)
- Agglutination
- ^ Couch, Robert B. (1996), Baron, Samuel (ed.), "Orthomyxoviruses", Medical Microbiology (4th ed.), Galveston (TX): University of Texas Medical Branch at Galveston, ISBN 978-0-9631172-1-2, PMID 21413353, retrieved 30 January 2024
- ^ "Paramyxoviridae - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 30 January 2024.
- ^ Nobusawa, E. (October 1997). "[Structure and function of the hemagglutinin of influenza viruses]". Nihon Rinsho. Japanese Journal of Clinical Medicine. 55 (10): 2562–2569. ISSN 0047-1852. PMID 9360372.
- ^ Bangaru, Sandhya; Lang, Shanshan; Schotsaert, Michael; Vanderven, Hillary A.; Zhu, Xueyong; Kose, Nurgun; Bombardi, Robin; Finn, Jessica A.; Kent, Stephen J.; Gilchuk, Pavlo; Gilchuk, Iuliia (2019). "A Site of Vulnerability on the Influenza Virus Hemagglutinin Head Domain Trimer Interface". Cell. 177 (5): 1136–1152.e18. doi: 10.1016/j.cell.2019.04.011. PMC 6629437. PMID 31100268.
- ^ Medeiros, R.; Escriou, N.; Naffakh, N.; Manuguerra, J. C.; van der Werf, S. (10 October 2001). "Hemagglutinin residues of recent human A(H3N2) influenza viruses that contribute to the inability to agglutinate chicken erythrocytes". Virology. 289 (1): 74–85. doi: 10.1006/viro.2001.1121. ISSN 0042-6822. PMID 11601919.
- ^ Kolata, Gina (26 January 1994). "George Keble Hirst, 84, Is Dead; A Pioneer in Molecular Virology". The New York Times. ISSN 0362-4331. Retrieved 14 May 2024.
- ^ Henry, Ronnie; Murphy, Frederick A. (October 2018). "Etymologia: Hemagglutinin and Neuraminidase". Emerging Infectious Diseases. 24 (10): 1849. doi: 10.3201/eid2410.ET2410. PMC 6154157.
- ^ Nelson DL, Cox MM (2005). Lehninger's Principles of Biochemistry (4th ed.). New York: WH Freeman.
- ^ CDC (2 November 2021). "Types of Influenza Viruses". Centers for Disease Control and Prevention. Retrieved 18 October 2022.
- ^ Hashiguchi, Takao; Maenaka, Katsumi; Yanagi, Yusuke (16 December 2011). "Measles Virus Hemagglutinin: Structural Insights into Cell Entry and Measles Vaccine". Frontiers in Microbiology. 2: 247. doi: 10.3389/fmicb.2011.00247. ISSN 1664-302X. PMC 3267179. PMID 22319511.
-
^ Pan CH, Jimenez GS, Nair N (21 August 2014) [August, 2008].
"Use of Vaxfectin Adjuvant with DNA Vaccine Encoding the Measles Virus Hemagglutinin and Fusion Proteins Protects Juvenile and Infant Rhesus Macaques against Measles Virus". Clinical and Vaccine Immunology. 15 (8): 1214–1221.
doi:
10.1128/CVI.00120-08.
PMC
2519314.
PMID
18524884.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ Tappert, Mary M.; Porterfield, J. Zachary; Mehta-D'Souza, Padmaja; Gulati, Shelly; Air, Gillian M. (August 2013). "Quantitative Comparison of Human Parainfluenza Virus Hemagglutinin-Neuraminidase Receptor Binding and Receptor Cleavage". Journal of Virology. 87 (16): 8962–8970. doi: 10.1128/JVI.00739-13. ISSN 0022-538X. PMC 3754076. PMID 23740997.
- ^ Kubota, Marie; Hashiguchi, Takao (2020). "Large-Scale Expression and Purification of Mumps Virus Hemagglutinin-Neuraminidase for Structural Analyses and Glycan-Binding Assays". Lectin Purification and Analysis. Methods in Molecular Biology. Vol. 2132. pp. 641–652. doi: 10.1007/978-1-0716-0430-4_55. ISBN 978-1-0716-0429-8. ISSN 1940-6029. PMID 32306363. S2CID 216030421.
- ^ Gamblin, Steven J.; Vachieri, Sébastien G.; Xiong, Xiaoli; Zhang, Jie; Martin, Stephen R.; Skehel, John J. (1 October 2021). "Hemagglutinin Structure and Activities". Cold Spring Harbor Perspectives in Medicine. 11 (10): a038638. doi: 10.1101/cshperspect.a038638. ISSN 2157-1422. PMC 8485738. PMID 32513673.
- ^ Gamblin, Steven J.; Vachieri, Sébastien G.; Xiong, Xiaoli; Zhang, Jie; Martin, Stephen R.; Skehel, John J. (1 October 2021). "Hemagglutinin Structure and Activities". Cold Spring Harbor Perspectives in Medicine. 11 (10): a038638. doi: 10.1101/cshperspect.a038638. ISSN 2157-1422. PMC 8485738. PMID 32513673.
-
^ Donald J. Benton, Andrea Nans, Lesley J. Calder, Jack Turner, Ursula Neu, Yi Pu Lin, Esther Ketelaars, Nicole L. Kallewaard, Davide Corti, Antonio Lanzavecchia, Steven J. Gamblin, Peter B. Rosenthal, John J. Skehel (2 October 2018) [Sep 17, 2018].
"Hemagglutinin membrane anchor". Proceedings of the National Academy of Sciences of the United States of America. 115 (40): 10112–10117.
doi:
10.1073/pnas.1810927115.
PMC
6176637.
PMID
30224494.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) - ^ Payne, Susan (2017). "Methods to Study Viruses". Viruses. pp. 37–52. doi: 10.1016/B978-0-12-803109-4.00004-0. ISBN 978-0-12-803109-4. S2CID 89981392.
- ^ Ashiba, Hiroki; Fujimaki, Makoto; Awazu, Koichi; Fu, Mengying; Ohki, Yoshimichi; Tanaka, Torahiko; Makishima, Makoto (March 2015). "Hemagglutination detection for blood typing based on waveguide-mode sensors". Sensing and Bio-Sensing Research. 3: 59–64. doi: 10.1016/j.sbsr.2014.12.003.
- ^ Theis, Samuel R.; Hashmi, Muhammad F. (2022), "Coombs Test", StatPearls, Treasure Island (FL): StatPearls Publishing, PMID 31613487, retrieved 16 December 2022
- ^ Focosi, Daniele; Franchini, Massimo; Maggi, Fabrizio (8 March 2022). "Modified Hemagglutination Tests for COVID-19 Serology in Resource-Poor Settings: Ready for Prime-Time?". Vaccines. 10 (3): 406. doi: 10.3390/vaccines10030406. ISSN 2076-393X. PMC 8953758. PMID 35335038.
-
 Media related to
Haemagglutinin at Wikimedia Commons
Media related to
Haemagglutinin at Wikimedia Commons