| C16orf82 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | C16orf82, TNT, chromosome 16 open reading frame 82 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | HomoloGene: 82387; GeneCards: C16orf82; OMA: C16orf82 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
C16orf82 is a protein that, in humans, is encoded by the C16orf82 gene. [3] C16orf82 encodes a 2285 nucleotide mRNA transcript which is translated into a 154 amino acid protein using a non-AUG (CUG) start codon. The gene has been shown to be largely expressed in the testis, tibial nerve, and the pituitary gland, although expression has been seen throughout a majority of tissue types. [4] [5] [6] The function of C16orf82 is not fully understood by the scientific community. [7]
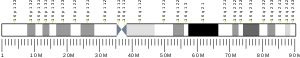
C16orf82 is located in humans at locus 16p12.1 on the positive strand.
The gene encodes for a 2285 nucleotide mRNA transcript that is intronless. Human intronless genes represent a unique subset of the genome that are often involved in signaling, sperm formation, immune responses, or development. [8] C16orf82 being such a gene indicates it may play a role in one of these processes. Translation of C16orf82 initiates at a non-AUG (CUG) start codon. The presence of the non-canonical start codon suggests possible increased regulation of C16orf82 translation and/or possibly could allow for the translation of protein products that start with leucine instead of methionine as seen in proteins coded for by some genes present in the major histocompatibility complex. [9] [10]
The C16orf82 promoter region has been predicted to contain a number of transcription factor binding sites including binding sites for transcription factors within the SOX family. [11] The presence of the SOX family transcription binding sites suggests that C16orf82 may play a role in sex determination. [12] Actual transcription factor functional studies show binding of the C16ORF82 promoter by ARNT, ELF5, SMAD4, and STAT3. [13]
C16orf82 expression in humans has been observed in major organ systems including the heart, liver, brain, and kidney at a constant level. [14] The tissue in which C16orf82 has been seen to be most highly expressed has been the testis, both by microarray experiments as well as RNA-seq. [4] [5] C16orf82 expression is also highly variable between individuals, with some expressing the gene in large amounts while others barely express the gene within the same tissue type. [6] [15] Micro RNA (miR-483) over expression has been shown to knock down C16orf82 expression. [16]
The C16orf82 protein is 154 amino acids in length with an approximate molecular weight of 16.46 kDa with a predicted isoelectric point of 6.06. [17] There are no known variants or isoforms of C16orf82.
C16orf82 contains one domain, DUF4694, which currently has a function that is uncharacterized. The domain spans from amino acid 8 to amino acid 153. [18] DUF4694 contains a SSGY (serine-serine-glycine-tyrosine) sequence motif that is found in a majority of the protein's orthologs. [19] [20] There is no presence of a transmembrane domain thus the protein is not a transmembrane protein. [21]

The localization of C16orf82 within a cell has been predicted to be nuclear. [21] A bipartite nuclear localization signal can be found starting at Arg107.

The human C16orf82 protein has been predicted to be phosphorylated at a number of serine residues. [26] O-linked glycosylation has also been predicted to happen at a number of sites, including some that overlap with the aforementioned phosphorylation sites. [27] The sites of overlap between the two types of post-translational modifications could play important regulatory roles in the activity and lifespan of the human C16orf82 protein. [28]
The secondary structure of the human C16orf82 protein has been predicted to be largely disordered by a number of modeling programs. [29] [30] [31] [32]
No paralogs of C16orf82 exist within humans. [20]
C16orf82 has over 100 predicted orthologs, which all reside in the class mammalia and more precisely the subclass eutheria. [33] [20] All of the orthologs contained the domain DUF4964. [33] The most distant ortholog detected was within the nine-banded armadillo (Dasypus novemcinctus) within the order Cingluata. Below is a table of 20 orthologs from various orders within the subclass eutheria with the sequence identity and time since divergence in relation to humans.
| Genus and Species | Common Name | Date of divergence (Mya) [34] | Accession number [35] | Protein Sequence length [35] | Sequence Identity (%) |
| Homo sapiens | Human | 0 | NP_001139017.1 | 154 | 100 |
| Gorilla gorilla gorilla | Gorilla | 9.06 | XP_004057433.1 | 217 | 97 |
| Saimiri boliviensis boliviensis | Bolivian squirrel monkey | 43.2 | XP_003945340.1 | 217 | 81 |
| Carlito syrichta | Philippine tarsier | 67.1 | XP_008059656.1 | 194 | 54 |
| Tupaia chinensis | Chinese tree shrew | 82 | XP_006148346.2 | 211 | 54 |
| Ochotona princeps | American pika | 90 | XP_004587173.1 | 184 | 46 |
| Oryctolagus cuniculus | Rabbit | 90 | XP_008256138.1 | 207 | 49 |
| Microtus ochrogaster | Prairie Vole | 90 | XP_005372535.1 | 180 | 48 |
| Fukomys damarensis | Damara mole-rat | 90 | XP_010621795.1 | 188 | 47 |
| Enhydra lutris kenyoni | Northern Sea Otter | 96 | XP_022382137.1 | 168 | 46 |
| Mustela putorius furo | domestic ferret | 96 | XP_012901961.1 | 173 | 46 |
| Canis lupus familiaris | Dog | 96 | NP_001139232.1 | 158 | 50 |
| Condylura cristata | star-nosed mole | 96 | XP_004696008.1 | 199 | 40 |
| Bos taurus | Cattle | 96 | NP_001139230.1 | 156 | 56 |
| Bison bison bison | American Bison | 96 | XP_010835728.1 | 197 | 55 |
| Capra hircus | Goat | 96 | XP_013830092.1 | 201 | 54 |
| Balaenoptera acutorostrata scammoni | Minke Whale | 96 | XP_007187042.1 | 206 | 52 |
| Equus Caballus | Horse | 96 | N/A | 153 | 47 |
| Hipposideros armiger | Great Roundleaf Bat | 96 | XP_019505352.1 | 192 | 63 |
| Loxodonta africana | African savanna elephant | 105 | XP_023414770.1 | 183 | 53 |
| Dasypus novemcinctus | nine-banded armadillo | 105 | XP_012377635.1 | 238 | 49 |

C16orf82's rate of evolution was determined to be relatively fast even in comparison to fibrinogen, a gene that has been shown to evolve quickly. [36]
C16orf82 has been associated with Schizophrenia through a genome-wide association study and autism based on copy number variation analysis. [37] [38] Currently, research has not shown if C16orf82 plays any direct role in either of these disorders.
- ^ a b c GRCh38: Ensembl release 89: ENSG00000234186 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b "C16orf82 chromosome 16 open reading frame 82 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-02-05.
- ^ a b Sato T, Kaneda A, Tsuji S, Isagawa T, Yamamoto S, Fujita T, Yamanaka R, Tanaka Y, Nukiwa T, Marquez VE, Ishikawa Y, Ichinose M, Aburatani H (2013-05-29). "PRC2 overexpression and PRC2-target gene repression relating to poorer prognosis in small cell lung cancer". Scientific Reports. 3: 1911. Bibcode: 2013NatSR...3E1911S. doi: 10.1038/srep01911. PMC 3665955. PMID 23714854.
- ^ a b Ardlie KG, Deluca DS, Segrè AV, Sullivan TJ, Young TR, Gelfand ET, et al. (GTEx Consortium) (May 2015). "Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans". Science. 348 (6235): 648–60. doi: 10.1126/science.1262110. PMC 4547484. PMID 25954001.
- ^ a b Jelinsky SA, Rodeo SA, Li J, Gulotta LV, Archambault JM, Seeherman HJ (May 2011). "Regulation of gene expression in human tendinopathy". BMC Musculoskeletal Disorders. 12: 86. doi: 10.1186/1471-2474-12-86. PMC 3095578. PMID 21539748.
- ^ Database, GeneCards Human Gene. "C16orf82 Gene - GeneCards | TNT Protein | TNT Antibody". www.genecards.org. Retrieved 2018-02-19.
- ^ Grzybowska EA (July 2012). "Human intronless genes: functional groups, associated diseases, evolution, and mRNA processing in absence of splicing". Biochemical and Biophysical Research Communications. 424 (1): 1–6. doi: 10.1016/j.bbrc.2012.06.092. PMID 22732409.
- ^ Glass NL (November 2017). "Near-Cognate Codons Contribute Complexity to Translation Regulation". mBio. 8 (6): e01820–17. doi: 10.1128/mbio.01820-17. PMC 5676045. PMID 29114030.
- ^ Starck SR, Jiang V, Pavon-Eternod M, Prasad S, McCarthy B, Pan T, Shastri N (June 2012). "Leucine-tRNA initiates at CUG start codons for protein synthesis and presentation by MHC class I". Science. 336 (6089): 1719–23. Bibcode: 2012Sci...336.1719S. doi: 10.1126/science.1220270. PMID 22745432. S2CID 206540614.
- ^ "Genomatix: Login Page". www.genomatix.de. Retrieved 2018-04-22.
- ^ Barrionuevo F, Scherer G (March 2010). "SOX E genes: SOX9 and SOX8 in mammalian testis development". The International Journal of Biochemistry & Cell Biology. 42 (3): 433–6. doi: 10.1016/j.biocel.2009.07.015. PMID 19647095.
- ^ Lachmann A, Xu H, Krishnan J, Berger SI, Mazloom AR, Ma'ayan A (October 2010). "ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments". Bioinformatics. 26 (19): 2438–44. doi: 10.1093/bioinformatics/btq466. PMC 2944209. PMID 20709693.
- ^ Yanai I, Benjamin H, Shmoish M, Chalifa-Caspi V, Shklar M, Ophir R, Bar-Even A, Horn-Saban S, Safran M, Domany E, Lancet D, Shmueli O (March 2005). "Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification". Bioinformatics. 21 (5): 650–9. doi: 10.1093/bioinformatics/bti042. PMID 15388519.
- ^ Goh SH, Josleyn M, Lee YT, Danner RL, Gherman RB, Cam MC, Miller JL (July 2007). "The human reticulocyte transcriptome". Physiological Genomics. 30 (2): 172–8. doi: 10.1152/physiolgenomics.00247.2006. PMID 17405831.
- ^ Liu M, Roth A, Yu M, Morris R, Bersani F, Rivera MN, Lu J, Shioda T, Vasudevan S, Ramaswamy S, Maheswaran S, Diederichs S, Haber DA (December 2013). "The IGF2 intronic miR-483 selectively enhances transcription from IGF2 fetal promoters and enhances tumorigenesis". Genes & Development. 27 (23): 2543–8. doi: 10.1101/gad.224170.113. PMC 3861668. PMID 24298054.
- ^ Walker, John M. (2005). Walker, John M (ed.). The Proteomics Protocols Handbook. pp. 571–607. doi: 10.1385/1592598900. ISBN 978-1-58829-343-5. S2CID 43080491.
- ^ "protein TNT [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ group, NIH/NLM/NCBI/IEB/CDD. "NCBI CDD Conserved Protein Domain DUF4694". www.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ a b c d "Protein BLAST: search protein databases using a protein query". blast.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ a b Nakai K, Horton P (January 1999). "PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization". Trends in Biochemical Sciences. 24 (1): 34–6. doi: 10.1016/S0968-0004(98)01336-X. PMID 10087920.
- ^ Sigrist CJ, de Castro E, Cerutti L, Cuche BA, Hulo N, Bridge A, Bougueleret L, Xenarios I (January 2013). "New and continuing developments at PROSITE". Nucleic Acids Research. 41 (Database issue): D344-7. doi: 10.1093/nar/gks1067. PMC 3531220. PMID 23161676.
- ^ Zhang Y (January 2008). "I-TASSER server for protein 3D structure prediction". BMC Bioinformatics. 9: 40. doi: 10.1186/1471-2105-9-40. PMC 2245901. PMID 18215316.
- ^ Roy A, Kucukural A, Zhang Y (April 2010). "I-TASSER: a unified platform for automated protein structure and function prediction". Nature Protocols. 5 (4): 725–38. doi: 10.1038/nprot.2010.5. PMC 2849174. PMID 20360767.
- ^ Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (January 2015). "The I-TASSER Suite: protein structure and function prediction". Nature Methods. 12 (1): 7–8. doi: 10.1038/nmeth.3213. PMC 4428668. PMID 25549265.
- ^ Blom N, Gammeltoft S, Brunak S (December 1999). "Sequence and structure-based prediction of eukaryotic protein phosphorylation sites". Journal of Molecular Biology. 294 (5): 1351–62. doi: 10.1006/jmbi.1999.3310. PMID 10600390.
- ^ Steentoft C, Vakhrushev SY, Joshi HJ, Kong Y, Vester-Christensen MB, Schjoldager KT, Lavrsen K, Dabelsteen S, Pedersen NB, Marcos-Silva L, Gupta R, Bennett EP, Mandel U, Brunak S, Wandall HH, Levery SB, Clausen H (May 2013). "Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology". The EMBO Journal. 32 (10): 1478–88. doi: 10.1038/emboj.2013.79. PMC 3655468. PMID 23584533.
- ^ Funakoshi Y, Suzuki T (February 2009). "Glycobiology in the cytosol: the bitter side of a sweet world". Biochimica et Biophysica Acta (BBA) - General Subjects. 1790 (2): 81–94. doi: 10.1016/j.bbagen.2008.09.009. PMID 18952151.
-
^ Zhou Y, Kloczkowski A, Faraggi E, Yang Y (2016-10-28). Prediction of protein secondary structure. New York, NY.
ISBN
9781493964048.
OCLC
961911230.
{{ cite book}}: CS1 maint: location missing publisher ( link) - ^ Drozdetskiy A, Cole C, Procter J, Barton GJ (July 2015). "JPred4: a protein secondary structure prediction server". Nucleic Acids Research. 43 (W1): W389-94. doi: 10.1093/nar/gkv332. PMC 4489285. PMID 25883141.
- ^ Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (June 2015). "The Phyre2 web portal for protein modeling, prediction and analysis". Nature Protocols. 10 (6): 845–58. doi: 10.1038/nprot.2015.053. PMC 5298202. PMID 25950237.
- ^ Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Gallo Cassarino T, Bertoni M, Bordoli L, Schwede T (July 2014). "SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information". Nucleic Acids Research. 42 (Web Server issue): W252-8. doi: 10.1093/nar/gku340. PMC 4086089. PMID 24782522.
- ^ a b "ortholog_gene_162083[group] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ Kumar S, Stecher G, Suleski M, Hedges SB (July 2017). "TimeTree: A Resource for Timelines, Timetrees, and Divergence Times". Molecular Biology and Evolution. 34 (7): 1812–1819. doi: 10.1093/molbev/msx116. PMID 28387841.
- ^ a b "Home - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-04-23.
- ^ a b Zhang J, Yang JR (July 2015). "Determinants of the rate of protein sequence evolution". Nature Reviews. Genetics. 16 (7): 409–20. doi: 10.1038/nrg3950. PMC 4523088. PMID 26055156.
- ^ McCarthy MJ, Nievergelt CM, Kelsoe JR, Welsh DK (2012-02-22). "A survey of genomic studies supports association of circadian clock genes with bipolar disorder spectrum illnesses and lithium response". PLOS ONE. 7 (2): e32091. Bibcode: 2012PLoSO...732091M. doi: 10.1371/journal.pone.0032091. PMC 3285204. PMID 22384149.
- ^ Wang LS, Hranilovic D, Wang K, Lindquist IE, Yurcaba L, Petkovic ZB, Gidaya N, Jernej B, Hakonarson H, Bucan M (September 2010). "Population-based study of genetic variation in individuals with autism spectrum disorders from Croatia". BMC Medical Genetics. 11: 134. doi: 10.1186/1471-2350-11-134. PMC 2954843. PMID 20858243.
| C16orf82 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | C16orf82, TNT, chromosome 16 open reading frame 82 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | HomoloGene: 82387; GeneCards: C16orf82; OMA: C16orf82 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
C16orf82 is a protein that, in humans, is encoded by the C16orf82 gene. [3] C16orf82 encodes a 2285 nucleotide mRNA transcript which is translated into a 154 amino acid protein using a non-AUG (CUG) start codon. The gene has been shown to be largely expressed in the testis, tibial nerve, and the pituitary gland, although expression has been seen throughout a majority of tissue types. [4] [5] [6] The function of C16orf82 is not fully understood by the scientific community. [7]
C16orf82 is located in humans at locus 16p12.1 on the positive strand.
The gene encodes for a 2285 nucleotide mRNA transcript that is intronless. Human intronless genes represent a unique subset of the genome that are often involved in signaling, sperm formation, immune responses, or development. [8] C16orf82 being such a gene indicates it may play a role in one of these processes. Translation of C16orf82 initiates at a non-AUG (CUG) start codon. The presence of the non-canonical start codon suggests possible increased regulation of C16orf82 translation and/or possibly could allow for the translation of protein products that start with leucine instead of methionine as seen in proteins coded for by some genes present in the major histocompatibility complex. [9] [10]
The C16orf82 promoter region has been predicted to contain a number of transcription factor binding sites including binding sites for transcription factors within the SOX family. [11] The presence of the SOX family transcription binding sites suggests that C16orf82 may play a role in sex determination. [12] Actual transcription factor functional studies show binding of the C16ORF82 promoter by ARNT, ELF5, SMAD4, and STAT3. [13]
C16orf82 expression in humans has been observed in major organ systems including the heart, liver, brain, and kidney at a constant level. [14] The tissue in which C16orf82 has been seen to be most highly expressed has been the testis, both by microarray experiments as well as RNA-seq. [4] [5] C16orf82 expression is also highly variable between individuals, with some expressing the gene in large amounts while others barely express the gene within the same tissue type. [6] [15] Micro RNA (miR-483) over expression has been shown to knock down C16orf82 expression. [16]
The C16orf82 protein is 154 amino acids in length with an approximate molecular weight of 16.46 kDa with a predicted isoelectric point of 6.06. [17] There are no known variants or isoforms of C16orf82.
C16orf82 contains one domain, DUF4694, which currently has a function that is uncharacterized. The domain spans from amino acid 8 to amino acid 153. [18] DUF4694 contains a SSGY (serine-serine-glycine-tyrosine) sequence motif that is found in a majority of the protein's orthologs. [19] [20] There is no presence of a transmembrane domain thus the protein is not a transmembrane protein. [21]

The localization of C16orf82 within a cell has been predicted to be nuclear. [21] A bipartite nuclear localization signal can be found starting at Arg107.

The human C16orf82 protein has been predicted to be phosphorylated at a number of serine residues. [26] O-linked glycosylation has also been predicted to happen at a number of sites, including some that overlap with the aforementioned phosphorylation sites. [27] The sites of overlap between the two types of post-translational modifications could play important regulatory roles in the activity and lifespan of the human C16orf82 protein. [28]
The secondary structure of the human C16orf82 protein has been predicted to be largely disordered by a number of modeling programs. [29] [30] [31] [32]
No paralogs of C16orf82 exist within humans. [20]
C16orf82 has over 100 predicted orthologs, which all reside in the class mammalia and more precisely the subclass eutheria. [33] [20] All of the orthologs contained the domain DUF4964. [33] The most distant ortholog detected was within the nine-banded armadillo (Dasypus novemcinctus) within the order Cingluata. Below is a table of 20 orthologs from various orders within the subclass eutheria with the sequence identity and time since divergence in relation to humans.
| Genus and Species | Common Name | Date of divergence (Mya) [34] | Accession number [35] | Protein Sequence length [35] | Sequence Identity (%) |
| Homo sapiens | Human | 0 | NP_001139017.1 | 154 | 100 |
| Gorilla gorilla gorilla | Gorilla | 9.06 | XP_004057433.1 | 217 | 97 |
| Saimiri boliviensis boliviensis | Bolivian squirrel monkey | 43.2 | XP_003945340.1 | 217 | 81 |
| Carlito syrichta | Philippine tarsier | 67.1 | XP_008059656.1 | 194 | 54 |
| Tupaia chinensis | Chinese tree shrew | 82 | XP_006148346.2 | 211 | 54 |
| Ochotona princeps | American pika | 90 | XP_004587173.1 | 184 | 46 |
| Oryctolagus cuniculus | Rabbit | 90 | XP_008256138.1 | 207 | 49 |
| Microtus ochrogaster | Prairie Vole | 90 | XP_005372535.1 | 180 | 48 |
| Fukomys damarensis | Damara mole-rat | 90 | XP_010621795.1 | 188 | 47 |
| Enhydra lutris kenyoni | Northern Sea Otter | 96 | XP_022382137.1 | 168 | 46 |
| Mustela putorius furo | domestic ferret | 96 | XP_012901961.1 | 173 | 46 |
| Canis lupus familiaris | Dog | 96 | NP_001139232.1 | 158 | 50 |
| Condylura cristata | star-nosed mole | 96 | XP_004696008.1 | 199 | 40 |
| Bos taurus | Cattle | 96 | NP_001139230.1 | 156 | 56 |
| Bison bison bison | American Bison | 96 | XP_010835728.1 | 197 | 55 |
| Capra hircus | Goat | 96 | XP_013830092.1 | 201 | 54 |
| Balaenoptera acutorostrata scammoni | Minke Whale | 96 | XP_007187042.1 | 206 | 52 |
| Equus Caballus | Horse | 96 | N/A | 153 | 47 |
| Hipposideros armiger | Great Roundleaf Bat | 96 | XP_019505352.1 | 192 | 63 |
| Loxodonta africana | African savanna elephant | 105 | XP_023414770.1 | 183 | 53 |
| Dasypus novemcinctus | nine-banded armadillo | 105 | XP_012377635.1 | 238 | 49 |

C16orf82's rate of evolution was determined to be relatively fast even in comparison to fibrinogen, a gene that has been shown to evolve quickly. [36]
C16orf82 has been associated with Schizophrenia through a genome-wide association study and autism based on copy number variation analysis. [37] [38] Currently, research has not shown if C16orf82 plays any direct role in either of these disorders.
- ^ a b c GRCh38: Ensembl release 89: ENSG00000234186 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b "C16orf82 chromosome 16 open reading frame 82 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-02-05.
- ^ a b Sato T, Kaneda A, Tsuji S, Isagawa T, Yamamoto S, Fujita T, Yamanaka R, Tanaka Y, Nukiwa T, Marquez VE, Ishikawa Y, Ichinose M, Aburatani H (2013-05-29). "PRC2 overexpression and PRC2-target gene repression relating to poorer prognosis in small cell lung cancer". Scientific Reports. 3: 1911. Bibcode: 2013NatSR...3E1911S. doi: 10.1038/srep01911. PMC 3665955. PMID 23714854.
- ^ a b Ardlie KG, Deluca DS, Segrè AV, Sullivan TJ, Young TR, Gelfand ET, et al. (GTEx Consortium) (May 2015). "Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans". Science. 348 (6235): 648–60. doi: 10.1126/science.1262110. PMC 4547484. PMID 25954001.
- ^ a b Jelinsky SA, Rodeo SA, Li J, Gulotta LV, Archambault JM, Seeherman HJ (May 2011). "Regulation of gene expression in human tendinopathy". BMC Musculoskeletal Disorders. 12: 86. doi: 10.1186/1471-2474-12-86. PMC 3095578. PMID 21539748.
- ^ Database, GeneCards Human Gene. "C16orf82 Gene - GeneCards | TNT Protein | TNT Antibody". www.genecards.org. Retrieved 2018-02-19.
- ^ Grzybowska EA (July 2012). "Human intronless genes: functional groups, associated diseases, evolution, and mRNA processing in absence of splicing". Biochemical and Biophysical Research Communications. 424 (1): 1–6. doi: 10.1016/j.bbrc.2012.06.092. PMID 22732409.
- ^ Glass NL (November 2017). "Near-Cognate Codons Contribute Complexity to Translation Regulation". mBio. 8 (6): e01820–17. doi: 10.1128/mbio.01820-17. PMC 5676045. PMID 29114030.
- ^ Starck SR, Jiang V, Pavon-Eternod M, Prasad S, McCarthy B, Pan T, Shastri N (June 2012). "Leucine-tRNA initiates at CUG start codons for protein synthesis and presentation by MHC class I". Science. 336 (6089): 1719–23. Bibcode: 2012Sci...336.1719S. doi: 10.1126/science.1220270. PMID 22745432. S2CID 206540614.
- ^ "Genomatix: Login Page". www.genomatix.de. Retrieved 2018-04-22.
- ^ Barrionuevo F, Scherer G (March 2010). "SOX E genes: SOX9 and SOX8 in mammalian testis development". The International Journal of Biochemistry & Cell Biology. 42 (3): 433–6. doi: 10.1016/j.biocel.2009.07.015. PMID 19647095.
- ^ Lachmann A, Xu H, Krishnan J, Berger SI, Mazloom AR, Ma'ayan A (October 2010). "ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments". Bioinformatics. 26 (19): 2438–44. doi: 10.1093/bioinformatics/btq466. PMC 2944209. PMID 20709693.
- ^ Yanai I, Benjamin H, Shmoish M, Chalifa-Caspi V, Shklar M, Ophir R, Bar-Even A, Horn-Saban S, Safran M, Domany E, Lancet D, Shmueli O (March 2005). "Genome-wide midrange transcription profiles reveal expression level relationships in human tissue specification". Bioinformatics. 21 (5): 650–9. doi: 10.1093/bioinformatics/bti042. PMID 15388519.
- ^ Goh SH, Josleyn M, Lee YT, Danner RL, Gherman RB, Cam MC, Miller JL (July 2007). "The human reticulocyte transcriptome". Physiological Genomics. 30 (2): 172–8. doi: 10.1152/physiolgenomics.00247.2006. PMID 17405831.
- ^ Liu M, Roth A, Yu M, Morris R, Bersani F, Rivera MN, Lu J, Shioda T, Vasudevan S, Ramaswamy S, Maheswaran S, Diederichs S, Haber DA (December 2013). "The IGF2 intronic miR-483 selectively enhances transcription from IGF2 fetal promoters and enhances tumorigenesis". Genes & Development. 27 (23): 2543–8. doi: 10.1101/gad.224170.113. PMC 3861668. PMID 24298054.
- ^ Walker, John M. (2005). Walker, John M (ed.). The Proteomics Protocols Handbook. pp. 571–607. doi: 10.1385/1592598900. ISBN 978-1-58829-343-5. S2CID 43080491.
- ^ "protein TNT [Homo sapiens] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ group, NIH/NLM/NCBI/IEB/CDD. "NCBI CDD Conserved Protein Domain DUF4694". www.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ a b c d "Protein BLAST: search protein databases using a protein query". blast.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ a b Nakai K, Horton P (January 1999). "PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization". Trends in Biochemical Sciences. 24 (1): 34–6. doi: 10.1016/S0968-0004(98)01336-X. PMID 10087920.
- ^ Sigrist CJ, de Castro E, Cerutti L, Cuche BA, Hulo N, Bridge A, Bougueleret L, Xenarios I (January 2013). "New and continuing developments at PROSITE". Nucleic Acids Research. 41 (Database issue): D344-7. doi: 10.1093/nar/gks1067. PMC 3531220. PMID 23161676.
- ^ Zhang Y (January 2008). "I-TASSER server for protein 3D structure prediction". BMC Bioinformatics. 9: 40. doi: 10.1186/1471-2105-9-40. PMC 2245901. PMID 18215316.
- ^ Roy A, Kucukural A, Zhang Y (April 2010). "I-TASSER: a unified platform for automated protein structure and function prediction". Nature Protocols. 5 (4): 725–38. doi: 10.1038/nprot.2010.5. PMC 2849174. PMID 20360767.
- ^ Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (January 2015). "The I-TASSER Suite: protein structure and function prediction". Nature Methods. 12 (1): 7–8. doi: 10.1038/nmeth.3213. PMC 4428668. PMID 25549265.
- ^ Blom N, Gammeltoft S, Brunak S (December 1999). "Sequence and structure-based prediction of eukaryotic protein phosphorylation sites". Journal of Molecular Biology. 294 (5): 1351–62. doi: 10.1006/jmbi.1999.3310. PMID 10600390.
- ^ Steentoft C, Vakhrushev SY, Joshi HJ, Kong Y, Vester-Christensen MB, Schjoldager KT, Lavrsen K, Dabelsteen S, Pedersen NB, Marcos-Silva L, Gupta R, Bennett EP, Mandel U, Brunak S, Wandall HH, Levery SB, Clausen H (May 2013). "Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology". The EMBO Journal. 32 (10): 1478–88. doi: 10.1038/emboj.2013.79. PMC 3655468. PMID 23584533.
- ^ Funakoshi Y, Suzuki T (February 2009). "Glycobiology in the cytosol: the bitter side of a sweet world". Biochimica et Biophysica Acta (BBA) - General Subjects. 1790 (2): 81–94. doi: 10.1016/j.bbagen.2008.09.009. PMID 18952151.
-
^ Zhou Y, Kloczkowski A, Faraggi E, Yang Y (2016-10-28). Prediction of protein secondary structure. New York, NY.
ISBN
9781493964048.
OCLC
961911230.
{{ cite book}}: CS1 maint: location missing publisher ( link) - ^ Drozdetskiy A, Cole C, Procter J, Barton GJ (July 2015). "JPred4: a protein secondary structure prediction server". Nucleic Acids Research. 43 (W1): W389-94. doi: 10.1093/nar/gkv332. PMC 4489285. PMID 25883141.
- ^ Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (June 2015). "The Phyre2 web portal for protein modeling, prediction and analysis". Nature Protocols. 10 (6): 845–58. doi: 10.1038/nprot.2015.053. PMC 5298202. PMID 25950237.
- ^ Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Gallo Cassarino T, Bertoni M, Bordoli L, Schwede T (July 2014). "SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information". Nucleic Acids Research. 42 (Web Server issue): W252-8. doi: 10.1093/nar/gku340. PMC 4086089. PMID 24782522.
- ^ a b "ortholog_gene_162083[group] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-02-19.
- ^ Kumar S, Stecher G, Suleski M, Hedges SB (July 2017). "TimeTree: A Resource for Timelines, Timetrees, and Divergence Times". Molecular Biology and Evolution. 34 (7): 1812–1819. doi: 10.1093/molbev/msx116. PMID 28387841.
- ^ a b "Home - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2018-04-23.
- ^ a b Zhang J, Yang JR (July 2015). "Determinants of the rate of protein sequence evolution". Nature Reviews. Genetics. 16 (7): 409–20. doi: 10.1038/nrg3950. PMC 4523088. PMID 26055156.
- ^ McCarthy MJ, Nievergelt CM, Kelsoe JR, Welsh DK (2012-02-22). "A survey of genomic studies supports association of circadian clock genes with bipolar disorder spectrum illnesses and lithium response". PLOS ONE. 7 (2): e32091. Bibcode: 2012PLoSO...732091M. doi: 10.1371/journal.pone.0032091. PMC 3285204. PMID 22384149.
- ^ Wang LS, Hranilovic D, Wang K, Lindquist IE, Yurcaba L, Petkovic ZB, Gidaya N, Jernej B, Hakonarson H, Bucan M (September 2010). "Population-based study of genetic variation in individuals with autism spectrum disorders from Croatia". BMC Medical Genetics. 11: 134. doi: 10.1186/1471-2350-11-134. PMC 2954843. PMID 20858243.